kwimage.structs package¶
Submodules¶
Module contents¶
mkinit ~/code/kwimage/kwimage/structs/__init__.py -w –relative –nomod
A common thread in many kwimage.structs / kwannot objects is that they attempt to store multiple data elements using a single data structure when possible e.g. the classes are Boxes, Points, Detections, Coords, and not Box, Detection, Coord. The exceptions are Polygon, Heatmap, and Mask, where it made more sense to have one object-per item because each individual item is a reasonably sized chuck of data.
Another commonality is that objects have only two main attributes: .data and .meta. These allow the underlying representation of the object to vary as needed.
Currently Boxes and Mask do not have a .meta attribute. They instead have a .format attribute which is a text-code indicating the underlying layout of the data.
The data and meta instance attributes in the Points, Detections, and Heatmaps classes are dictionaries. These classes also have a __datakeys__ and __metakeys__ class attribute, which are lists of strings. These lists specify which keys are expected in each dictionary. For instance, Points.__datakeys__ = [‘xy’, ‘class_idxs’, ‘visible’] and Points.__metakeys__ = [‘classes’]. All objects in the data dictionary are expected to be aligned, whereas the meta dictionary is for auxillay data. For example in Points, the xy position data[‘xy’][i] is expected to have the class index data[‘class_idxs’][i]. By convention, a class index indexes into the list of category names stored in meta[‘classes’].
The Heatmap.data behaves slighly different than Points. Its data dictionary stores different per-pixel attributes like class probability scores, or offset vectors. The meta dictionary stores data like the originaly image dimensions (heatmaps are usually downsampled wrt the image that they correspond to) and the transformation matrices would warp the “data” space back onto the original image space.
Note that the developer can add any extra data or meta keys that they like, but they should keep in mind that all items in data should be aligned, whereas meta can contain arbitrary information.
- class kwimage.structs.Boxes(data, format=None, check=True)[source]¶
Bases:
_BoxConversionMixins,_BoxPropertyMixins,_BoxTransformMixins,_BoxDrawMixins,NiceReprConverts boxes between different formats as long as the last dimension contains 4 coordinates and the format is specified.
This is a convinience class, and should not not store the data for very long. The general idiom should be create class, convert data, and then get the raw data and let the class be garbage collected. This will help ensure that your code is portable and understandable if this class is not available.
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> import kwimage >>> import numpy as np >>> # Given an array / tensor that represents one or more boxes >>> data = np.array([[ 0, 0, 10, 10], >>> [ 5, 5, 50, 50], >>> [20, 0, 30, 10]]) >>> # The kwimage.Boxes data structure is a thin fast wrapper >>> # that provides methods for operating on the boxes. >>> # It requires that the user explicitly provide a code that denotes >>> # the format of the boxes (i.e. what each column represents) >>> boxes = kwimage.Boxes(data, 'ltrb') >>> # This means that there is no ambiguity about box format >>> # The representation string of the Boxes object demonstrates this >>> print('boxes = {!r}'.format(boxes)) boxes = <Boxes(ltrb, array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]))> >>> # if you pass this data around. You can convert to other formats >>> # For docs on available format codes see :class:`BoxFormat`. >>> # In this example we will convert (left, top, right, bottom) >>> # to (left-x, top-y, width, height). >>> boxes.toformat('xywh') <Boxes(xywh, array([[ 0, 0, 10, 10], [ 5, 5, 45, 45], [20, 0, 10, 10]]))> >>> # In addition to format conversion there are other operations >>> # We can quickly (using a C-backend) find IoUs >>> ious = boxes.ious(boxes) >>> print('{}'.format(ub.repr2(ious, nl=1, precision=2, with_dtype=False))) np.array([[1. , 0.01, 0. ], [0.01, 1. , 0.02], [0. , 0.02, 1. ]]) >>> # We can ask for the area of each box >>> print('boxes.area = {}'.format(ub.repr2(boxes.area, nl=0, with_dtype=False))) boxes.area = np.array([[ 100],[2025],[ 100]]) >>> # We can ask for the center of each box >>> print('boxes.center = {}'.format(ub.repr2(boxes.center, nl=1, with_dtype=False))) boxes.center = ( np.array([[ 5. ],[27.5],[25. ]]), np.array([[ 5. ],[27.5],[ 5. ]]), ) >>> # We can translate / scale the boxes >>> boxes.translate((10, 10)).scale(100) <Boxes(ltrb, array([[1000., 1000., 2000., 2000.], [1500., 1500., 6000., 6000.], [3000., 1000., 4000., 2000.]]))> >>> # We can clip the bounding boxes >>> boxes.translate((10, 10)).scale(100).clip(1200, 1200, 1700, 1800) <Boxes(ltrb, array([[1200., 1200., 1700., 1800.], [1500., 1500., 1700., 1800.], [1700., 1200., 1700., 1800.]]))> >>> # We can perform arbitrary warping of the boxes >>> # (note that if the transform is not axis aligned, the axis aligned >>> # bounding box of the transform result will be returned) >>> transform = np.array([[-0.83907153, 0.54402111, 0. ], >>> [-0.54402111, -0.83907153, 0. ], >>> [ 0. , 0. , 1. ]]) >>> boxes.warp(transform) <Boxes(ltrb, array([[ -8.3907153 , -13.8309264 , 5.4402111 , 0. ], [-39.23347095, -69.154632 , 23.00569785, -6.9154632 ], [-25.1721459 , -24.7113486 , -11.3412195 , -10.8804222 ]]))> >>> # Note, that we can transform the box to a Polygon for more >>> # accurate warping. >>> transform = np.array([[-0.83907153, 0.54402111, 0. ], >>> [-0.54402111, -0.83907153, 0. ], >>> [ 0. , 0. , 1. ]]) >>> warped_polys = boxes.to_polygons().warp(transform) >>> print(ub.repr2(warped_polys.data, sv=1)) [ <Polygon({ 'exterior': <Coords(data= array([[ 0. , 0. ], [ 5.4402111, -8.3907153], [ -2.9505042, -13.8309264], [ -8.3907153, -5.4402111], [ 0. , 0. ]]))>, 'interiors': [], })>, <Polygon({ 'exterior': <Coords(data= array([[ -1.4752521 , -6.9154632 ], [ 23.00569785, -44.67368205], [-14.752521 , -69.154632 ], [-39.23347095, -31.39641315], [ -1.4752521 , -6.9154632 ]]))>, 'interiors': [], })>, <Polygon({ 'exterior': <Coords(data= array([[-16.7814306, -10.8804222], [-11.3412195, -19.2711375], [-19.7319348, -24.7113486], [-25.1721459, -16.3206333], [-16.7814306, -10.8804222]]))>, 'interiors': [], })>, ] >>> # The kwimage.Boxes data structure is also convertable to >>> # several alternative data structures, like shapely, coco, and imgaug. >>> print(ub.repr2(boxes.to_shapely(), sv=1)) [ POLYGON ((0 0, 0 10, 10 10, 10 0, 0 0)), POLYGON ((5 5, 5 50, 50 50, 50 5, 5 5)), POLYGON ((20 0, 20 10, 30 10, 30 0, 20 0)), ] >>> # xdoctest: +REQUIRES(module:imgaug) >>> print(ub.repr2(boxes[0:1].to_imgaug(shape=(100, 100)), sv=1)) BoundingBoxesOnImage([BoundingBox(x1=0.0000, y1=0.0000, x2=10.0000, y2=10.0000, label=None)], shape=(100, 100)) >>> # xdoctest: -REQUIRES(module:imgaug) >>> print(ub.repr2(list(boxes.to_coco()), sv=1)) [ [0, 0, 10, 10], [5, 5, 45, 45], [20, 0, 10, 10], ] >>> # Finally, when you are done with your boxes object, you can >>> # unwrap the raw data by using the ``.data`` attribute >>> # all operations are done on this data, which gives the >>> # kwiamge.Boxes data structure almost no overhead when >>> # inserted into existing code. >>> print('boxes.data =\n{}'.format(ub.repr2(boxes.data, nl=1))) boxes.data = np.array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]], dtype=np.int64) >>> # xdoctest: +REQUIRES(module:torch) >>> # This data structure was designed for use with both torch >>> # and numpy, the underlying data can be either an array or tensor. >>> boxes.tensor() <Boxes(ltrb, tensor([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]))> >>> boxes.numpy() <Boxes(ltrb, array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]))>
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> # Demo of conversion methods >>> import kwimage >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh') <Boxes(xywh, array([[25, 30, 15, 10]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_xywh() <Boxes(xywh, array([[25, 30, 15, 10]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_cxywh() <Boxes(cxywh, array([[32.5, 35. , 15. , 10. ]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_ltrb() <Boxes(ltrb, array([[25, 30, 40, 40]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').scale(2).to_ltrb() <Boxes(ltrb, array([[50., 60., 80., 80.]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> kwimage.Boxes(torch.FloatTensor([[25, 30, 15, 20]]), 'xywh').scale(.1).to_ltrb() <Boxes(ltrb, tensor([[ 2.5000, 3.0000, 4.0000, 5.0000]]))>
Note
In the following examples we show cases where
Boxescan hold a single 1-dimensional box array. This is a holdover from an older codebase, and some functions may assume that the input is at least 2-D. Thus when representing a single bounding box it is best practice to view it as a list of 1 box. While many function will work in the 1-D case, not all functions have been tested and thus we cannot gaurentee correctness.Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes([25, 30, 15, 10], 'xywh') <Boxes(xywh, array([25, 30, 15, 10]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_xywh() <Boxes(xywh, array([25, 30, 15, 10]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_cxywh() <Boxes(cxywh, array([32.5, 35. , 15. , 10. ]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_ltrb() <Boxes(ltrb, array([25, 30, 40, 40]))> >>> Boxes([25, 30, 15, 10], 'xywh').scale(2).to_ltrb() <Boxes(ltrb, array([50., 60., 80., 80.]))> >>> # xdoctest: +REQUIRES(module:torch) >>> Boxes(torch.FloatTensor([[25, 30, 15, 20]]), 'xywh').scale(.1).to_ltrb() <Boxes(ltrb, tensor([[ 2.5000, 3.0000, 4.0000, 5.0000]]))>
Example
>>> datas = [ >>> [1, 2, 3, 4], >>> [[1, 2, 3, 4], [4, 5, 6, 7]], >>> [[[1, 2, 3, 4], [4, 5, 6, 7]]], >>> ] >>> formats = BoxFormat.cannonical >>> for format1 in formats: >>> for data in datas: >>> self = box1 = Boxes(data, format1) >>> for format2 in formats: >>> box2 = box1.toformat(format2) >>> back = box2.toformat(format1) >>> assert box1 == back
- classmethod random(num=1, scale=1.0, format='xywh', anchors=None, anchor_std=0.16666666666666666, tensor=False, rng=None)[source]¶
Makes random boxes; typically for testing purposes
- Parameters
num (int) – number of boxes to generate
scale (float | Tuple[float, float]) – size of imgdims
format (str) – format of boxes to be created (e.g. ltrb, xywh)
anchors (ndarray) – normalized width / heights of anchor boxes to perterb and randomly place. (must be in range 0-1)
anchor_std (float) – magnitude of noise applied to anchor shapes
tensor (bool) – if True, returns boxes in tensor format
rng (None | int | RandomState) – initial random seed
- Returns
random boxes
- Return type
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes.random(3, rng=0, scale=100) <Boxes(xywh, array([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> Boxes.random(3, rng=0, scale=100).tensor() <Boxes(xywh, tensor([[ 54, 54, 6, 17], [ 42, 64, 1, 25], [ 79, 38, 17, 14]]))> >>> anchors = np.array([[.5, .5], [.3, .3]]) >>> Boxes.random(3, rng=0, scale=100, anchors=anchors) <Boxes(xywh, array([[ 2, 13, 51, 51], [32, 51, 32, 36], [36, 28, 23, 26]]))>
Example
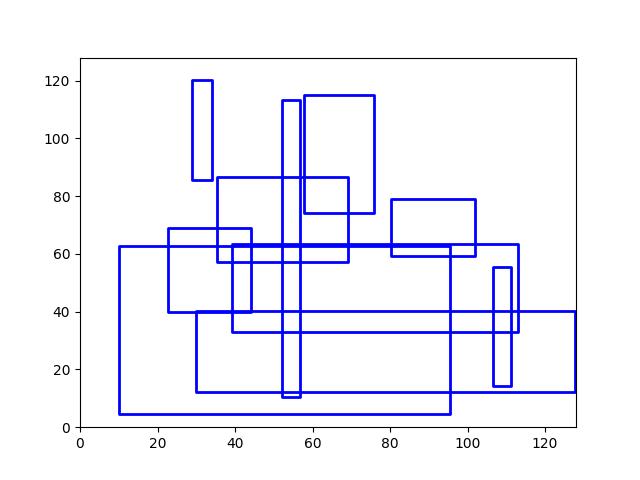
>>> # Boxes position/shape within 0-1 space should be uniform. >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> fig.gca().set_xlim(0, 128) >>> fig.gca().set_ylim(0, 128) >>> import kwimage >>> kwimage.Boxes.random(num=10).scale(128).draw()
- classmethod concatenate(boxes, axis=0)[source]¶
Concatenates multiple boxes together
- Parameters
boxes (Sequence[Boxes]) – list of boxes to concatenate
axis (int) – axis to stack on. Defaults to 0.
- Returns
stacked boxes
- Return type
Example
>>> boxes = [Boxes.random(3) for _ in range(3)] >>> new = Boxes.concatenate(boxes) >>> assert len(new) == 9 >>> assert np.all(new.data[3:6] == boxes[1].data)
Example
>>> boxes = [Boxes.random(3) for _ in range(3)] >>> boxes[0].data = boxes[0].data[0] >>> boxes[1].data = boxes[0].data[0:0] >>> new = Boxes.concatenate(boxes) >>> assert len(new) == 4 >>> # xdoctest: +REQUIRES(module:torch) >>> new = Boxes.concatenate([b.tensor() for b in boxes]) >>> assert len(new) == 4
- compress(flags, axis=0, inplace=False)[source]¶
Filters boxes based on a boolean criterion
- Parameters
flags (ArrayLike) – true for items to be kept. Extended type: ArrayLike[bool]
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns
the boxes corresponding to where flags were true
- Return type
Example
>>> self = Boxes([[25, 30, 15, 10]], 'ltrb') >>> self.compress([True]) <Boxes(ltrb, array([[25, 30, 15, 10]]))> >>> self.compress([False]) <Boxes(ltrb, array([], shape=(0, 4), dtype=int64))>
- take(idxs, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
- Parameters
indices (ArrayLike) – Indexes of items to take. Extended type ArrayLike[int].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns
the boxes corresponding to the specified indices
- Return type
Example
>>> self = Boxes([[25, 30, 15, 10]], 'ltrb') >>> self.take([0]) <Boxes(ltrb, array([[25, 30, 15, 10]]))> >>> self.take([]) <Boxes(ltrb, array([], shape=(0, 4), dtype=int64))>
- is_tensor()[source]¶
is the backend fueled by torch?
- Returns
True if the Boxes are torch tensors
- Return type
- is_numpy()[source]¶
is the backend fueled by numpy?
- Returns
True if the Boxes are numpy arrays
- Return type
- property device¶
If the backend is torch returns the data device, otherwise None
- astype(dtype)[source]¶
Changes the type of the internal array used to represent the boxes
Note
this operation is not inplace
- Returns
the boxes with the chosen type
- Return type
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> # xdoctest: +REQUIRES(module:torch) >>> Boxes.random(3, 100, rng=0).tensor().astype('int32') <Boxes(xywh, tensor([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]], dtype=torch.int32))> >>> Boxes.random(3, 100, rng=0).numpy().astype('int32') <Boxes(xywh, array([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]], dtype=int32))> >>> Boxes.random(3, 100, rng=0).tensor().astype('float32') >>> Boxes.random(3, 100, rng=0).numpy().astype('float32')
- round(inplace=False)[source]¶
Rounds data coordinates to the nearest integer.
This operation is applied directly to the box coordinates, so its output will depend on the format the boxes are stored in.
- Parameters
inplace (bool) – if True, modifies this object. Defaults to False.
- Returns
the boxes with rounded coordinates
- Return type
- SeeAlso:
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0).scale(10) >>> new = self.round() >>> print('self = {!r}'.format(self)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[5.48813522, 5.44883192, 0.53949833, 1.70306146], [4.23654795, 6.4589411 , 0.13932407, 2.45878875], [7.91725039, 3.83441508, 1.71937704, 1.45453393]]))> new = <Boxes(xywh, array([[5., 5., 1., 2.], [4., 6., 0., 2.], [8., 4., 2., 1.]]))>
- quantize(inplace=False, dtype=<class 'numpy.int32'>)[source]¶
Converts the box to integer coordinates.
This operation takes the floor of the left side and the ceil of the right side. Thus the area of the box will never decreases. But this will often increase the width / height of the box by a pixel.
- Parameters
inplace (bool) – if True, modifies this object
dtype (type) – type to cast as
- Returns
the boxes with quantized coordinates
- Return type
- SeeAlso:
Boxes.round()Boxes.resize()if you need to ensure the size does not change
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0).scale(10) >>> new = self.quantize() >>> print('self = {!r}'.format(self)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[5.48813522, 5.44883192, 0.53949833, 1.70306146], [4.23654795, 6.4589411 , 0.13932407, 2.45878875], [7.91725039, 3.83441508, 1.71937704, 1.45453393]]))> new = <Boxes(xywh, array([[5, 5, 2, 3], [4, 6, 1, 3], [7, 3, 3, 3]], dtype=int32))>
Example
>>> import kwimage >>> # Be careful if it is important to preserve the width/height >>> self = kwimage.Boxes([[0, 0, 10, 10]], 'xywh') >>> aff = kwimage.Affine.coerce(offset=(0.5, 0.0)) >>> warped = self.warp(aff) >>> new = warped.quantize(dtype=int) >>> print('self = {!r}'.format(self)) >>> print('warped = {!r}'.format(warped)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[ 0, 0, 10, 10]]))> warped = <Boxes(xywh, array([[ 0.5, 0. , 10. , 10. ]]))> new = <Boxes(xywh, array([[ 0, 0, 11, 10]]))>
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0) >>> orig = self.copy() >>> self.quantize(inplace=True) >>> assert np.any(self.data != orig.data)
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
- Returns
the boxes with a numpy backend
- Return type
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(3).tensor() >>> newself = self.numpy() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
- Parameters
device (int | None | torch.device) – The torch device to put the backend tensors on
- Returns
the boxes with a torch backend
- Return type
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(3) >>> # xdoctest: +REQUIRES(module:torch) >>> newself = self.tensor() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- ious(other, bias=0, impl='auto', mode=None)[source]¶
Intersection over union.
Compute IOUs (intersection area over union area) between these boxes and another set of boxes. This is a symmetric measure of similarity between boxes.
Todo
- [ ] Add pairwise flag to toggle between one-vs-one and all-vs-all
computation. I.E. Add option for componentwise calculation.
- Parameters
other (Boxes) – boxes to compare IoUs against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
impl (str) – code to specify implementation used to ious. Can be either torch, py, c, or auto. Efficiency and the exact result will vary by implementation, but they will always be close. Some implementations only accept certain data types (e.g. impl=’c’, only accepts float32 numpy arrays). See ~/code/kwimage/dev/bench_bbox.py for benchmark details. On my system the torch impl was fastest (when the data was on the GPU). Defaults to ‘auto’
mode (str) – depricated, use impl
- Returns
the ious
- Return type
ndarray
- SeeAlso:
iooas - for a measure of coverage between boxes
Examples
>>> import kwimage >>> self = kwimage.Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = kwimage.Boxes(np.array([6, 2, 20, 10]), 'ltrb') >>> overlaps = self.ious(other, bias=1).round(2) >>> assert np.all(np.isclose(overlaps, [0.21, 0.63, 0.04])), repr(overlaps)
Examples
>>> import kwimage >>> boxes1 = kwimage.Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = kwimage.Boxes(np.array([[6, 2, 20, 10], >>> [100, 200, 300, 300]]), 'ltrb') >>> overlaps = boxes1.ious(other) >>> print('{}'.format(ub.repr2(overlaps, precision=2, nl=1))) np.array([[0.18, 0. ], [0.61, 0. ], [0. , 0. ]]...)
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes(np.empty(0), 'xywh').ious(Boxes(np.empty(4), 'xywh')).shape (0,) >>> #Boxes(np.empty(4), 'xywh').ious(Boxes(np.empty(0), 'xywh')).shape >>> Boxes(np.empty((0, 4)), 'xywh').ious(Boxes(np.empty((0, 4)), 'xywh')).shape (0, 0) >>> Boxes(np.empty((1, 4)), 'xywh').ious(Boxes(np.empty((0, 4)), 'xywh')).shape (1, 0) >>> Boxes(np.empty((0, 4)), 'xywh').ious(Boxes(np.empty((1, 4)), 'xywh')).shape (0, 1)
Examples
>>> # xdoctest: +REQUIRES(module:torch) >>> formats = BoxFormat.cannonical >>> istensors = [False, True] >>> results = {} >>> for format in formats: >>> for tensor in istensors: >>> boxes1 = Boxes.random(5, scale=10.0, rng=0, format=format, tensor=tensor) >>> boxes2 = Boxes.random(7, scale=10.0, rng=1, format=format, tensor=tensor) >>> ious = boxes1.ious(boxes2) >>> results[(format, tensor)] = ious >>> results = {k: v.numpy() if torch.is_tensor(v) else v for k, v in results.items() } >>> results = {k: v.tolist() for k, v in results.items()} >>> print(ub.repr2(results, sk=True, precision=3, nl=2)) >>> from functools import partial >>> assert ub.allsame(results.values(), partial(np.allclose, atol=1e-07))
- iooas(other, bias=0)[source]¶
Intersection over other area.
This is an asymetric measure of coverage. How much of the “other” boxes are covered by these boxes. It is the area of intersection between each pair of boxes and the area of the “other” boxes.
- SeeAlso:
ious - for a measure of similarity between boxes
- Parameters
other (Boxes) – boxes to compare IoOA against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
- Returns
the iooas
- Return type
ndarray
Examples
>>> self = Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = Boxes(np.array([[6, 2, 20, 10], [0, 0, 0, 3]]), 'xywh') >>> coverage = self.iooas(other, bias=0).round(2) >>> print('coverage = {!r}'.format(coverage))
- isect_area(other, bias=0)[source]¶
Intersection part of intersection over union computation
- Parameters
other (Boxes) – boxes to compare IoOA against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
- Returns
the iooas
- Return type
ndarray
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> self = Boxes.random(5, scale=10.0, rng=0, format='ltrb') >>> other = Boxes.random(3, scale=10.0, rng=1, format='ltrb') >>> isect = self.isect_area(other, bias=0) >>> ious_v1 = isect / ((self.area + other.area.T) - isect) >>> ious_v2 = self.ious(other, bias=0) >>> assert np.allclose(ious_v1, ious_v2)
- intersection(other)[source]¶
Componentwise intersection between two sets of Boxes
intersections of boxes are always boxes, so this works
- Parameters
other (Boxes) – boxes to intersect with this object. (must be of same length)
- Returns
the intersection geometry
- Return type
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.intersection(other) >>> new_area = np.nan_to_num(new.area).ravel() >>> alt_area = np.diag(self.isect_area(other)) >>> close = np.isclose(new_area, alt_area) >>> assert np.all(close)
- union_hull(other)[source]¶
Componentwise hull union between two sets of Boxes
NOTE: convert to polygon to do a real union.
- Parameters
other (Boxes) – boxes to union with this object. (must be of same length)
- Returns
unioned boxes
- Return type
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.union_hull(other) >>> new_area = np.nan_to_num(new.area).ravel()
- bounding_box()[source]¶
Returns the box that bounds all of the contained boxes
- Returns
a single box
- Return type
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.union_hull(other) >>> new_area = np.nan_to_num(new.area).ravel()
- contains(other)[source]¶
Determine of points are completely contained by these boxes
- Parameters
other (kwimage.Points) – points to test for containment. TODO: support generic data types
- Returns
- flags - N x M boolean matrix indicating which box
contains which points, where N is the number of boxes and M is the number of points.
- Return type
ArrayLike
Examples
>>> import kwimage >>> self = kwimage.Boxes.random(10).scale(10).round() >>> other = kwimage.Points.random(10).scale(10).round() >>> flags = self.contains(other) >>> flags = self.contains(self.xy_center) >>> assert np.all(np.diag(flags))
- view(*shape)[source]¶
Passthrough method to view or reshape
- Parameters
*shape (Tuple[int, …]) – new shape
- Returns
data with a different view
- Return type
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(6, scale=10.0, rng=0, format='xywh').tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4] >>> self = Boxes.random(6, scale=10.0, rng=0, format='ltrb').tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4]
- class kwimage.structs.Coords(data=None, meta=None)[source]¶
-
A data structure to store n-dimensional coordinate geometry.
Currently it is up to the user to maintain what coordinate system this geometry belongs to.
Note
This class was designed to hold coordinates in r/c format, but in general this class is anostic to dimension ordering as long as you are consistent. However, there are two places where this matters: (1) drawing and (2) gdal/imgaug-warping. In these places we will assume x/y for legacy reasons. This may change in the future.
The term axes with resepct to
Coordsalways refers to the final numpy axis. In other words the final numpy-axis represents ALL of the coordinate-axes.CommandLine
xdoctest -m kwimage.structs.coords Coords
Example
>>> from kwimage.structs.coords import * # NOQA >>> import kwarray >>> rng = kwarray.ensure_rng(0) >>> self = Coords.random(num=4, dim=3, rng=rng) >>> print('self = {}'.format(self)) self = <Coords(data= array([[0.5488135 , 0.71518937, 0.60276338], [0.54488318, 0.4236548 , 0.64589411], [0.43758721, 0.891773 , 0.96366276], [0.38344152, 0.79172504, 0.52889492]]))> >>> matrix = rng.rand(4, 4) >>> self.warp(matrix) <Coords(data= array([[0.71037426, 1.25229659, 1.39498435], [0.60799503, 1.26483447, 1.42073131], [0.72106004, 1.39057144, 1.38757508], [0.68384299, 1.23914654, 1.29258196]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[3.5488135 , 3.71518937, 3.60276338], [3.54488318, 3.4236548 , 3.64589411], [3.43758721, 3.891773 , 3.96366276], [3.38344152, 3.79172504, 3.52889492]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[6.5488135 , 6.71518937, 6.60276338], [6.54488318, 6.4236548 , 6.64589411], [6.43758721, 6.891773 , 6.96366276], [6.38344152, 6.79172504, 6.52889492]]))> >>> self.scale(2) <Coords(data= array([[13.09762701, 13.43037873, 13.20552675], [13.08976637, 12.8473096 , 13.29178823], [12.87517442, 13.783546 , 13.92732552], [12.76688304, 13.58345008, 13.05778984]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> self.tensor() >>> self.tensor().tensor().numpy().numpy() >>> self.numpy() >>> #self.draw_on()
- property dtype¶
- property dim¶
- property shape¶
- classmethod random(num=1, dim=2, rng=None, meta=None)[source]¶
Makes random coordinates; typically for testing purposes
- compress(flags, axis=0, inplace=False)[source]¶
Filters items based on a boolean criterion
- Parameters
flags (ArrayLike) – true for items to be kept. Extended type: ArrayLike[bool].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns
filtered coords
- Return type
Example
>>> import kwimage >>> self = kwimage.Coords.random(10, rng=0) >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self)) <Coords(data=array([], shape=(0, 2), dtype=float64))> >>> # xdoctest: +REQUIRES(module:torch) >>> self = self.tensor() >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self))
- take(indices, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
- Parameters
indices (ArrayLike) – indexes of items to take. Extended type ArrayLike[int].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns
filtered coords
- Return type
Example
>>> import kwimage >>> self = kwimage.Coords(np.array([[25, 30, 15, 10]])) >>> self.take([0]) <Coords(data=array([[25, 30, 15, 10]]))> >>> self.take([]) <Coords(data=array([], shape=(0, 4), dtype=int64))>
- astype(dtype, inplace=False)[source]¶
Changes the data type
- Parameters
dtype – new type
inplace (bool) – if True, modifies this object
- Returns
modified coordinates
- Return type
- round(decimals=0, inplace=False)[source]¶
Rounds data to the specified decimal place
- Parameters
inplace (bool) – if True, modifies this object
decimals (int) – number of decimal places to round to
- Returns
modified coordinates
- Return type
Example
>>> import kwimage >>> self = kwimage.Coords.random(3).scale(10) >>> self.round()
- view(*shape)[source]¶
Passthrough method to view or reshape
- Parameters
*shape – new shape of the data
- Returns
modified coordinates
- Return type
Example
>>> self = Coords.random(6, dim=4).numpy() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4] >>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(6, dim=4).tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4]
- classmethod concatenate(coords, axis=0)[source]¶
Concatenates lists of coordinates together
- Parameters
coords (Sequence[Coords]) – list of coords to concatenate
axis (int) – axis to stack on. Defaults to 0.
- Returns
stacked coords
- Return type
CommandLine
xdoctest -m kwimage.structs.coords Coords.concatenate
Example
>>> coords = [Coords.random(3) for _ in range(3)] >>> new = Coords.concatenate(coords) >>> assert len(new) == 9 >>> assert np.all(new.data[3:6] == coords[1].data)
- property device¶
If the backend is torch returns the data device, otherwise None
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
- Returns
modified coordinates
- Return type
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).numpy() >>> newself = self.tensor() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
- Returns
modified coordinates
- Return type
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).tensor() >>> newself = self.numpy() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- reorder_axes(new_order, inplace=False)[source]¶
Change the ordering of the coordinate axes.
- Parameters
new_order (Tuple[int]) –
new_order[i]should specify which axes in the original coordinates should be mapped to thei-thposition in the returned axes.inplace (bool) – if True, modifies data inplace
- Returns
modified coordinates
- Return type
Note
This is the ordering of the “columns” in final numpy axis, not the numpy axes themselves.
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords(data=np.array([ >>> [7, 11], >>> [13, 17], >>> [21, 23], >>> ])) >>> new = self.reorder_axes((1, 0)) >>> print('new = {!r}'.format(new)) new = <Coords(data= array([[11, 7], [17, 13], [23, 21]]))>
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.reorder_axes((1, 0)) >>> # Remapping using 1, 0 reverses the axes >>> assert np.all(new.data[:, 0] == self.data[:, 1]) >>> assert np.all(new.data[:, 1] == self.data[:, 0]) >>> # Remapping using 0, 1 does nothing >>> eye = self.reorder_axes((0, 1)) >>> assert np.all(eye.data == self.data) >>> # Remapping using 0, 0, destroys the 1-th column >>> bad = self.reorder_axes((0, 0)) >>> assert np.all(bad.data[:, 0] == self.data[:, 0]) >>> assert np.all(bad.data[:, 1] == self.data[:, 0])
- warp(transform, input_dims=None, output_dims=None, inplace=False)[source]¶
Generalized coordinate transform.
- Parameters
transform (GeometricTransform | ArrayLike | Augmenter | Callable) – scikit-image tranform, a 3x3 transformation matrix, an imgaug Augmenter, or generic callable which transforms an NxD ndarray.
input_dims (Tuple) – shape of the image these objects correspond to (only needed / used when transform is an imgaug augmenter)
output_dims (Tuple) – unused in non-raster structures, only exists for compatibility.
inplace (bool) – if True, modifies data inplace
- Returns
modified coordinates
- Return type
Note
Let D = self.dims
- transformation matrices can be either:
(D + 1) x (D + 1) # for homog
D x D # for scale / rotate
D x (D + 1) # for affine
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> transform = skimage.transform.AffineTransform(scale=(2, 2)) >>> new = self.warp(transform) >>> assert np.all(new.data == self.scale(2).data)
Doctest
>>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(np.eye(3)).data == self.data) >>> assert np.all(self.warp(np.eye(2)).data == self.data)
Doctest
>>> # xdoctest: +REQUIRES(module:osgeo) >>> from osgeo import osr >>> wgs84_crs = osr.SpatialReference() >>> wgs84_crs.ImportFromEPSG(4326) >>> dst_crs = osr.SpatialReference() >>> dst_crs.ImportFromEPSG(2927) >>> transform = osr.CoordinateTransformation(wgs84_crs, dst_crs) >>> self = Coords.random(10, rng=0) >>> new = self.warp(transform) >>> assert np.all(new.data != self.data)
>>> # Alternative using generic func >>> def _gdal_coord_tranform(pts): ... return np.array([transform.TransformPoint(x, y, 0)[0:2] ... for x, y in pts]) >>> alt = self.warp(_gdal_coord_tranform) >>> assert np.all(alt.data != self.data) >>> assert np.all(alt.data == new.data)
Doctest
>>> # can use a generic function >>> def func(xy): ... return np.zeros_like(xy) >>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(func).data == 0)
- to_imgaug(input_dims)[source]¶
Translate to an imgaug object
- Returns
imgaug data structure
- Return type
imgaug.KeypointsOnImage
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> import kwimage >>> import numpy as np >>> self = kwimage.Coords.random(10) >>> input_dims = (10, 10) >>> kpoi = self.to_imgaug(input_dims) >>> new = kwimage.Coords.from_imgaug(kpoi) >>> assert np.allclose(new.data, self.data)
- scale(factor, about=None, output_dims=None, inplace=False)[source]¶
Scale coordinates by a factor
- Parameters
factor (float | Tuple[float, float]) – scale factor as either a scalar or per-dimension tuple.
about (Tuple | None) – if unspecified scales about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns
modified coordinates
- Return type
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.scale(10) >>> assert new.data.max() <= 10
>>> self = Coords.random(10, rng=0) >>> self.data = (self.data * 10).astype(int) >>> new = self.scale(10) >>> assert new.data.dtype.kind == 'i' >>> new = self.scale(10.0) >>> assert new.data.dtype.kind == 'f'
- translate(offset, output_dims=None, inplace=False)[source]¶
Shift the coordinates
- Parameters
offset (float | Tuple[float, float]) – transation offset as either a scalar or a per-dimension tuple.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns
modified coordinates
- Return type
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=3, rng=0) >>> new = self.translate(10) >>> assert new.data.min() >= 10 >>> assert new.data.max() <= 11 >>> Coords.random(3, dim=3, rng=0) >>> Coords.random(3, dim=3, rng=0).translate((1, 2, 3))
- rotate(theta, about=None, output_dims=None, inplace=False)[source]¶
Rotate the coordinates about a point.
- Parameters
theta (float) – rotation angle in radians
about (Tuple | None) – if unspecified rotates about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns
modified coordinates
- Return type
Todo
[ ] Generalized ND Rotations?
References
https://math.stackexchange.com/questions/197772/gen-rot-matrix
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=2, rng=0) >>> theta = np.pi / 2 >>> new = self.rotate(theta)
>>> # Test rotate agrees with warp >>> sin_ = np.sin(theta) >>> cos_ = np.cos(theta) >>> rot_ = np.array([[cos_, -sin_], [sin_, cos_]]) >>> new2 = self.warp(rot_) >>> assert np.allclose(new.data, new2.data)
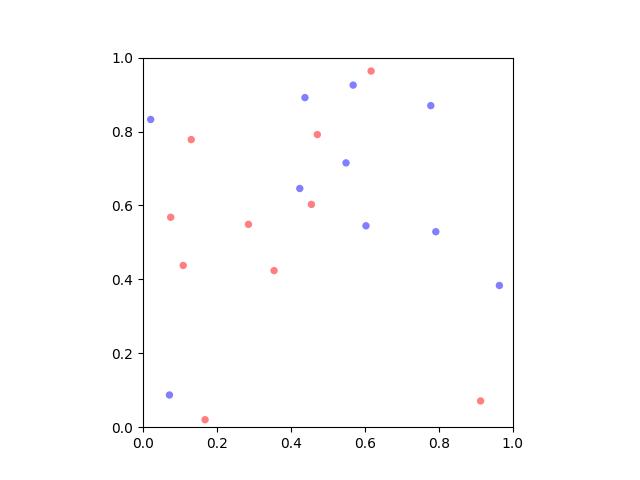
>>> # >>> # Rotate about a custom point >>> theta = np.pi / 2 >>> new3 = self.rotate(theta, about=(0.5, 0.5)) >>> # >>> # Rotate about the center of mass >>> about = self.data.mean(axis=0) >>> new4 = self.rotate(theta, about=about) >>> # xdoc: +REQUIRES(--show) >>> # xdoc: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> plt = kwplot.autoplt() >>> self.draw(radius=0.01, color='blue', alpha=.5, coord_axes=[1, 0], setlim='grow') >>> plt.gca().set_aspect('equal') >>> new3.draw(radius=0.01, color='red', alpha=.5, coord_axes=[1, 0], setlim='grow')
- fill(image, value, coord_axes=None, interp='bilinear')[source]¶
Sets sub-coordinate locations in a grid to a particular value
- Parameters
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns
image with coordinates rasterized on it
- Return type
ndarray
- soft_fill(image, coord_axes=None, radius=5)[source]¶
Used for drawing keypoint truth in heatmaps
- Parameters
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns
image with coordinates rasterized on it
- Return type
ndarray
References
https://stackoverflow.com/questions/54726703/generating-keypoint-heatmaps-in-tensorflow
Example
>>> from kwimage.structs.coords import * # NOQA >>> s = 64 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> # Put points on edges to to verify "edge cases" >>> self.data[1] = [0, 0] # top left >>> self.data[2] = [s, s] # bottom right >>> self.data[3] = [0, s + 10] # bottom left >>> self.data[4] = [-3, s // 2] # middle left >>> self.data[5] = [s + 1, -1] # top right >>> # Put points in the middle to verify overlap blending >>> self.data[6] = [32.5, 32.5] # middle >>> self.data[7] = [34.5, 34.5] # middle >>> fill_value = 1 >>> coord_axes = [1, 0] >>> radius = 10 >>> image1 = np.zeros((s, s)) >>> self.soft_fill(image1, coord_axes=coord_axes, radius=radius) >>> radius = 3.0 >>> image2 = np.zeros((s, s)) >>> self.soft_fill(image2, coord_axes=coord_axes, radius=radius) >>> # xdoc: +REQUIRES(--show) >>> # xdoc: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image1, pnum=(1, 2, 1)) >>> kwplot.imshow(image2, pnum=(1, 2, 2))

- draw_on(image=None, fill_value=1, coord_axes=[1, 0], interp='bilinear')[source]¶
Note
unlike other methods, the defaults assume x/y internal data
- Parameters
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns
image with coordinates drawn on it
- Return type
ndarray
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> s = 256 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> self.data[0] = [10, 10] >>> self.data[1] = [20, 40] >>> image = np.zeros((s, s)) >>> fill_value = 1 >>> image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[0, 1], interp='nearest') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='nearest') >>> # xdoc: +REQUIRES(--show) >>> # xdoc: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(image) >>> self.draw(radius=3, alpha=.5, coord_axes=[1, 0])

- draw(color='blue', ax=None, alpha=None, coord_axes=[1, 0], radius=1, setlim=False)[source]¶
Draw these coordinates via matplotlib
Note
unlike other methods, the defaults assume x/y internal data
- Parameters
setlim (bool) – if True ensures the limits of the axes contains the polygon
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns
drawn matplotlib objects
- Return type
List[mpl.collections.PatchCollection]
Example
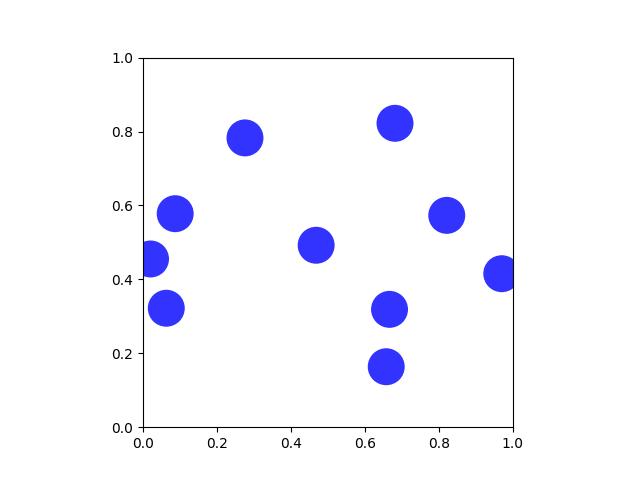
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> self.draw(radius=0.05, alpha=0.8) >>> plt.gca().set_xlim(0, 1) >>> plt.gca().set_ylim(0, 1) >>> plt.gca().set_aspect('equal')
- class kwimage.structs.Detections(data=None, meta=None, datakeys=None, metakeys=None, checks=True, **kwargs)[source]¶
Bases:
NiceRepr,_DetAlgoMixin,_DetDrawMixinContainer for holding and manipulating multiple detections.
- Variables
data (Dict) –
dictionary containing corresponding lists. The length of each list is the number of detections. This contains the bounding boxes, confidence scores, and class indices. Details of the most common keys and types are as follows:
boxes (kwimage.Boxes[ArrayLike]): multiple bounding boxes scores (ArrayLike): associated scores class_idxs (ArrayLike): associated class indices segmentations (ArrayLike): segmentations masks for each box, members can be
MaskorMultiPolygon. keypoints (ArrayLike): keypoints for each box. Members should bePoints.Additional custom keys may be specified as long as (a) the values are array-like and the first axis corresponds to the standard data values and (b) are custom keys are listed in the datakeys kwargs when constructing the Detections.
meta (Dict) – This contains contextual information about the detections. This includes the class names, which can be indexed into via the class indexes.
Example
>>> import kwimage >>> dets = kwimage.Detections( >>> # there are expected keys that do not need registration >>> boxes=kwimage.Boxes.random(3), >>> class_idxs=[0, 1, 1], >>> classes=['a', 'b'], >>> # custom data attrs must align with boxes >>> myattr1=np.random.rand(3), >>> myattr2=np.random.rand(3, 2, 8), >>> # there are no restrictions on metadata >>> mymeta='a custom metadata string', >>> # Note that any key not in kwimage.Detections.__datakeys__ or >>> # kwimage.Detections.__metakeys__ must be registered at the >>> # time of construction. >>> datakeys=['myattr1', 'myattr2'], >>> metakeys=['mymeta'], >>> checks=True, >>> ) >>> print('dets = {}'.format(dets)) dets = <Detections(3)>
- classmethod coerce(data=None, **kwargs)[source]¶
The “try-anything to get what I want” constructor
- Parameters
data
**kwargs – currently boxes and cnames
Example
>>> from kwimage.structs.detections import * # NOQA >>> import kwimage >>> kwargs = dict( >>> boxes=kwimage.Boxes.random(4), >>> cnames=['a', 'b', 'c', 'c'], >>> ) >>> data = {} >>> self = kwimage.Detections.coerce(data, **kwargs)
- classmethod from_coco_annots(anns, cats=None, classes=None, kp_classes=None, shape=None, dset=None)[source]¶
Create a Detections object from a list of coco-like annotations.
- Parameters
anns (List[Dict]) – list of coco-like annotation objects
dset (kwcoco.CocoDataset) – if specified, cats, classes, and kp_classes can are ignored.
cats (List[Dict]) – coco-format category information. Used only if dset is not specified.
classes (kwcoco.CategoryTree) – category tree with coco class info. Used only if dset is not specified.
kp_classes (kwcoco.CategoryTree) – keypoint category tree with coco keypoint class info. Used only if dset is not specified.
shape (tuple) – shape of parent image
- Returns
a detections object
- Return type
Example
>>> from kwimage.structs.detections import * # NOQA >>> # xdoctest: +REQUIRES(--module:ndsampler) >>> anns = [{ >>> 'id': 0, >>> 'image_id': 1, >>> 'category_id': 2, >>> 'bbox': [2, 3, 10, 10], >>> 'keypoints': [4.5, 4.5, 2], >>> 'segmentation': { >>> 'counts': '_11a04M2O0O20N101N3L_5', >>> 'size': [20, 20], >>> }, >>> }] >>> dataset = { >>> 'images': [], >>> 'annotations': [], >>> 'categories': [ >>> {'id': 0, 'name': 'background'}, >>> {'id': 2, 'name': 'class1', 'keypoints': ['spot']} >>> ] >>> } >>> #import ndsampler >>> #dset = ndsampler.CocoDataset(dataset) >>> cats = dataset['categories'] >>> dets = Detections.from_coco_annots(anns, cats)
Example
>>> # xdoctest: +REQUIRES(--module:ndsampler) >>> # Test case with no category information >>> from kwimage.structs.detections import * # NOQA >>> anns = [{ >>> 'id': 0, >>> 'image_id': 1, >>> 'category_id': None, >>> 'bbox': [2, 3, 10, 10], >>> 'prob': [.1, .9], >>> }] >>> cats = [ >>> {'id': 0, 'name': 'background'}, >>> {'id': 2, 'name': 'class1'} >>> ] >>> dets = Detections.from_coco_annots(anns, cats)
Example
>>> import kwimage >>> # xdoctest: +REQUIRES(--module:ndsampler) >>> import ndsampler >>> sampler = ndsampler.CocoSampler.demo('photos') >>> iminfo, anns = sampler.load_image_with_annots(1) >>> shape = iminfo['imdata'].shape[0:2] >>> kp_classes = sampler.dset.keypoint_categories() >>> dets = kwimage.Detections.from_coco_annots( >>> anns, sampler.dset.dataset['categories'], sampler.catgraph, >>> kp_classes, shape=shape)
- to_coco(cname_to_cat=None, style='orig', image_id=None, dset=None)[source]¶
Converts this set of detections into coco-like annotation dictionaries.
Note
Not all aspects of the MS-COCO format can be accurately represented, so some liberties are taken. The MS-COCO standard defines that annotations should specifiy a category_id field, but in some cases this information is not available so we will populate a ‘category_name’ field if possible and in the worst case fall back to ‘category_index’.
Additionally, detections may contain additional information beyond the MS-COCO standard, and this information (e.g. weight, prob, score) is added as forign fields.
- Parameters
cname_to_cat – currently ignored.
style (str) – either ‘orig’ (for the original coco format) or ‘new’ for the more general kwcoco-style coco format. Defaults to ‘orig’
image_id (int) – if specified, populates the image_id field of each image.
dset (kwcoco.CocoDataset | None) – if specified, attempts to populate the category_id field to be compatible with this coco dataset.
- Yields
dict – coco-like annotation structures
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> from kwimage.structs.detections import * >>> self = Detections.demo()[0] >>> cname_to_cat = None >>> list(self.to_coco())
- property boxes¶
- property class_idxs¶
- property scores¶
typically only populated for predicted detections
- property probs¶
typically only populated for predicted detections
- property weights¶
typically only populated for groundtruth detections
- property classes¶
- warp(transform, input_dims=None, output_dims=None, inplace=False)[source]¶
Spatially warp the detections.
- Parameters
transform (kwimage.Affine | ndarray | Callable | Any) – Something coercable to a transform. Usually a kwimage.Affine object
input_dims (Tuple[int, int]) – shape of the expected input canvas
output_dims (Tuple[int, int]) – shape of the expected output canvas
inplace (bool) – if true operate inplace
- Returns
the warped detections object
- Return type
Example
>>> import skimage >>> transform = skimage.transform.AffineTransform(scale=(2, 3), translation=(4, 5)) >>> self = Detections.random(2) >>> new = self.warp(transform) >>> assert new.boxes == self.boxes.warp(transform) >>> assert new != self
- scale(factor, output_dims=None, inplace=False)[source]¶
Spatially scale the detections.
Example
>>> import skimage >>> transform = skimage.transform.AffineTransform(scale=(2, 3), translation=(4, 5)) >>> self = Detections.random(2) >>> new = self.warp(transform) >>> assert new.boxes == self.boxes.warp(transform) >>> assert new != self
- translate(offset, output_dims=None, inplace=False)[source]¶
Spatially translate the detections.
Example
>>> import skimage >>> self = Detections.random(2) >>> new = self.translate(10)
- classmethod concatenate(dets)[source]¶
- Parameters
boxes (Sequence[Detections]) – list of detections to concatenate
- Returns
stacked detections
- Return type
Example
>>> self = Detections.random(2) >>> other = Detections.random(3) >>> dets = [self, other] >>> new = Detections.concatenate(dets) >>> assert new.num_boxes() == 5
>>> self = Detections.random(2, segmentations=True) >>> other = Detections.random(3, segmentations=True) >>> dets = [self, other] >>> new = Detections.concatenate(dets) >>> assert new.num_boxes() == 5
- argsort(reverse=True)[source]¶
Sorts detection indices by descending (or ascending) scores
- Returns
sorted indices torch.Tensor: sorted indices if using torch backends
- Return type
ndarray[Shape[‘*’], Integer]
- sort(reverse=True)[source]¶
Sorts detections by descending (or ascending) scores
- Returns
sorted copy of self
- Return type
- compress(flags, axis=0)[source]¶
Returns a subset where corresponding locations are True.
- Parameters
flags (ndarray[Any, Bool] | torch.Tensor) – mask marking selected items
- Returns
subset of self
- Return type
CommandLine
xdoctest -m kwimage.structs.detections Detections.compress
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> dets = kwimage.Detections.random(keypoints='dense') >>> flags = np.random.rand(len(dets)) > 0.5 >>> subset = dets.compress(flags) >>> assert len(subset) == flags.sum() >>> subset = dets.tensor().compress(flags) >>> assert len(subset) == flags.sum()
- take(indices, axis=0)[source]¶
Returns a subset specified by indices
- Parameters
indices (ndarray[Any, Integer]) – indices to select
- Returns
subset of self
- Return type
Example
>>> import kwimage >>> dets = kwimage.Detections(boxes=kwimage.Boxes.random(10)) >>> subset = dets.take([2, 3, 5, 7]) >>> assert len(subset) == 4 >>> # xdoctest: +REQUIRES(module:torch) >>> subset = dets.tensor().take([2, 3, 5, 7]) >>> assert len(subset) == 4
- property device¶
If the backend is torch returns the data device, otherwise None
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Detections.random(3).tensor() >>> newself = self.numpy() >>> self.scores[0] = 0 >>> assert newself.scores[0] == 0 >>> self.scores[0] = 1 >>> assert self.scores[0] == 1 >>> self.numpy().numpy()
- property dtype¶
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.detections import * >>> self = Detections.random(3) >>> newself = self.tensor() >>> self.scores[0] = 0 >>> assert newself.scores[0] == 0 >>> self.scores[0] = 1 >>> assert self.scores[0] == 1 >>> self.tensor().tensor()
- classmethod random(num=10, scale=1.0, classes=3, keypoints=False, segmentations=False, tensor=False, rng=None)[source]¶
Creates dummy data, suitable for use in tests and benchmarks
- Parameters
num (int) – number of boxes
scale (float | tuple) – bounding image size. Defaults to 1.0
classes (int | Sequence) – list of class labels or number of classes
keypoints (bool) – if True include random keypoints for each box. Defaults to False.
segmentations (bool) – if True include random segmentations for each box. Defaults to False.
tensor (bool) – determines backend. DEPRECATED. Call
.tensor()on resulting object instead.rng (np.random.RandomState) – random state
Example
>>> import kwimage >>> dets = kwimage.Detections.random(keypoints='jagged') >>> dets.data['keypoints'].data[0].data >>> dets.data['keypoints'].meta >>> dets = kwimage.Detections.random(keypoints='dense') >>> dets = kwimage.Detections.random(keypoints='dense', segmentations=True).scale(1000) >>> # xdoctest:+REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets.draw(setlim=True)
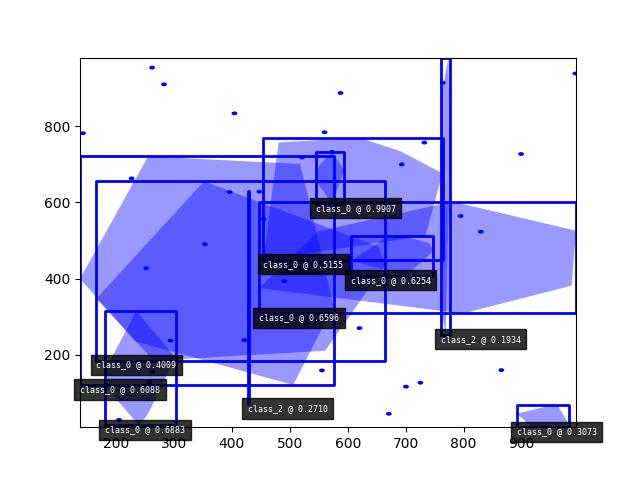
Example
>>> import kwimage >>> dets = kwimage.Detections.random( >>> keypoints='jagged', segmentations=True, rng=0).scale(1000) >>> print('dets = {}'.format(dets)) dets = <Detections(10)> >>> dets.data['boxes'].quantize(inplace=True) >>> print('dets.data = {}'.format(ub.repr2( >>> dets.data, nl=1, with_dtype=False, strvals=True))) dets.data = { 'boxes': <Boxes(xywh, array([[548, 544, 55, 172], [423, 645, 15, 247], [791, 383, 173, 146], [ 71, 87, 498, 839], [ 20, 832, 759, 39], [461, 780, 518, 20], [118, 639, 26, 306], [264, 414, 258, 361], [ 18, 568, 439, 50], [612, 616, 332, 66]], dtype=int32))>, 'class_idxs': [1, 2, 0, 0, 2, 0, 0, 0, 0, 0], 'keypoints': <PointsList(n=10)>, 'scores': [0.3595079 , 0.43703195, 0.6976312 , 0.06022547, 0.66676672, 0.67063787,0.21038256, 0.1289263 , 0.31542835, 0.36371077], 'segmentations': <SegmentationList(n=10)>, } >>> # xdoctest:+REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets.draw(setlim=True)
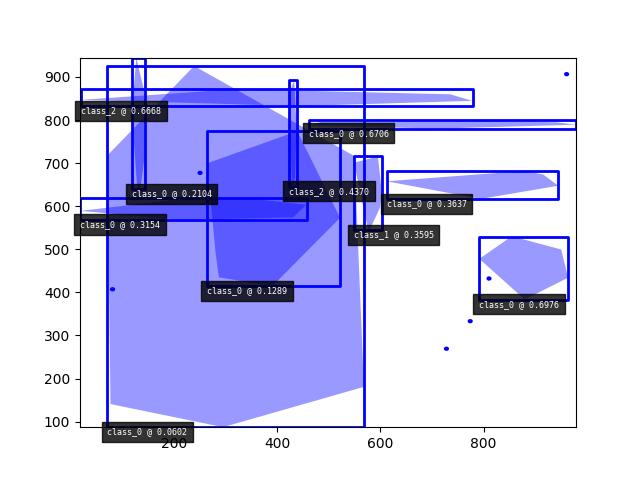
Example
>>> # Boxes position/shape within 0-1 space should be uniform. >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> fig.gca().set_xlim(0, 128) >>> fig.gca().set_ylim(0, 128) >>> import kwimage >>> kwimage.Detections.random(num=10, segmentations=True).scale(128).draw()
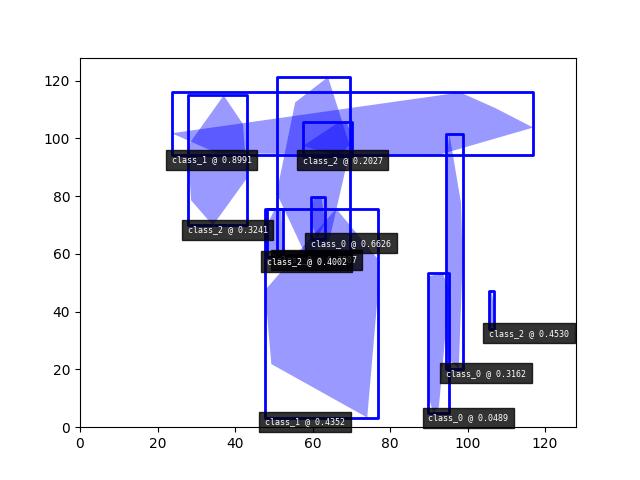
- class kwimage.structs.Heatmap(data=None, meta=None, **kwargs)[source]¶
Bases:
Spatial,_HeatmapDrawMixin,_HeatmapWarpMixin,_HeatmapAlgoMixinKeeps track of a downscaled heatmap and how to transform it to overlay the original input image. Heatmaps generally are used to estimate class probabilites at each pixel. This data struction additionally contains logic to augment pixel with offset (dydx) and scale (diamter) information.
- Variables
data (Dict[str, ArrayLike]) –
dictionary containing spatially aligned heatmap data. Valid keys are as follows.
- class_probs (ArrayLike[C, H, W] | ArrayLike[C, D, H, W]):
A probability map for each class. C is the number of classes.
- offset (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object center position offset in y,x / t,y,x coordinates
- diamter (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object bounding box sizes in h,w / d,h,w coordinates
- keypoints (ArrayLike[2, K, H, W] | ArrayLike[3, K, D, H, W], optional):
y/x offsets for K different keypoint classes
dictionary containing miscellanious metadata about the heatmap data. Valid keys are as follows.
- img_dims (Tuple[H, W] | Tuple[D, H, W]):
original image dimension
- tf_data_to_image (skimage.transform._geometric.GeometricTransform):
transformation matrix (typically similarity or affine) that projects the given, heatmap onto the image dimensions such that the image and heatmap are spatially aligned.
- classes (List[str] | ndsampler.CategoryTree):
information about which index in
data['class_probs']corresponds to which semantic class.
dims (Tuple) – dimensions of the heatmap (See
image_dims) for the original image dimensions.**kwargs – any key that is accepted by the data or meta dictionaries can be specified as a keyword argument to this class and it will be properly placed in the appropriate internal dictionary.
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/heatmap.py Heatmap --show
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> import kwimage >>> class_probs = kwimage.grab_test_image(dsize=(32, 32), space='gray')[None, ..., 0] / 255.0 >>> img_dims = (220, 220) >>> tf_data_to_img = skimage.transform.AffineTransform(translation=(-18, -18), scale=(8, 8)) >>> self = Heatmap(class_probs=class_probs, img_dims=img_dims, >>> tf_data_to_img=tf_data_to_img) >>> aligned = self.upscale() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(aligned[0]) >>> kwplot.show_if_requested()

Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> self = Heatmap.random() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw()

- property shape¶
- property bounds¶
- property dims¶
space-time dimensions of this heatmap
- classmethod random(dims=(10, 10), classes=3, diameter=True, offset=True, keypoints=False, img_dims=None, dets=None, nblips=10, noise=0.0, smooth_k=3, rng=None, ensure_background=True)[source]¶
Creates dummy data, suitable for use in tests and benchmarks
- Parameters
dims (Tuple[int, int]) – dimensions of the heatmap
classes (int | List[str] | kwcoco.CategoryTree) – foreground classes
diameter (bool) – if True, include a “diameter” heatmap
offset (bool) – if True, include an “offset” heatmap
keypoints (bool)
smooth_k (int) – kernel size for gaussian blur to smooth out the heatmaps.
img_dims (Tuple) – dimensions of an upscaled image the heatmap corresponds to. (This should be removed and simply handled with a transform
in the future).
- Returns
Heatmap
Example
>>> from kwimage.structs.heatmap import * # NOQA >>> self = Heatmap.random((128, 128), img_dims=(200, 200), >>> classes=3, nblips=10, rng=0, noise=0.1) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(self.colorize(0, imgspace=0), fnum=1, pnum=(1, 4, 1), doclf=1) >>> kwplot.imshow(self.colorize(1, imgspace=0), fnum=1, pnum=(1, 4, 2)) >>> kwplot.imshow(self.colorize(2, imgspace=0), fnum=1, pnum=(1, 4, 3)) >>> kwplot.imshow(self.colorize(3, imgspace=0), fnum=1, pnum=(1, 4, 4))
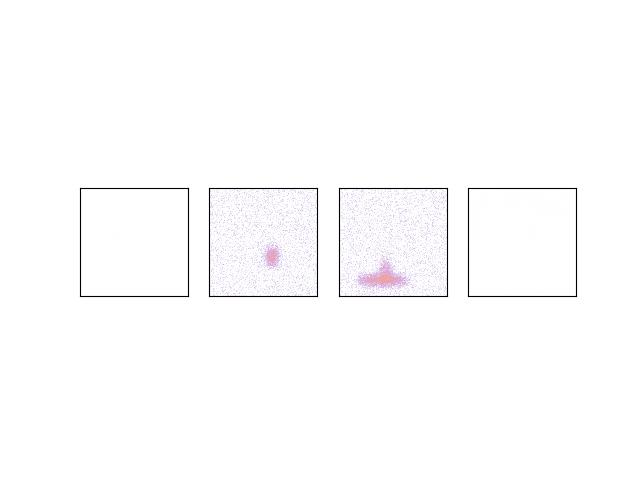
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> self = kwimage.Heatmap.random(dims=(50, 200), dets='coco', >>> keypoints=True) >>> image = np.zeros(self.img_dims) >>> # xdoctest: +REQUIRES(module:kwplot) >>> toshow = self.draw_on(image, 1, vecs=True, kpts=0, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(toshow)
- property class_probs¶
- property offset¶
- property diameter¶
- property img_dims¶
- property tf_data_to_img¶
- property classes¶
- class kwimage.structs.Mask(data=None, format=None)[source]¶
Bases:
NiceRepr,_MaskConversionMixin,_MaskConstructorMixin,_MaskTransformMixin,_MaskDrawMixinManages a single segmentation mask and can convert to and from multiple formats including:
bytes_rle - byte encoded run length encoding
array_rle - raw run length encoding
c_mask - c-style binary mask
f_mask - fortran-style binary mask
Example
>>> # xdoc: +REQUIRES(--mask) >>> # a ms-coco style compressed bytes rle segmentation >>> segmentation = {'size': [5, 9], 'counts': ';?1B10O30O4'} >>> mask = Mask(segmentation, 'bytes_rle') >>> # convert to binary numpy representation >>> binary_mask = mask.to_c_mask().data >>> print(ub.repr2(binary_mask.tolist(), nl=1, nobr=1)) [0, 0, 0, 1, 1, 1, 1, 1, 0], [0, 0, 1, 1, 1, 0, 0, 0, 0], [0, 0, 1, 1, 1, 1, 1, 1, 0], [0, 0, 1, 1, 1, 0, 1, 1, 0], [0, 0, 1, 1, 1, 0, 1, 1, 0],
- property dtype¶
- classmethod random(rng=None, shape=(32, 32))[source]¶
Create a random binary mask object
- Parameters
rng (int | RandomState | None) – the random seed
shape (Tuple[int, int]) – the height / width of the returned mask
- Returns
the random mask
- Return type
Example
>>> import kwimage >>> mask = kwimage.Mask.random() >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> mask.draw() >>> kwplot.show_if_requested()
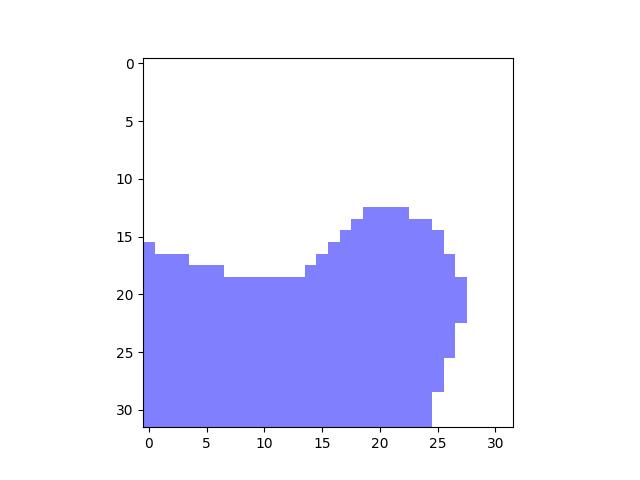
- classmethod demo()[source]¶
Demo mask with holes and disjoint shapes
- Returns
the demo mask
- Return type
- classmethod from_text(text, zero_chr='.', shape=None, has_border=False)[source]¶
Construct a mask from a text art representation
- Parameters
text (str) – the text representing a mask
zero_chr (str) – the character that represents a zero
shape (None | Tuple[int, int]) – if specified force a specific height / width, otherwise the character extent determines this.
has_border (bool) – if True, assume the characters at the edge are representing a border and remove them.
Example
>>> import kwimage >>> import ubelt as ub >>> text = ub.indent(ub.codeblock( >>> ''' >>> ooo >>> ooo >>> ooooo >>> o >>> ''')) >>> mask = kwimage.Mask.from_text(text, zero_chr=' ') >>> print(mask.data) [[0 0 0 0 1 1 1 0 0] [0 0 0 0 1 1 1 0 0] [0 0 0 0 1 1 1 1 1] [0 0 0 0 0 0 0 0 1]]
Example
>>> import kwimage >>> import ubelt as ub >>> text = ub.codeblock( >>> ''' >>> +------------+ >>> | | >>> | ooo | >>> | ooo | >>> | ooooo | >>> | o | >>> | | >>> +------------+ >>> ''') >>> mask = kwimage.Mask.from_text(text, has_border=True, zero_chr=' ') >>> print(mask.data) [[0 0 0 0 0 0 0 0 0 0 0 0] [0 0 0 0 1 1 1 0 0 0 0 0] [0 0 0 0 1 1 1 0 0 0 0 0] [0 0 0 0 1 1 1 1 1 0 0 0] [0 0 0 0 0 0 0 0 1 0 0 0] [0 0 0 0 0 0 0 0 0 0 0 0]]
- copy()[source]¶
Performs a deep copy of the mask data
- Returns
the copied mask
- Return type
Example
>>> self = Mask.random(shape=(8, 8), rng=0) >>> other = self.copy() >>> assert other.data is not self.data
- union(*others)[source]¶
This can be used as a staticmethod or an instancemethod
- Parameters
*others – multiple input masks to union
- Returns
the unioned mask
- Return type
Example
>>> # xdoc: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> masks = [Mask.random(shape=(8, 8), rng=i) for i in range(2)] >>> mask = Mask.union(*masks) >>> print(mask.area) >>> masks = [m.to_c_mask() for m in masks] >>> mask = Mask.union(*masks) >>> print(mask.area)
>>> masks = [m.to_bytes_rle() for m in masks] >>> mask = Mask.union(*masks) >>> print(mask.area)
- intersection(*others)[source]¶
This can be used as a staticmethod or an instancemethod
- Parameters
*others – multiple input masks to intersect
- Returns
the intersection of the masks
- Return type
Example
>>> n = 3 >>> masks = [Mask.random(shape=(8, 8), rng=i) for i in range(n)] >>> items = masks >>> mask = Mask.intersection(*masks) >>> areas = [item.area for item in items] >>> print('areas = {!r}'.format(areas)) >>> print(mask.area) >>> print(Mask.intersection(*masks).area / Mask.union(*masks).area)
- property shape¶
- property area¶
Returns the number of non-zero pixels
- Returns
the number of non-zero pixels
- Return type
Example
>>> self = Mask.demo() >>> self.area 150
- get_patch()[source]¶
Extract the patch with non-zero data
Example
>>> # xdoc: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.random(shape=(8, 8), rng=0) >>> self.get_patch()
- get_xywh()[source]¶
Gets the bounding xywh box coordinates of this mask
- Returns
- x, y, w, h: Note we dont use a Boxes object because
a general singular version does not yet exist.
- Return type
ndarray
Example
>>> # xdoc: +REQUIRES(--mask) >>> self = Mask.random(shape=(8, 8), rng=0) >>> self.get_xywh().tolist() >>> self = Mask.random(rng=0).translate((10, 10)) >>> self.get_xywh().tolist()
Example
>>> # test empty case >>> import kwimage >>> self = kwimage.Mask(np.empty((0, 0), dtype=np.uint8), format='c_mask') >>> assert self.get_xywh().tolist() == [0, 0, 0, 0]
- get_polygon()[source]¶
DEPRECATED: USE to_multi_polygon
Returns a list of (x,y)-coordinate lists. The length of the list is equal to the number of disjoint regions in the mask.
- Returns
- polygon around each connected component of the
mask. Each ndarray is an Nx2 array of xy points.
- Return type
List[ndarray]
Note
The returned polygon may not surround points that are only one pixel thick.
Example
>>> # xdoc: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.random(shape=(8, 8), rng=0) >>> polygons = self.get_polygon() >>> print('polygons = ' + ub.repr2(polygons)) >>> polygons = self.get_polygon() >>> self = self.to_bytes_rle() >>> other = Mask.from_polygons(polygons, self.shape) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> image = np.ones(self.shape) >>> image = self.draw_on(image, color='blue') >>> image = other.draw_on(image, color='red') >>> kwplot.imshow(image)
- to_mask(dims=None)[source]¶
Converts to a mask object (which does nothing because this already is mask object!)
- Returns
kwimage.Mask
- to_multi_polygon(pixels_are='points')[source]¶
Returns a MultiPolygon object fit around this raster including disjoint pieces and holes.
- Parameters
pixel_are (str) – Can either be “points” or “areas”.
If pixels are “points”, the we treat each pixel (i, j) as a single infinitely small point at (i, j). As such, some polygons may have zero area.
If pixels are “areas”, then each pixel (i, j) represents a square with coordinates ([i - 0.5, j - 0.5], [i + 0.5, j - 0.5], [i + 0.5, j + 0.5], and [i - 0.5, j + 0.5]). Must have rasterio installed to use this method.
- Returns
vectorized representation
- Return type
Note
The OpenCV (and thus this function) coordinate system places coordinates at the center of pixels, and the polygon is traced tightly around these coordinates. A single pixel is not considered to have any width, so polygon edges will directly trace through the centers of pixels, and in the case where an object is only 1 pixel thick, this will produce a polygon that is not a valid shapely polygon.
Todo
[x] add a flag where polygons consider pixels to have width and the resulting polygon is traced around the pixel edges, not the pixel centers.
[ ] Polygons and Masks should keep track of what “pixels_are”
Example
>>> # xdoc: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.demo() >>> self = self.scale(5) >>> multi_poly = self.to_multi_polygon() >>> # xdoc: +REQUIRES(module:kwplot) >>> # xdoc: +REQUIRES(--show) >>> self.draw(color='red') >>> multi_poly.scale(1.1).draw(color='blue')
>>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> image = np.ones(self.shape) >>> image = self.draw_on(image, color='blue') >>> #image = other.draw_on(image, color='red') >>> kwplot.imshow(image) >>> multi_poly.draw()
Example
>>> # Test empty cases >>> import kwimage >>> mask0 = kwimage.Mask(np.zeros((0, 0), dtype=np.uint8), format='c_mask') >>> mask1 = kwimage.Mask(np.zeros((1, 1), dtype=np.uint8), format='c_mask') >>> mask2 = kwimage.Mask(np.zeros((2, 2), dtype=np.uint8), format='c_mask') >>> mask3 = kwimage.Mask(np.zeros((3, 3), dtype=np.uint8), format='c_mask') >>> pixels_are = 'points' >>> poly0 = mask0.to_multi_polygon(pixels_are=pixels_are) >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert len(poly0) == 0 >>> assert len(poly1) == 0 >>> assert len(poly2) == 0 >>> assert len(poly3) == 0 >>> # xdoctest: +REQUIRES(module:rasterio) >>> pixels_are = 'areas' >>> poly0 = mask0.to_multi_polygon(pixels_are=pixels_are) >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert len(poly0) == 0 >>> assert len(poly1) == 0 >>> assert len(poly2) == 0 >>> assert len(poly3) == 0
Example
>>> # Test full ones cases >>> import kwimage >>> mask1 = kwimage.Mask(np.ones((1, 1), dtype=np.uint8), format='c_mask') >>> mask2 = kwimage.Mask(np.ones((2, 2), dtype=np.uint8), format='c_mask') >>> mask3 = kwimage.Mask(np.ones((3, 3), dtype=np.uint8), format='c_mask') >>> pixels_are = 'points' >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert np.all(poly1.to_mask(mask1.shape).data == 1) >>> assert np.all(poly2.to_mask(mask2.shape).data == 1) >>> assert np.all(poly3.to_mask(mask3.shape).data == 1) >>> # xdoctest: +REQUIRES(module:rasterio) >>> pixels_are = 'areas' >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert np.all(poly1.to_mask(mask1.shape).data == 1) >>> assert np.all(poly2.to_mask(mask2.shape).data == 1) >>> assert np.all(poly3.to_mask(mask3.shape).data == 1)
Example
>>> # Corner case, only two pixels are on >>> import kwimage >>> self = kwimage.Mask(np.zeros((768, 768), dtype=np.uint8), format='c_mask') >>> x_coords = np.array([621, 752]) >>> y_coords = np.array([366, 292]) >>> self.data[y_coords, x_coords] = 1 >>> poly = self.to_multi_polygon()
Example
>>> # xdoctest: +REQUIRES(module:rasterio) >>> import kwimage >>> dims = (10, 10) >>> data = np.zeros(dims, dtype=np.uint8) >>> data[0, 3:5] = 1 >>> data[9, 1:3] = 1 >>> data[3:5, 0:2] = 1 >>> data[1, 1] = 1 >>> # 1 pixel L shape >>> data[3, 5] = 1 >>> data[4, 5] = 1 >>> data[4, 6] = 1 >>> data[1, 5] = 1 >>> data[2, 6] = 1 >>> data[3, 7] = 1 >>> data[6, 1] = 1 >>> data[7, 1] = 1 >>> data[7, 2] = 1 >>> data[6:10, 5] = 1 >>> data[6:10, 8] = 1 >>> data[9, 5:9] = 1 >>> data[6, 5:9] = 1 >>> #data = kwimage.imresize(data, scale=2.0, interpolation='nearest') >>> self = kwimage.Mask.coerce(data) >>> #self = self.translate((0, 0), output_dims=(10, 9)) >>> self = self.translate((0, 1), output_dims=(11, 11)) >>> dims = self.shape[0:2] >>> multi_poly1 = self.to_multi_polygon(pixels_are='points') >>> multi_poly2 = self.to_multi_polygon(pixels_are='areas') >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pretty_data = kwplot.make_heatmask(self.data/1.0, cmap='magma')[..., 0:3] >>> def _pixel_grid_lines(self, ax): >>> h, w = self.data.shape[0:2] >>> ybasis = np.arange(0, h) + 0.5 >>> xbasis = np.arange(0, w) + 0.5 >>> xmin = 0 - 0.5 >>> xmax = w - 0.5 >>> ymin = 0 - 0.5 >>> ymax = h - 0.5 >>> ax.hlines(y=ybasis, xmin=xmin, xmax=xmax, color="gainsboro") >>> ax.vlines(x=xbasis, ymin=ymin, ymax=ymax, color="gainsboro") >>> def _setup_grid(self, pnum): >>> ax = kwplot.imshow(pretty_data, show_ticks=True, pnum=pnum)[1] >>> # The gray ticks show the center of the pixels >>> ax.grid(color='dimgray', linewidth=0.5) >>> ax.set_xticks(np.arange(self.data.shape[1])) >>> ax.set_yticks(np.arange(self.data.shape[0])) >>> # Also draw black lines around the edges of the pixels >>> _pixel_grid_lines(self, ax=ax) >>> return ax >>> # Overlay the extracted polygons >>> ax = _setup_grid(self, pnum=(2, 3, 1)) >>> ax.set_title('input binary mask data') >>> ax = _setup_grid(self, pnum=(2, 3, 2)) >>> multi_poly1.draw(linewidth=5, alpha=0.5, radius=0.2, ax=ax, fill=False, vertex=0.2) >>> ax.set_title('opencv "point" polygons') >>> ax = _setup_grid(self, pnum=(2, 3, 3)) >>> multi_poly2.draw(linewidth=5, alpha=0.5, radius=0.2, color='limegreen', ax=ax, fill=False, vertex=0.2) >>> ax.set_title('raterio "area" polygons') >>> ax.figure.suptitle(ub.codeblock( >>> ''' >>> Gray lines are coordinates and pass through pixel centers (integer coords) >>> White lines trace pixel boundaries (fractional coords) >>> ''')) >>> raster1 = multi_poly1.to_mask(dims, pixels_are='points') >>> raster2 = multi_poly2.to_mask(dims, pixels_are='areas') >>> kwplot.imshow(raster1.draw_on(), pnum=(2, 3, 5), title='rasterized') >>> kwplot.imshow(raster2.draw_on(), pnum=(2, 3, 6), title='rasterized')
- get_convex_hull()[source]¶
Returns a list of xy points around the convex hull of this mask
Note
The returned polygon may not surround points that are only one pixel thick.
Example
>>> # xdoc: +REQUIRES(--mask) >>> self = Mask.random(shape=(8, 8), rng=0) >>> polygons = self.get_convex_hull() >>> print('polygons = ' + ub.repr2(polygons)) >>> other = Mask.from_polygons(polygons, self.shape)
- iou(other)[source]¶
The area of intersection over the area of union
Todo
[ ] Write plural Masks version of this class, which should be able to perform this operation more efficiently.
CommandLine
xdoctest -m kwimage.structs.mask Mask.iou
Example
>>> # xdoc: +REQUIRES(--mask) >>> self = Mask.demo() >>> other = self.translate(1) >>> iou = self.iou(other) >>> print('iou = {:.4f}'.format(iou)) iou = 0.0830 >>> iou2 = self.intersection(other).area / self.union(other).area >>> print('iou2 = {:.4f}'.format(iou2))
- classmethod coerce(data, dims=None)[source]¶
Attempts to auto-inspect the format of the data and conver to Mask
- Parameters
data (Any) – the data to coerce
dims (Tuple) – required for certain formats like polygons height / width of the source image
- Returns
the constructed mask object
- Return type
Example
>>> # xdoc: +REQUIRES(--mask) >>> segmentation = {'size': [5, 9], 'counts': ';?1B10O30O4'} >>> polygon = [ >>> [np.array([[3, 0],[2, 1],[2, 4],[4, 4],[4, 3],[7, 0]])], >>> [np.array([[2, 1],[2, 2],[4, 2],[4, 1]])], >>> ] >>> dims = (9, 5) >>> mask = (np.random.rand(32, 32) > .5).astype(np.uint8) >>> Mask.coerce(polygon, dims).to_bytes_rle() >>> Mask.coerce(segmentation).to_bytes_rle() >>> Mask.coerce(mask).to_bytes_rle()
- to_coco(style='orig')[source]¶
Convert the Mask to a COCO json representation based on the current format.
A COCO mask is formatted as a run-length-encoding (RLE), of which there are two variants: (1) a array RLE, which is slightly more readable and extensible, and (2) a bytes RLE, which is slightly more concise. The returned format will depend on the current format of the Mask object. If it is in “bytes_rle” format, it will be returned in that format, otherwise it will be converted to the “array_rle” format and returned as such.
- Parameters
style (str) – Does nothing for this particular method, exists for API compatibility and if alternate encoding styles are implemented in the future.
- Returns
- either a bytes-rle or array-rle encoding, depending
on the current mask format. The keys in this dictionary are as follows:
counts (List[int] | str): the array or bytes rle encoding
- size (Tuple[int]): the height and width of the encoded mask
see note.
- shape (Tuple[int]): only present in array-rle mode. This
is also the height/width of the underlying encoded array. This exists for semantic consistency with other kwimage conventions, and is not part of the original coco spec.
- order (str): only present in array-rle mode.
Either C or F, indicating if counts is aranged in row-major or column-major order. For COCO-compatibility this is always returned in F (column-major) order.
- binary (bool): only present in array-rle mode.
For COCO-compatibility this is always returned as False, indicating the mask only contains binary 0 or 1 values.
- Return type
Note
The output dictionary will contain a key named “size”, this is the only location in kwimage where “size” refers to a tuple in (height/width) order, in order to be backwards compatible with the original coco spec. In all other locations in kwimage a “size” will refer to a (width/height) ordered tuple.
- SeeAlso:
- func
kwimage.im_runlen.encode_run_length - backend function that does array-style run length encoding.
Example
>>> # xdoc: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.demo() >>> coco_data1 = self.toformat('array_rle').to_coco() >>> coco_data2 = self.toformat('bytes_rle').to_coco() >>> print('coco_data1 = {}'.format(ub.repr2(coco_data1, nl=1))) >>> print('coco_data2 = {}'.format(ub.repr2(coco_data2, nl=1))) coco_data1 = { 'binary': True, 'counts': [47, 5, 3, 1, 14, ... 1, 4, 19, 141], 'order': 'F', 'shape': (23, 32), 'size': (23, 32), } coco_data2 = { 'counts': '_153L;4EL...ON3060L0N060L0Nb0Y4', 'size': [23, 32], }
- class kwimage.structs.MaskList(data, meta=None)[source]¶
Bases:
ObjectListStore and manipulate multiple masks, usually within the same image
- to_polygon_list()[source]¶
Converts all mask objects to multi-polygon objects
- Returns
kwimage.PolygonList
- class kwimage.structs.MultiPolygon(data, meta=None)[source]¶
Bases:
ObjectListData structure for storing multiple polygons (typically related to the same underlying but potentitally disjoing object)
- Variables
data (List[Polygon]) –
- property area¶
Computes are via shapley conversion
- Returns
float
- classmethod random(n=3, n_holes=0, rng=None, tight=False)[source]¶
Create a random MultiPolygon
- Returns
MultiPolygon
- fill(image, value=1, pixels_are='points')[source]¶
Inplace fill in an image based on this multi-polyon.
- Parameters
image (ndarray) – image to draw on (inplace)
value (int | Tuple[int, …]) – value fill in with. Defaults to 1.0
- Returns
the image that has been modified in place
- Return type
ndarray
- bounding_box()[source]¶
Return the bounding box of the multi polygon
- Returns
- a Boxes object with one box that encloses all
polygons
- Return type
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(rng=0, n=10) >>> boxes = self.to_boxes() >>> sub_boxes = [d.to_boxes() for d in self.data] >>> areas1 = np.array([s.intersection(boxes).area[0] for s in sub_boxes]) >>> areas2 = np.array([s.area[0] for s in sub_boxes]) >>> assert np.allclose(areas1, areas2)
- to_mask(dims=None, pixels_are='points')[source]¶
Returns a mask object indication regions occupied by this multipolygon
- Returns
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> s = 100 >>> self = MultiPolygon.random(rng=0).scale(s) >>> dims = (s, s) >>> mask = self.to_mask(dims) >>> # xdoc: +REQUIRES(--show) >>> # xdoc: +REQUIRES(module:kwplot) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> ax = plt.gca() >>> ax.set_xlim(0, s) >>> ax.set_ylim(0, s) >>> self.draw(color='red', alpha=.4) >>> mask.draw(color='blue', alpha=.4)
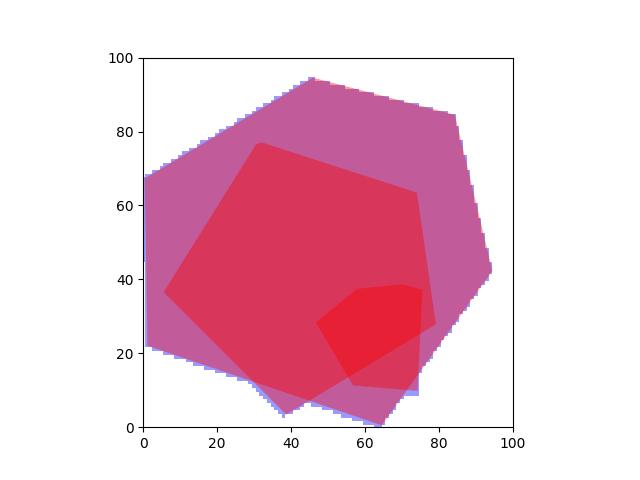
- to_relative_mask(return_offset=False)[source]¶
Returns a translated mask such the mask dimensions are minimal.
In other words, we move the polygon all the way to the top-left and return a mask just big enough to fit the polygon.
- Returns
kwimage.Mask
- classmethod coerce(data, dims=None)[source]¶
Attempts to construct a MultiPolygon instance from the input data
See Segmentation.coerce
- Returns
None | MultiPolygon
Example
>>> import kwimage >>> dims = (32, 32) >>> kw_poly = kwimage.Polygon.random().scale(dims) >>> kw_multi_poly = kwimage.MultiPolygon.random().scale(dims) >>> forms = [kw_poly, kw_multi_poly] >>> forms.append(kw_poly.to_shapely()) >>> forms.append(kw_poly.to_mask((32, 32))) >>> forms.append(kw_poly.to_geojson()) >>> forms.append(kw_poly.to_coco(style='orig')) >>> forms.append(kw_poly.to_coco(style='new')) >>> forms.append(kw_multi_poly.to_shapely()) >>> forms.append(kw_multi_poly.to_mask((32, 32))) >>> forms.append(kw_multi_poly.to_geojson()) >>> forms.append(kw_multi_poly.to_coco(style='orig')) >>> forms.append(kw_multi_poly.to_coco(style='new')) >>> for data in forms: >>> result = kwimage.MultiPolygon.coerce(data, dims=dims) >>> assert isinstance(result, kwimage.MultiPolygon)
- to_shapely()[source]¶
- Returns
shapely.geometry.MultiPolygon
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> # xdoc: +REQUIRES(module:shapely) >>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(rng=0) >>> geom = self.to_shapely() >>> print('geom = {!r}'.format(geom))
- classmethod from_shapely(geom)[source]¶
Convert a shapely polygon or multipolygon to a kwimage.MultiPolygon
- Parameters
geom (shapely.geometry.MultiPolygon | shapely.geometry.Polygon)
- Returns
MultiPolygon
Example
>>> import kwimage >>> sh_poly = kwimage.Polygon.random().to_shapely() >>> sh_multi_poly = kwimage.MultiPolygon.random().to_shapely() >>> kwimage.MultiPolygon.from_shapely(sh_poly) >>> kwimage.MultiPolygon.from_shapely(sh_multi_poly)
- classmethod from_geojson(data_geojson)[source]¶
Convert a geojson polygon or multipolygon to a kwimage.MultiPolygon
- Parameters
data_geojson (Dict) – geojson data
- Returns
MultiPolygon
Example
>>> import kwimage >>> orig = kwimage.MultiPolygon.random() >>> data_geojson = orig.to_geojson() >>> self = kwimage.MultiPolygon.from_geojson(data_geojson)
- classmethod from_coco(data, dims=None)[source]¶
Accepts either new-style or old-style coco multi-polygons
- Parameters
data (List[List[Number] | Dict]) – a new or old style coco multi polygon
dims (None | Tuple[int, …]) – the shape dimensions of the canvas. Unused. Exists for compatibility with masks.
- Returns
MultiPolygon
- class kwimage.structs.Points(data=None, meta=None, datakeys=None, metakeys=None, **kwargs)[source]¶
Bases:
Spatial,_PointsWarpMixinStores multiple keypoints for a single object.
This stores both the geometry and the class metadata if available
Example
>>> from kwimage.structs.points import * # NOQA >>> xy = np.random.rand(10, 2) >>> pts = Points(xy=xy) >>> print('pts = {!r}'.format(pts))
- property shape¶
- property xy¶
- classmethod random(num=1, classes=None, rng=None)[source]¶
Makes random points; typically for testing purposes
Example
>>> import kwimage >>> self = kwimage.Points.random(classes=[1, 2, 3]) >>> self.data >>> print('self.data = {!r}'.format(self.data))
- tensor(device=NoParam)[source]¶
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10) >>> self.tensor()
- round(inplace=False)[source]¶
Rounds data to the nearest integer
- Parameters
inplace (bool) – if True, modifies this object
Example
>>> import kwimage >>> self = kwimage.Points.random(3).scale(10) >>> self.round()
- numpy()[source]¶
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10) >>> self.tensor().numpy().tensor().numpy()
- draw_on(image=None, color='white', radius=None, copy=False)[source]¶
- Parameters
image (ndarray) – image to draw points on.
color (str | Any | List[Any]) – one color for all boxes or a list of colors for each box Can be any type accepted by kwimage.Color.coerce. Extended types: str | ColorLike | List[ColorLike]
radius (None | int) – if an integer, an circle is drawn at each xy point with this radius. if None, attempts to fill a single point with subpixel accuracy, which generally means 4 pixels will be given some weight. Note: color can only be a single value for all points in this case.
copy (bool) – if True, force a copy of the image, otherwise try to draw inplace (may not work depending on dtype).
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/points.py Points.draw_on --show
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> s = 128 >>> image = np.zeros((s, s)) >>> self = Points.random(10).scale(s) >>> image = self.draw_on(image) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.autompl() >>> kwplot.imshow(image) >>> self.draw(radius=3, alpha=.5) >>> kwplot.show_if_requested()

Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> s = 128 >>> image = np.zeros((s, s)) >>> self = Points.random(10).scale(s) >>> image = self.draw_on(image, radius=3, color='distinct') >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.autompl() >>> kwplot.imshow(image) >>> #self.draw(radius=3, alpha=.5, color='classes') >>> kwplot.show_if_requested()

Example
>>> import kwimage >>> s = 32 >>> self = kwimage.Points.random(10).scale(s) >>> color = 'kitware_green' >>> # Test drawing on all channel + dtype combinations >>> im3 = np.zeros((s, s, 3), dtype=np.float32) >>> im_chans = { >>> 'im3': im3, >>> 'im1': kwimage.convert_colorspace(im3, 'rgb', 'gray'), >>> 'im4': kwimage.convert_colorspace(im3, 'rgb', 'rgba'), >>> } >>> inputs = {} >>> for k, im in im_chans.items(): >>> inputs[k + '_01'] = (kwimage.ensure_float01(im.copy()), {'radius': None}) >>> inputs[k + '_255'] = (kwimage.ensure_uint255(im.copy()), {'radius': None}) >>> outputs = {} >>> for k, v in inputs.items(): >>> im, kw = v >>> outputs[k] = self.draw_on(im, color=color, **kw) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=2, doclf=True) >>> plt = kwplot.autoplt() >>> pnum_ = kwplot.PlotNums(nRows=2, nSubplots=len(inputs)) >>> for k in inputs.keys(): >>> kwplot.imshow(outputs[k], fnum=2, pnum=pnum_(), title=k) >>> plt.gcf().suptitle('Test draw points on channel + dtype combos') >>> kwplot.show_if_requested()

Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10).scale(32) >>> image = self.draw_on(radius=3, color='distinct') >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image) >>> kwplot.show_if_requested()

Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> # Test cases where single and multiple colors are given >>> # with radius=None and radius=scalar >>> from kwimage.structs.points import * # NOQA >>> import kwimage >>> self = kwimage.Points.random(10).scale(32) >>> image1 = self.draw_on(radius=2, color='blue') >>> image2 = self.draw_on(radius=None, color='blue') >>> image3 = self.draw_on(radius=2, color='distinct') >>> image4 = self.draw_on(radius=None, color='distinct') >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> canvas = kwimage.stack_images_grid( >>> [image1, image2, image3, image4], >>> pad=3, bg_value=(1, 1, 1)) >>> kwplot.autompl() >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()

- draw(color='blue', ax=None, alpha=None, radius=1, **kwargs)[source]¶
TODO: can use kwplot.draw_points
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> pts = Points.random(10) >>> # xdoc: +REQUIRES(--show) >>> pts.draw(radius=0.01)
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10, classes=['a', 'b', 'c']) >>> self.draw(radius=0.01, color='classes')
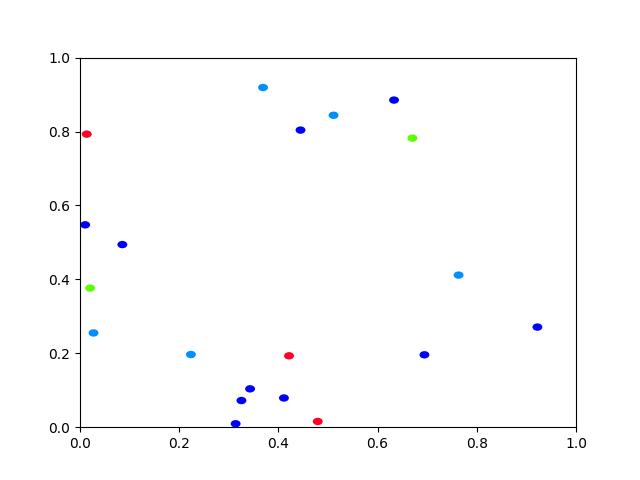
- compress(flags, axis=0, inplace=False)[source]¶
Filters items based on a boolean criterion
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4) >>> flags = [1, 0, 1, 1] >>> other = self.compress(flags) >>> assert len(self) == 4 >>> assert len(other) == 3
>>> # xdoctest: +REQUIRES(module:torch) >>> other = self.tensor().compress(flags) >>> assert len(other) == 3
- take(indices, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4) >>> indices = [1, 3] >>> other = self.take(indices) >>> assert len(self) == 4 >>> assert len(other) == 2
>>> # xdoctest: +REQUIRES(module:torch) >>> other = self.tensor().take(indices) >>> assert len(other) == 2
- to_coco(style='orig')[source]¶
Converts to an mscoco-like representation
Note
items that are usually id-references to other objects may need to be rectified.
- Parameters
style (str) – either orig, new, new-id, or new-name
- Returns
mscoco-like representation
- Return type
Dict
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4, classes=['a', 'b']) >>> orig = self._to_coco(style='orig') >>> print('orig = {!r}'.format(orig)) >>> new_name = self._to_coco(style='new-name') >>> print('new_name = {}'.format(ub.repr2(new_name, nl=-1))) >>> # xdoctest: +REQUIRES(module:kwcoco) >>> import kwcoco >>> self.meta['classes'] = kwcoco.CategoryTree.coerce(self.meta['classes']) >>> new_id = self._to_coco(style='new-id') >>> print('new_id = {}'.format(ub.repr2(new_id, nl=-1)))
- classmethod from_coco(coco_kpts, class_idxs=None, classes=None, warn=False)[source]¶
- Parameters
coco_kpts (list | dict) – either the original list keypoint encoding or the new dict keypoint encoding.
class_idxs (list) – only needed if using old style
classes (list | kwcoco.CategoryTree) – list of all keypoint category names
warn (bool) – if True raise warnings
Example
>>> ## >>> classes = ['mouth', 'left-hand', 'right-hand'] >>> coco_kpts = [ >>> {'xy': (0, 0), 'visible': 2, 'keypoint_category': 'left-hand'}, >>> {'xy': (1, 2), 'visible': 2, 'keypoint_category': 'mouth'}, >>> ] >>> Points.from_coco(coco_kpts, classes=classes) >>> # Test without classes >>> Points.from_coco(coco_kpts) >>> # Test without any category info >>> coco_kpts2 = [ub.dict_diff(d, {'keypoint_category'}) for d in coco_kpts] >>> Points.from_coco(coco_kpts2) >>> # Test without category instead of keypoint_category >>> coco_kpts3 = [ub.map_keys(lambda x: x.replace('keypoint_', ''), d) for d in coco_kpts] >>> Points.from_coco(coco_kpts3) >>> # >>> # Old style >>> coco_kpts = [0, 0, 2, 0, 1, 2] >>> Points.from_coco(coco_kpts) >>> # Fail case >>> coco_kpts4 = [{'xy': [4686.5, 1341.5], 'category': 'dot'}] >>> Points.from_coco(coco_kpts4, classes=[])
Example
>>> # xdoctest: +REQUIRES(module:kwcoco) >>> import kwcoco >>> classes = kwcoco.CategoryTree.from_coco([ >>> {'name': 'mouth', 'id': 2}, {'name': 'left-hand', 'id': 3}, {'name': 'right-hand', 'id': 5} >>> ]) >>> coco_kpts = [ >>> {'xy': (0, 0), 'visible': 2, 'keypoint_category_id': 5}, >>> {'xy': (1, 2), 'visible': 2, 'keypoint_category_id': 2}, >>> ] >>> pts = Points.from_coco(coco_kpts, classes=classes) >>> assert pts.data['class_idxs'].tolist() == [2, 0]
- class kwimage.structs.PointsList(data, meta=None)[source]¶
Bases:
ObjectListStores a list of Points, each item usually corresponds to a different object.
Note
# TODO: when the data is homogenous we can use a more efficient # representation, otherwise we have to use heterogenous storage.
- class kwimage.structs.Polygon(data=None, meta=None, datakeys=None, metakeys=None, **kwargs)[source]¶
Bases:
Spatial,_PolyArrayBackend,_PolyWarpMixin,NiceReprRepresents a single polygon as set of exterior boundary points and a list of internal polygons representing holes.
By convention exterior boundaries should be counterclockwise and interior holes should be clockwise.
Example
>>> import kwimage >>> data = { >>> 'exterior': np.array([[13, 1], [13, 19], [25, 19], [25, 1]]), >>> 'interiors': [ >>> np.array([[13, 13], [14, 12], [24, 12], [25, 13], [25, 18], >>> [24, 19], [14, 19], [13, 18]]), >>> np.array([[13, 2], [14, 1], [24, 1], [25, 2], [25, 11], >>> [24, 12], [14, 12], [13, 11]])] >>> } >>> self = kwimage.Polygon(**data) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=True)
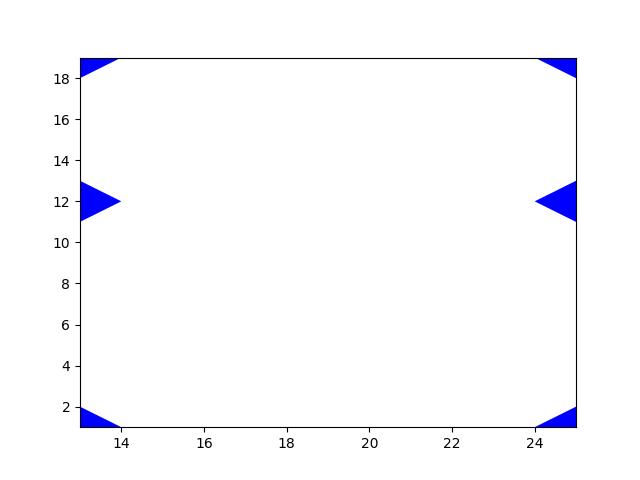
Example
>>> import kwimage >>> self = kwimage.Polygon.random( >>> n=5, n_holes=1, convex=False, rng=0) >>> print('self = {}'.format(self)) self = <Polygon({ 'exterior': <Coords(data= array([[0.30371392, 0.97195856], [0.24372304, 0.60568445], [0.21408694, 0.34884262], [0.5799477 , 0.44020379], [0.83720288, 0.78367234]]))>, 'interiors': [<Coords(data= array([[0.50164209, 0.83520279], [0.25835064, 0.40313428], [0.28778562, 0.74758761], [0.30341266, 0.93748088]]))>], })> >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=True)
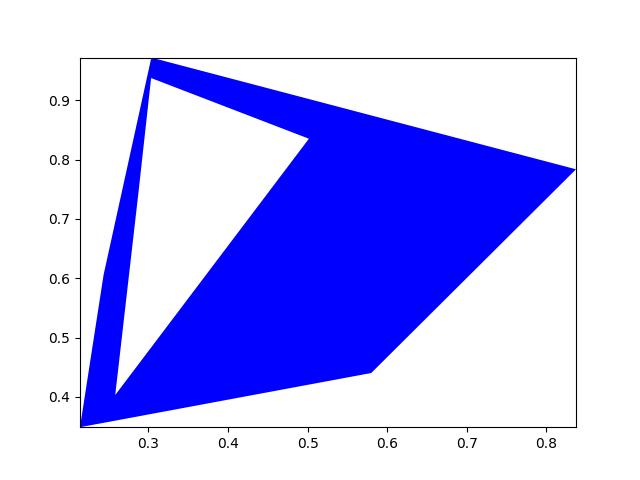
Example
>>> # Test empty polygon >>> import kwimage >>> data = { >>> 'exterior': np.array([]), >>> 'interiors': [],} >>> self = kwimage.Polygon(**data) >>> geos = self.to_geojson() >>> kwimage.Polygon.from_geojson(geos) >>> geom = self.to_shapely() >>> kwimage.Polygon.from_shapely(geom) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=True)
- property exterior¶
Returns: kwimage.Coords
- property interiors¶
Returns: List[kwimage.Coords]
- classmethod circle(xy, r, resolution=64)[source]¶
Create a circular or elliptical polygon.
Might rename to ellipse later?
- Parameters
xy (Iterable[Number]) – x and y center coordinate
r (Number | Tuple[Number, Number]) – circular radius or major and minor elliptical radius
resolution (int) – number of sides
- Returns
Polygon
Example
>>> import kwimage >>> xy = (0.5, 0.5) >>> r = .3 >>> # Demo with circle >>> circle = kwimage.Polygon.circle(xy, r, resolution=6) >>> # Demo with ellipse >>> xy = (0.5, 0.5) >>> r = (.4, .7) >>> ellipse1 = kwimage.Polygon.circle(xy, r, resolution=12) >>> ellipse2 = kwimage.Polygon.circle(xy, (.7, .4), resolution=12) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> circle.draw(setlim=True, border=1, fill=0, color='kitware_orange') >>> ellipse1.draw(setlim=True, border=1, fill=0, color='kitware_blue') >>> ellipse2.draw(setlim=True, border=1, fill=0, color='kitware_green') >>> plt.gca().set_xlim(-0.5, 1.5) >>> plt.gca().set_ylim(-0.5, 1.5) >>> plt.gca().set_aspect('equal')
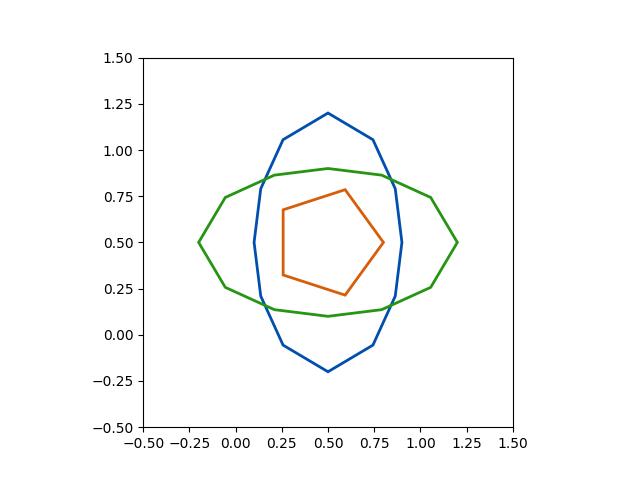
- classmethod random(n=6, n_holes=0, convex=True, tight=False, rng=None)[source]¶
- Parameters
n (int) – number of points in the polygon (must be 3 or more)
n_holes (int) – number of holes
tight (bool) – fits the minimum and maximum points between 0 and 1
convex (bool) – force resulting polygon will be convex (may remove exterior points)
- Returns
Polygon
CommandLine
xdoctest -m kwimage.structs.polygon Polygon.random
Example
>>> rng = None >>> n = 4 >>> n_holes = 1 >>> cls = Polygon >>> self = Polygon.random(n=n, rng=rng, n_holes=n_holes, convex=1) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> self.draw()
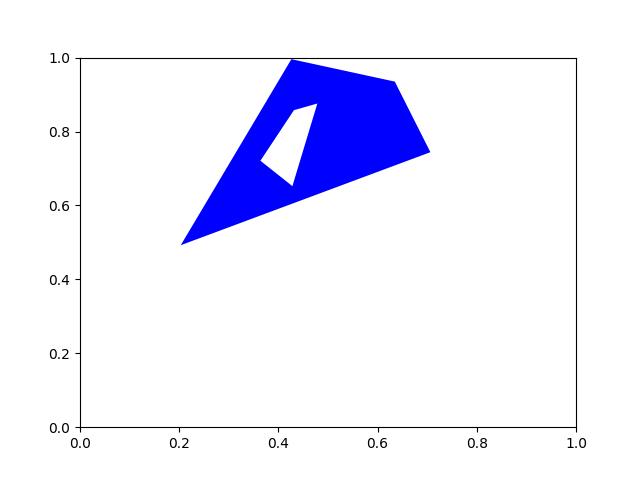
References
https://gis.stackexchange.com/questions/207731/random-multipolygon https://stackoverflow.com/questions/8997099/random-polygon https://stackoverflow.com/questions/27548363/from-voronoi-tessellation-to-shapely-polygons https://stackoverflow.com/questions/8997099/algorithm-to-generate-random-2d-polygon
- to_mask(dims=None, pixels_are='points')[source]¶
Convert this polygon to a mask
Todo
[ ] currently not efficient
- Parameters
dims (Tuple) – height and width of the output mask
pixels_are (str) – either “points” or “areas”
- Returns
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1).scale(128) >>> mask = self.to_mask((128, 128)) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> mask.draw(color='blue') >>> mask.to_multi_polygon().draw(color='red', alpha=.5)
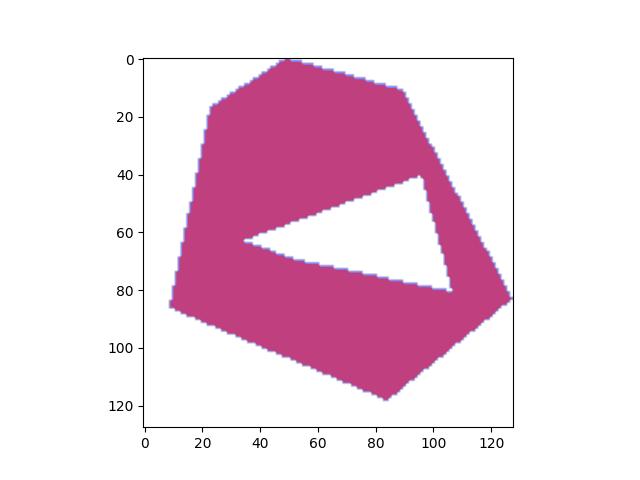
- to_relative_mask(return_offset=False)[source]¶
Returns a translated mask such the mask dimensions are minimal.
In other words, we move the polygon all the way to the top-left and return a mask just big enough to fit the polygon.
- Returns
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random().scale(8).translate(100, 100) >>> mask = self.to_relative_mask() >>> assert mask.shape <= (8, 8) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> mask.draw(color='blue') >>> mask.to_multi_polygon().draw(color='red', alpha=.5)
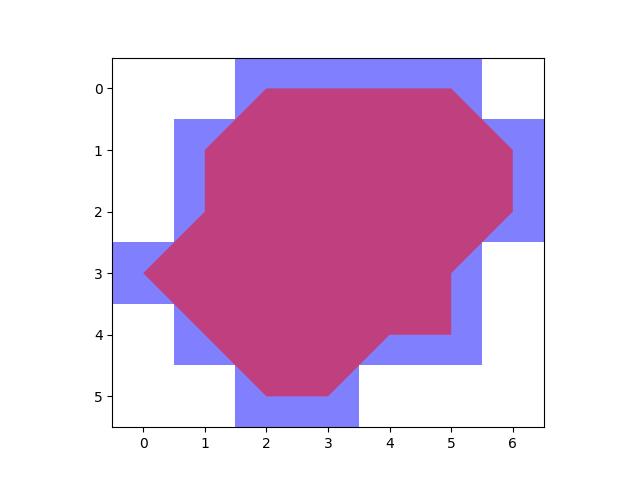
- classmethod coerce(data)[source]¶
Routes the input to the proper constructor
Try to autodetermine format of input polygon and coerce it into a kwimage.Polygon.
- Parameters
data (object) – some type of data that can be interpreted as a polygon.
- Returns
kwimage.Polygon
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> kwimage.Polygon.coerce(self) >>> kwimage.Polygon.coerce(self.exterior) >>> kwimage.Polygon.coerce(self.exterior.data) >>> kwimage.Polygon.coerce(self.data) >>> kwimage.Polygon.coerce(self.to_geojson()) >>> kwimage.Polygon.coerce('POLYGON ((0.11 0.61, 0.07 0.588, 0.015 0.50, 0.11 0.61))')
- classmethod from_shapely(geom)[source]¶
Convert a shapely polygon to a kwimage.Polygon
- Parameters
geom (shapely.geometry.polygon.Polygon) – a shapely polygon
- Returns
kwimage.Polygon
- classmethod from_wkt(data)[source]¶
Convert a WKT string to a kwimage.Polygon
- Parameters
data (str) – a WKT polygon string
- Returns
kwimage.Polygon
Example
>>> import kwimage >>> data = 'POLYGON ((0.11 0.61, 0.07 0.588, 0.015 0.50, 0.11 0.61))' >>> self = kwimage.Polygon.from_wkt(data) >>> assert len(self.exterior) == 4
- classmethod from_geojson(data_geojson)[source]¶
Convert a geojson polygon to a kwimage.Polygon
- Parameters
data_geojson (dict) – geojson data
- Returns
Polygon
References
https://geojson.org/geojson-spec.html
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=2) >>> data_geojson = self.to_geojson() >>> new = Polygon.from_geojson(data_geojson)
- to_shapely()[source]¶
- Returns
shapely.geometry.polygon.Polygon
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> # xdoc: +REQUIRES(module:shapely) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1) >>> self = self.scale(100) >>> geom = self.to_shapely() >>> print('geom = {!r}'.format(geom))
- property area¶
Computes are via shapley conversion
- Returns
float
- to_geojson()[source]¶
Converts polygon to a geojson structure
- Returns
Dict[str, object]
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> print(self.to_geojson())
- to_wkt()[source]¶
Convert a kwimage.Polygon to WKT string
- Returns
str
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> print(self.to_wkt())
- classmethod from_coco(data, dims=None)[source]¶
Accepts either new-style or old-style coco polygons
- Parameters
data (List[Number] | Dict) – A new or old-style coco polygon
dims (None | Tuple[int, …]) – the shape dimensions of the canvas. Unused. Exists for compatibility with masks.
- Returns
Polygon
- to_coco(style='orig')[source]¶
- Parameters
style (str) – can be “orig” or “new”
- Returns
coco-style polygons
- Return type
List | Dict
- property centroid¶
Returns: Tuple[Number, Number]
- bounding_box()[source]¶
Returns an axis-aligned bounding box for the segmentation
- Returns
kwimage.Boxes
- bounding_box_polygon()[source]¶
Returns an axis-aligned bounding polygon for the segmentation.
Note
This Polygon will be a Box, not a convex hull! Use shapely for convex hulls.
- Returns
kwimage.Polygon
- clip(x_min, y_min, x_max, y_max, inplace=False)[source]¶
Clip polygon to specified boundaries.
- Returns
clipped polygon
- Return type
Example
>>> from kwimage.structs.polygon import * >>> self = Polygon.random().scale(10).translate(-1) >>> self2 = self.clip(1, 1, 3, 3) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self2.draw(setlim=True)
- fill(image, value=1, pixels_are='points')[source]¶
Inplace fill in an image based on this polyon.
- Parameters
image (ndarray) – image to draw on
value (int | Tuple[int]) – value fill in with. Defaults to 1.
pixels_are (str) – either points or areas
- Returns
the image that has been modified in place
- Return type
ndarray
Example
>>> # xdoctest: +REQUIRES(module:rasterio) >>> import kwimage >>> mask = kwimage.Mask.random() >>> self = mask.to_multi_polygon(pixels_are='areas').data[0] >>> image = np.zeros_like(mask.data) >>> self.fill(image, pixels_are='areas')
Example
>>> # Test case where there are multiple channels >>> import kwimage >>> mask = kwimage.Mask.random(shape=(4, 4), rng=0) >>> self = mask.to_multi_polygon() >>> image = np.zeros(mask.shape[0:2] + (2,)) >>> fill_v1 = self.fill(image.copy(), value=1) >>> fill_v2 = self.fill(image.copy(), value=(1, 2)) >>> assert np.all((fill_v1 > 0) == (fill_v2 > 0))
- draw_on(image, color='blue', fill=True, border=False, alpha=1.0, edgecolor=None, facecolor=None, copy=False)[source]¶
Rasterizes a polygon on an image. See draw for a vectorized matplotlib version.
- Parameters
image (ndarray) – image to raster polygon on.
color (str | tuple) – data coercable to a color
fill (bool) – draw the center mass of the polygon. Note: this will be deprecated. Use facecolor instead.
border (bool) – draw the border of the polygon Note: this will be deprecated. Use edgecolor instead.
alpha (float) – polygon transparency (setting alpha < 1 makes this function much slower). Defaults to 1.0
copy (bool) – if False only copies if necessary
edgecolor (str | tuple) – color for the border
facecolor (str | tuple) – color for the fill
- Returns
np.ndarray
Note
This function will only be inplace if alpha=1.0 and the input has 3 or 4 channels. Otherwise the output canvas is coerced so colors can be drawn on it. In the case where alpha < 1.0,
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1).scale(128) >>> image_in = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image_in) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image_out, fnum=1)

Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> # Demo drawing on a RGBA canvas >>> # If you initialize an zero rgba canvas, the alpha values are >>> # filled correctly. >>> from kwimage.structs.polygon import * # NOQA >>> s = 16 >>> self = Polygon.random(n_holes=1, rng=32).scale(s) >>> image_in = np.zeros((s, s, 4), dtype=np.float32) >>> image_out = self.draw_on(image_in, color='black') >>> assert np.all(image_out[..., 0:3] == 0) >>> assert not np.all(image_out[..., 3] == 1) >>> assert not np.all(image_out[..., 3] == 0)
Example
>>> import kwimage >>> color = 'blue' >>> self = kwimage.Polygon.random(n_holes=1).scale(128) >>> image = np.zeros((128, 128), dtype=np.float32) >>> # Test drawing on all channel + dtype combinations >>> im3 = np.random.rand(128, 128, 3) >>> im_chans = { >>> 'im3': im3, >>> 'im1': kwimage.convert_colorspace(im3, 'rgb', 'gray'), >>> #'im0': im3[..., 0], >>> 'im4': kwimage.convert_colorspace(im3, 'rgb', 'rgba'), >>> } >>> inputs = {} >>> for k, im in im_chans.items(): >>> inputs[k + '_f01'] = (kwimage.ensure_float01(im.copy()), {'alpha': None}) >>> inputs[k + '_u255'] = (kwimage.ensure_uint255(im.copy()), {'alpha': None}) >>> inputs[k + '_f01_a'] = (kwimage.ensure_float01(im.copy()), {'alpha': 0.5}) >>> inputs[k + '_u255_a'] = (kwimage.ensure_uint255(im.copy()), {'alpha': 0.5}) >>> # Check cases when image is/isnot written inplace Construct images >>> # with different dtypes / channels and run a draw_on with different >>> # keyword args. For each combination, demo if that results in an >>> # implace operation or not. >>> rows = [] >>> outputs = {} >>> for k, v in inputs.items(): >>> im, kw = v >>> outputs[k] = self.draw_on(im, color=color, **kw) >>> inplace = outputs[k] is im >>> rows.append({'key': k, 'inplace': inplace}) >>> # xdoc: +REQUIRES(module:pandas) >>> import pandas as pd >>> df = pd.DataFrame(rows).sort_values('inplace') >>> print(df.to_string()) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=2, doclf=True) >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nCols=2, nRows=len(inputs)) >>> for k in inputs.keys(): >>> kwplot.imshow(inputs[k][0], fnum=2, pnum=pnum_(), title=k) >>> kwplot.imshow(outputs[k], fnum=2, pnum=pnum_(), title=k) >>> kwplot.show_if_requested()
Example
>>> # Test empty polygon draw >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.from_coco([]) >>> image_in = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image_in)
Example
>>> # Test stupid large polygon draw >>> from kwimage.structs.polygon import * # NOQA >>> from kwimage.structs.polygon import _generic >>> import kwimage >>> self = kwimage.Polygon.random().scale(2e11) >>> image = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image)
- draw(color='blue', ax=None, alpha=1.0, radius=1, setlim=False, border=None, linewidth=None, edgecolor=None, facecolor=None, fill=True, vertex=False, vertexcolor=None)[source]¶
Draws polygon in a matplotlib axes. See draw_on for in-memory image modification.
- Parameters
setlim (bool) – if True ensures the limits of the axes contains the polygon
color (str | Tuple) – coercable color. Default color if specific colors are not given.
alpha (float) – fill transparency
fill (bool) – if True fill the polygon with facecolor, otherwise just draw the border if linewidth > 0
setlim (bool) – if True, modify the x and y limits of the matplotlib axes such that the polygon is can be seen.
border (bool) – if True, draws an edge border on the polygon. DEPRECATED. Use linewidth instead.
linewidth (bool) – width of the border
edgecolor (None | Any) – if None, uses the value of
color. Otherwise the color of the border when linewidth > 0. Extended types Coercable[kwimage.Color].facecolor (None | Any) – if None, uses the value of
color. Otherwise, color of the border when fill=True. Extended types Coercable[kwimage.Color].vertex (float) – if non-zero, draws vertexes on the polygon with this radius.
vertexcolor (Any) – color of vertexes Extended types Coercable[kwimage.Color].
- Returns
None for am empty polygon
- Return type
matplotlib.patches.PathPatch | None
Todo
[ ] Rework arguments in favor of matplotlib standards
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1) >>> self = self.scale(100) >>> # xdoc: +REQUIRES(--show) >>> kwargs = dict(edgecolor='orangered', facecolor='dodgerblue', linewidth=10) >>> self.draw(**kwargs) >>> import kwplot >>> kwplot.autompl() >>> from matplotlib import pyplot as plt >>> kwplot.figure(fnum=2) >>> self.draw(setlim=True, **kwargs)
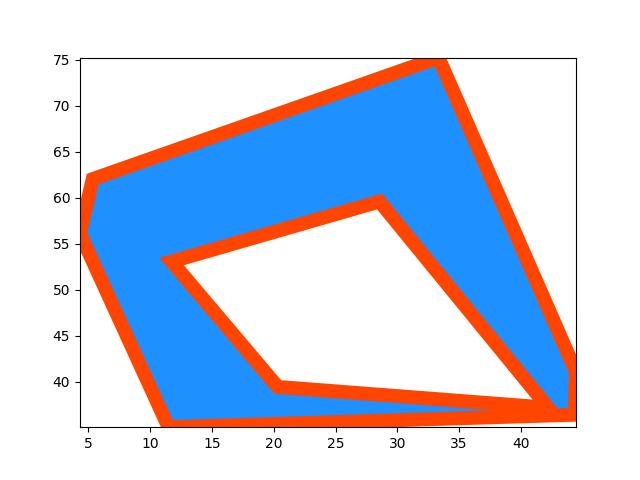
Example
>>> # xdoc: +REQUIRES(module:kwplot) >>> # xdoc: +REQUIRES(--show) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1, rng=33202) >>> import textwrap >>> # Test over a range of parameters >>> basis = { >>> 'linewidth': [0, 4], >>> 'edgecolor': [None, 'gold'], >>> 'facecolor': ['purple'], >>> 'fill': [True, False], >>> 'alpha': [1.0, 0.5], >>> 'vertex': [0, 0.01], >>> 'vertexcolor': ['green'], >>> } >>> grid = list(ub.named_product(basis)) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nSubplots=len(grid)) >>> for kwargs in grid: >>> fig = kwplot.figure(fnum=1, pnum=pnum_()) >>> ax = fig.gca() >>> self.draw(ax=ax, **kwargs) >>> title = ub.repr2(kwargs, compact=True) >>> title = '\n'.join(textwrap.wrap( >>> title.replace(',', ' '), break_long_words=False, >>> width=60)) >>> ax.set_title(title, fontdict={'fontsize': 8}) >>> ax.grid(False) >>> ax.set_xticks([]) >>> ax.set_yticks([]) >>> fig.subplots_adjust(wspace=0.5, hspace=0.3, bottom=0.001, top=0.97) >>> kwplot.show_if_requested()

- class kwimage.structs.PolygonList(data, meta=None)[source]¶
Bases:
ObjectListStores and allows manipluation of multiple polygons, usually within the same image.
- to_mask_list(dims=None, pixels_are='points')[source]¶
Converts all items to masks
- Returns
kwimage.MaskList
- to_segmentation_list()[source]¶
Converts all items to segmentation objects
- Returns
kwimage.SegmentationList
- to_geojson(as_collection=False)[source]¶
Converts a list of polygons/multipolygons to a geojson structure
- Parameters
as_collection (bool) – if True, wraps the polygon geojson items in a geojson feature collection, otherwise just return a list of items.
- Returns
items or geojson data
- Return type
List[Dict] | Dict
Example
>>> import kwimage >>> data = [kwimage.Polygon.random(), >>> kwimage.Polygon.random(n_holes=1), >>> kwimage.MultiPolygon.random(n_holes=1), >>> kwimage.MultiPolygon.random()] >>> self = kwimage.PolygonList(data) >>> geojson = self.to_geojson(as_collection=True) >>> items = self.to_geojson(as_collection=False) >>> print('geojson = {}'.format(ub.repr2(geojson, nl=-2, precision=1))) >>> print('items = {}'.format(ub.repr2(items, nl=-2, precision=1)))
- class kwimage.structs.Segmentation(data, format=None)[source]¶
Bases:
_WrapperObjectEither holds a MultiPolygon, Polygon, or Mask
- Parameters
data (object) – the underlying object
format (str) – either ‘mask’, ‘polygon’, or ‘multipolygon’
- classmethod random(rng=None)[source]¶
Example
>>> self = Segmentation.random() >>> print('self = {!r}'.format(self)) >>> # xdoc: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> self.draw() >>> kwplot.show_if_requested()
- property meta¶
- class kwimage.structs.SegmentationList(data, meta=None)[source]¶
Bases:
ObjectListStore and manipulate multiple segmentations (masks or polygons), usually within the same image
- kwimage.structs.smooth_prob(prob, k=3, inplace=False, eps=1e-09)[source]¶
Smooths the probability map, but preserves the magnitude of the peaks.
Note
even if inplace is true, we still need to make a copy of the input array, however, we do ensure that it is cleaned up before we leave the function scope.
sigma=0.8 @ k=3, sigma=1.1 @ k=5, sigma=1.4 @ k=7