kwimage package¶
Subpackages¶
- kwimage.algo package
- kwimage.cli package
- kwimage.gis package
- kwimage.structs package
- Submodules
- kwimage.structs._generic module
- kwimage.structs.boxes module
- kwimage.structs.coords module
- kwimage.structs.detections module
- kwimage.structs.heatmap module
- kwimage.structs.mask module
- kwimage.structs.points module
- kwimage.structs.polygon module
- kwimage.structs.segmentation module
- kwimage.structs.single_box module
- Module contents
BoxBox.formatBox.dataBox.random()Box.from_slice()Box.from_shapely()Box.from_dsize()Box.from_data()Box.coerce()Box.dsizeBox.translate()Box.warp()Box.scale()Box.clip()Box.quantize()Box.copy()Box.round()Box.pad()Box.resize()Box.intersection()Box.union_hull()Box.to_ltrb()Box.to_xywh()Box.to_cxywh()Box.toformat()Box.astype()Box.corners()Box.to_boxes()Box.aspect_ratioBox.centerBox.center_xBox.center_yBox.widthBox.heightBox.tl_xBox.tl_yBox.br_xBox.br_yBox.dtypeBox.areaBox.to_slice()Box.to_shapely()Box.to_polygon()Box.to_coco()Box.draw_on()Box.draw()
BoxesBoxes.random()Boxes.copy()Boxes.concatenate()Boxes.compress()Boxes.take()Boxes.is_tensor()Boxes.is_numpy()Boxes._implBoxes.deviceBoxes.astype()Boxes.round()Boxes.quantize()Boxes.numpy()Boxes.tensor()Boxes.ious()Boxes.iooas()Boxes.isect_area()Boxes.intersection()Boxes.union_hull()Boxes.bounding_box()Boxes.contains()Boxes.view()Boxes._ensure_nonnegative_extent()
CoordsCoords.dtypeCoords.dimCoords.shapeCoords.copy()Coords.random()Coords.is_numpy()Coords.is_tensor()Coords.compress()Coords.take()Coords.astype()Coords.round()Coords.view()Coords.concatenate()Coords.deviceCoords._implCoords.tensor()Coords.numpy()Coords.reorder_axes()Coords.warp()Coords._warp_imgaug()Coords.to_imgaug()Coords.from_imgaug()Coords.scale()Coords.translate()Coords.rotate()Coords._rectify_about()Coords.fill()Coords.soft_fill()Coords.draw_on()Coords.draw()
DetectionsDetections.copy()Detections.coerce()Detections.from_coco_annots()Detections.to_coco()Detections.boxesDetections.class_idxsDetections.scoresDetections.probsDetections.weightsDetections.classesDetections.num_boxes()Detections.warp()Detections.scale()Detections.translate()Detections.concatenate()Detections.argsort()Detections.sort()Detections.compress()Detections.take()Detections.deviceDetections.is_tensor()Detections.is_numpy()Detections.numpy()Detections.dtypeDetections.tensor()Detections.demo()Detections.random()
HeatmapMaskMask.dtypeMask.random()Mask.demo()Mask.from_text()Mask.copy()Mask.union()Mask.intersection()Mask.shapeMask.areaMask.get_patch()Mask.get_xywh()Mask.bounding_box()Mask.get_polygon()Mask.to_mask()Mask.to_boxes()Mask.to_multi_polygon()Mask.get_convex_hull()Mask.iou()Mask.coerce()Mask._to_coco()Mask.to_coco()
MaskListMultiPolygonMultiPolygon.random()MultiPolygon.fill()MultiPolygon.to_multi_polygon()MultiPolygon.to_boxes()MultiPolygon.to_box()MultiPolygon.bounding_box()MultiPolygon.box()MultiPolygon.to_mask()MultiPolygon.to_relative_mask()MultiPolygon.coerce()MultiPolygon.to_shapely()MultiPolygon.from_shapely()MultiPolygon.from_geojson()MultiPolygon.to_geojson()MultiPolygon.from_coco()MultiPolygon._to_coco()MultiPolygon.to_coco()MultiPolygon.swap_axes()MultiPolygon.draw_on()
PointsPoints.shapePoints.xyPoints.random()Points.is_numpy()Points.is_tensor()Points._implPoints.tensor()Points.round()Points.numpy()Points.draw_on()Points.draw()Points.compress()Points.take()Points.concatenate()Points.to_coco()Points._to_coco()Points.coerce()Points._from_coco()Points.from_coco()
PointsListPolygonPolygon.exteriorPolygon.interiorsPolygon.circle()Polygon.regular()Polygon.star()Polygon.random()Polygon._implPolygon.to_mask()Polygon.to_relative_mask()Polygon._to_cv_countours()Polygon.coerce()Polygon.from_shapely()Polygon.from_wkt()Polygon.from_geojson()Polygon.to_shapely()Polygon.to_geojson()Polygon.to_wkt()Polygon.from_coco()Polygon._to_coco()Polygon.to_coco()Polygon.to_multi_polygon()Polygon.to_boxes()Polygon.centroidPolygon.to_box()Polygon.bounding_box()Polygon.box()Polygon.bounding_box_polygon()Polygon.copy()Polygon.clip()Polygon.fill()Polygon.draw_on()Polygon.draw()Polygon._ensure_vertex_order()Polygon.interpolate()Polygon.morph()
PolygonListSegmentationSegmentationListsmooth_prob()
- Submodules
Submodules¶
- kwimage._im_color_data module
- kwimage._internal module
- kwimage.im_alphablend module
- kwimage.im_color module
ColorColor.coerce()Color.forimage()Color._forimage()Color.ashex()Color.as255()Color.as01()Color._is_base01()Color._is_base255()Color._hex_to_01()Color._ensure_color01()Color._255_to_01()Color._string_to_01()Color.named_colors()Color.distinct()Color.random()Color.distance()Color.interpolate()Color.to_image()Color.adjust()
- kwimage.im_core module
- kwimage.im_cv2 module
- kwimage.im_demodata module
- kwimage.im_draw module
_draw_text_on_image_pil()draw_text_on_image()_text_sizes()draw_clf_on_image()draw_boxes_on_image()draw_line_segments_on_image()_broadcast_colors()make_heatmask()make_orimask()make_vector_field()draw_vector_field()draw_header_text()fill_nans_with_checkers()_masked_checkerboard()nodata_checkerboard()
- kwimage.im_filter module
- kwimage.im_io module
- kwimage.im_runlen module
- kwimage.im_stack module
- kwimage.transform module
TransformMatrixLinearAffineAffine.shapeAffine.concise()Affine.from_shapely()Affine.from_affine()Affine.from_gdal()Affine.from_skimage()Affine.coerce()Affine.eccentricity()Affine.to_affine()Affine.to_gdal()Affine.to_shapely()Affine.to_skimage()Affine.scale()Affine.translate()Affine._scale_translate()Affine.rotate()Affine.random()Affine.random_params()Affine.decompose()Affine.affine()Affine.fit()Affine.fliprot()
Projective
- kwimage.util_warp module
_coordinate_grid()warp_tensor()subpixel_align()subpixel_set()subpixel_accum()subpixel_maximum()subpixel_minimum()subpixel_slice()subpixel_translate()_padded_slice()_ensure_arraylike()_rectify_slice()_warp_tensor_cv2()warp_points()remove_homog()add_homog()subpixel_getvalue()subpixel_setvalue()_bilinear_coords()
Module contents¶
The Kitware Image Module (kwimage) contains functions to accomplish lower-level image operations via a high level API.
Read the docs |
|
Gitlab (main) |
|
Github (mirror) |
|
Pypi |
Module features:
Image reader / writer functions with multiple backends
Wrapers around opencv that simplify and extend its functionality
Annotation datastructure with configurable backends.
Many function have awareness of torch tensors and can be used interchangably with ndarrays.
Misc image manipulation functions
- class kwimage.Affine(matrix)[source]¶
Bases:
ProjectiveA thin wraper around a 3x3 matrix that represents an affine transform
- Implements methods for:
creating random affine transforms
decomposing the matrix
finding a best-fit transform between corresponding points
TODO: - [ ] fully rational transform
Example
>>> import kwimage >>> import math >>> image = kwimage.grab_test_image() >>> theta = 0.123 * math.tau >>> components = { >>> 'rotate': kwimage.Affine.affine(theta=theta), >>> 'scale': kwimage.Affine.affine(scale=0.5), >>> 'shear': kwimage.Affine.affine(shearx=0.2), >>> 'translation': kwimage.Affine.affine(offset=(100, 200)), >>> 'rotate+translate': kwimage.Affine.affine(theta=0.123 * math.tau, about=(256, 256)), >>> 'random composed': kwimage.Affine.random(scale=(0.5, 1.5), translate=(-20, 20), theta=(-theta, theta), shearx=(0, .4), rng=900558176210808600), >>> } >>> warp_stack = [] >>> for key, aff in components.items(): ... warp = kwimage.warp_affine(image, aff) ... warp = kwimage.draw_text_on_image( ... warp, ... ub.urepr(aff.matrix, nl=1, nobr=1, precision=2, si=1, sv=1, with_dtype=0), ... org=(1, 1), ... valign='top', halign='left', ... fontScale=0.8, color='kw_blue', ... border={'thickness': 3}, ... ) ... warp = kwimage.draw_header_text(warp, key, color='kw_green') ... warp_stack.append(warp) >>> warp_canvas = kwimage.stack_images_grid(warp_stack, chunksize=3, pad=10, bg_value='kitware_gray') >>> # xdoctest: +REQUIRES(module:sympy) >>> import sympy >>> # Shows the symbolic construction of the code >>> # https://groups.google.com/forum/#!topic/sympy/k1HnZK_bNNA >>> from sympy.abc import theta >>> params = x0, y0, sx, sy, theta, shearx, tx, ty = sympy.symbols( >>> 'x0, y0, sx, sy, theta, shearx, tx, ty') >>> theta = sympy.symbols('theta') >>> # move the center to 0, 0 >>> tr1_ = np.array([[1, 0, -x0], >>> [0, 1, -y0], >>> [0, 0, 1]]) >>> # Define core components of the affine transform >>> S = np.array([ # scale >>> [sx, 0, 0], >>> [ 0, sy, 0], >>> [ 0, 0, 1]]) >>> E = np.array([ # x-shear >>> [1, shearx, 0], >>> [0, 1, 0], >>> [0, 0, 1]]) >>> R = np.array([ # rotation >>> [sympy.cos(theta), -sympy.sin(theta), 0], >>> [sympy.sin(theta), sympy.cos(theta), 0], >>> [ 0, 0, 1]]) >>> T = np.array([ # translation >>> [ 1, 0, tx], >>> [ 0, 1, ty], >>> [ 0, 0, 1]]) >>> # Contruct the affine 3x3 about the origin >>> aff0 = np.array(sympy.simplify(T @ R @ E @ S)) >>> # move 0, 0 back to the specified origin >>> tr2_ = np.array([[1, 0, x0], >>> [0, 1, y0], >>> [0, 0, 1]]) >>> # combine transformations >>> aff = tr2_ @ aff0 @ tr1_ >>> print('aff = {}'.format(ub.urepr(aff.tolist(), nl=1))) >>> # This could be prettier >>> texts = { >>> 'Translation': sympy.pretty(R), >>> 'Rotation': sympy.pretty(R), >>> 'shEar-X': sympy.pretty(E), >>> 'Scale': sympy.pretty(S), >>> } >>> print(ub.urepr(texts, nl=2, sv=1)) >>> equation_stack = [] >>> for text, m in texts.items(): >>> render_canvas = kwimage.draw_text_on_image(None, m, color='kw_blue', fontScale=1.0) >>> render_canvas = kwimage.draw_header_text(render_canvas, text, color='kw_green') >>> render_canvas = kwimage.imresize(render_canvas, scale=1.3) >>> equation_stack.append(render_canvas) >>> equation_canvas = kwimage.stack_images(equation_stack, pad=10, axis=1, bg_value='kitware_gray') >>> render_canvas = kwimage.draw_text_on_image(None, sympy.pretty(aff), color='kw_blue', fontScale=1.0) >>> render_canvas = kwimage.draw_header_text(render_canvas, 'Full Equation With Pre-Shift', color='kw_green') >>> # xdoctest: -REQUIRES(module:sympy) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> canvas = kwimage.stack_images([warp_canvas, equation_canvas, render_canvas], pad=20, axis=0, bg_value='kitware_gray', resize='larger') >>> canvas = kwimage.draw_header_text(canvas, 'Affine matrixes can represent', color='kw_green') >>> kwplot.imshow(canvas) >>> fig = plt.gcf() >>> fig.set_size_inches(13, 13)

Example
>>> import kwimage >>> self = kwimage.Affine(np.eye(3)) >>> m1 = np.eye(3) @ self >>> m2 = self @ np.eye(3)
Example
>>> from kwimage.transform import * # NOQA >>> m = {} >>> # Works, and returns a Affine >>> m[len(m)] = x = Affine.random() @ np.eye(3) >>> assert isinstance(x, Affine) >>> m[len(m)] = x = Affine.random() @ None >>> assert isinstance(x, Affine) >>> # Works, and returns an ndarray >>> m[len(m)] = x = np.eye(3) @ Affine.random() >>> assert isinstance(x, np.ndarray) >>> # Works, and returns an Matrix >>> m[len(m)] = x = Affine.random() @ Matrix.random(3) >>> assert isinstance(x, Matrix) >>> m[len(m)] = x = Matrix.random(3) @ Affine.random() >>> assert isinstance(x, Matrix) >>> print('m = {}'.format(ub.urepr(m)))
- property shape¶
- concise()[source]¶
Return a concise coercable dictionary representation of this matrix
- Returns:
- a small serializable dict that can be passed
to
Affine.coerce()to reconstruct this object.
- Return type:
- Returns:
dictionary with consise parameters
- Return type:
Dict
Example
>>> import kwimage >>> self = kwimage.Affine.random(rng=0, scale=1) >>> params = self.concise() >>> assert np.allclose(Affine.coerce(params).matrix, self.matrix) >>> print('params = {}'.format(ub.urepr(params, nl=1, precision=2))) params = { 'offset': (0.08, 0.38), 'theta': 0.08, 'type': 'affine', }
Example
>>> import kwimage >>> self = kwimage.Affine.random(rng=0, scale=2, offset=0) >>> params = self.concise() >>> assert np.allclose(Affine.coerce(params).matrix, self.matrix) >>> print('params = {}'.format(ub.urepr(params, nl=1, precision=2))) params = { 'scale': 2.00, 'theta': 0.04, 'type': 'affine', }
- classmethod from_shapely(sh_aff)[source]¶
Shapely affine tuples are in the format (a, b, d, e, x, y)
- classmethod coerce(data=None, **kwargs)[source]¶
Attempt to coerce the data into an affine object
- Parameters:
data – some data we attempt to coerce to an Affine matrix
**kwargs – some data we attempt to coerce to an Affine matrix, mutually exclusive with data.
- Returns:
Affine
Example
>>> import kwimage >>> import skimage.transform >>> kwimage.Affine.coerce({'type': 'affine', 'matrix': [[1, 0, 0], [0, 1, 0]]}) >>> kwimage.Affine.coerce({'scale': 2}) >>> kwimage.Affine.coerce({'offset': 3}) >>> kwimage.Affine.coerce(np.eye(3)) >>> kwimage.Affine.coerce(None) >>> kwimage.Affine.coerce({}) >>> kwimage.Affine.coerce(skimage.transform.AffineTransform(scale=30))
- eccentricity()[source]¶
Eccentricity of the ellipse formed by this affine matrix
- Returns:
- large when there are big scale differences in principle
directions or skews.
- Return type:
References
[WikiConic]Example
>>> import kwimage >>> kwimage.Affine.random(rng=432).eccentricity()
- to_gdal()[source]¶
Convert to a gdal tuple (c, a, b, f, d, e)
- Returns:
Tuple[float, float, float, float, float, float]
- to_shapely()[source]¶
Returns a matrix suitable for shapely.affinity.affine_transform
- Returns:
Tuple[float, float, float, float, float, float]
Example
>>> import kwimage >>> self = kwimage.Affine.random() >>> sh_transform = self.to_shapely() >>> # Transform points with kwimage and shapley >>> import shapely >>> from shapely.affinity import affine_transform >>> kw_poly = kwimage.Polygon.random() >>> kw_warp_poly = kw_poly.warp(self) >>> sh_poly = kw_poly.to_shapely() >>> sh_warp_poly = affine_transform(sh_poly, sh_transform) >>> kw_warp_poly_recon = kwimage.Polygon.from_shapely(sh_warp_poly) >>> assert np.allclose(kw_warp_poly_recon.exterior.data, kw_warp_poly_recon.exterior.data)
- to_skimage()[source]¶
- Returns:
skimage.transform.AffineTransform
Example
>>> import kwimage >>> self = kwimage.Affine.random() >>> tf = self.to_skimage() >>> # Transform points with kwimage and scikit-image >>> kw_poly = kwimage.Polygon.random() >>> kw_warp_xy = kw_poly.warp(self.matrix).exterior.data >>> sk_warp_xy = tf(kw_poly.exterior.data) >>> assert np.allclose(sk_warp_xy, sk_warp_xy)
- classmethod scale(scale)[source]¶
Create a scale Affine object
- Parameters:
scale (float | Tuple[float, float]) – x, y scale factor
- Returns:
Affine
- classmethod translate(offset)[source]¶
Create a translation Affine object
- Parameters:
offset (float | Tuple[float, float]) – x, y translation factor
- Returns:
Affine
Benchmark
>>> # xdoctest: +REQUIRES(--benchmark) >>> # It is ~3x faster to use the more specific method >>> import timerit >>> import kwimage >>> # >>> offset = np.random.rand(2) >>> ti = timerit.Timerit(100, bestof=10, verbose=2) >>> for timer in ti.reset('time'): >>> with timer: >>> kwimage.Affine.translate(offset) >>> # >>> for timer in ti.reset('time'): >>> with timer: >>> kwimage.Affine.affine(offset=offset)
- classmethod rotate(theta)[source]¶
Create a rotation Affine object
- Parameters:
theta (float) – counter-clockwise rotation angle in radians
- Returns:
Affine
- classmethod random(shape=None, rng=None, **kw)[source]¶
Create a random Affine object
- Parameters:
rng – random number generator
**kw – passed to
Affine.random_params(). can contain coercable random distributions for scale, offset, about, theta, and shearx.
- Returns:
Affine
- classmethod random_params(rng=None, **kw)[source]¶
- Parameters:
rng – random number generator
**kw – can contain coercable random distributions for scale, offset, about, theta, and shearx.
- Returns:
affine parameters suitable to be passed to Affine.affine
- Return type:
Dict
Todo
[ ] improve kwargs parameterization
- decompose()[source]¶
Decompose the affine matrix into its individual scale, translation, rotation, and skew parameters.
- Returns:
decomposed offset, scale, theta, and shearx params
- Return type:
Dict
References
[SE3521141][SE70357473]https://stackoverflow.com/questions/70357473/how-to-decompose-a-2x2-affine-matrix-with-sympy
[WikiTranMat][WikiShear]Example
>>> from kwimage.transform import * # NOQA >>> self = Affine.random() >>> params = self.decompose() >>> recon = Affine.coerce(**params) >>> params2 = recon.decompose() >>> pt = np.vstack([np.random.rand(2, 1), [1]]) >>> result1 = self.matrix[0:2] @ pt >>> result2 = recon.matrix[0:2] @ pt >>> assert np.allclose(result1, result2)
>>> self = Affine.scale(0.001) @ Affine.random() >>> params = self.decompose() >>> self.det()
Example
>>> # xdoctest: +REQUIRES(module:sympy) >>> # Test decompose with symbolic matrices >>> from kwimage.transform import * # NOQA >>> self = Affine.random().rationalize() >>> self.decompose()
Example
>>> # xdoctest: +REQUIRES(module:pandas) >>> from kwimage.transform import * # NOQA >>> import kwimage >>> import pandas as pd >>> # Test consistency of decompose + reconstruct >>> param_grid = list(ub.named_product({ >>> 'theta': np.linspace(-4 * np.pi, 4 * np.pi, 3), >>> 'shearx': np.linspace(- 10 * np.pi, 10 * np.pi, 4), >>> })) >>> def normalize_angle(radian): >>> return np.arctan2(np.sin(radian), np.cos(radian)) >>> for pextra in param_grid: >>> params0 = dict(scale=(3.05, 3.07), offset=(10.5, 12.1), **pextra) >>> self = recon0 = kwimage.Affine.affine(**params0) >>> self.decompose() >>> # Test drift with multiple decompose / reconstructions >>> params_list = [params0] >>> recon_list = [recon0] >>> n = 4 >>> for _ in range(n): >>> prev = recon_list[-1] >>> params = prev.decompose() >>> recon = kwimage.Affine.coerce(**params) >>> params_list.append(params) >>> recon_list.append(recon) >>> params_df = pd.DataFrame(params_list) >>> #print('params_list = {}'.format(ub.urepr(params_list, nl=1, precision=5))) >>> print(params_df) >>> assert ub.allsame(normalize_angle(params_df['theta']), eq=np.isclose) >>> assert ub.allsame(params_df['shearx'], eq=np.allclose) >>> assert ub.allsame(params_df['scale'], eq=np.allclose) >>> assert ub.allsame(params_df['offset'], eq=np.allclose)
- classmethod affine(scale=None, offset=None, theta=None, shear=None, about=None, shearx=None, array_cls=None, math_mod=None, **kwargs)[source]¶
Create an affine matrix from high-level parameters
- Parameters:
scale (float | Tuple[float, float]) – x, y scale factor
offset (float | Tuple[float, float]) – x, y translation factor
theta (float) – counter-clockwise rotation angle in radians
shearx (float) – shear factor parallel to the x-axis.
about (float | Tuple[float, float]) – x, y location of the origin
shear (float) – BROKEN, dont use. counter-clockwise shear angle in radians
Todo
- [ ] Add aliases? -
origin : alias for about rotation : alias for theta translation : alias for offset
- Returns:
the constructed Affine object
- Return type:
Example
>>> from kwimage.transform import * # NOQA >>> rng = kwarray.ensure_rng(None) >>> scale = rng.randn(2) * 10 >>> offset = rng.randn(2) * 10 >>> about = rng.randn(2) * 10 >>> theta = rng.randn() * 10 >>> shearx = rng.randn() * 10 >>> # Create combined matrix from all params >>> F = Affine.affine( >>> scale=scale, offset=offset, theta=theta, shearx=shearx, >>> about=about) >>> # Test that combining components matches >>> S = Affine.affine(scale=scale) >>> T = Affine.affine(offset=offset) >>> R = Affine.affine(theta=theta) >>> E = Affine.affine(shearx=shearx) >>> O = Affine.affine(offset=about) >>> # combine (note shear must be on the RHS of rotation) >>> alt = O @ T @ R @ E @ S @ O.inv() >>> print('F = {}'.format(ub.urepr(F.matrix.tolist(), nl=1))) >>> print('alt = {}'.format(ub.urepr(alt.matrix.tolist(), nl=1))) >>> assert np.all(np.isclose(alt.matrix, F.matrix)) >>> pt = np.vstack([np.random.rand(2, 1), [[1]]]) >>> warp_pt1 = (F.matrix @ pt) >>> warp_pt2 = (alt.matrix @ pt) >>> assert np.allclose(warp_pt2, warp_pt1)
Sympy
>>> # xdoctest: +SKIP >>> import sympy >>> # Shows the symbolic construction of the code >>> # https://groups.google.com/forum/#!topic/sympy/k1HnZK_bNNA >>> from sympy.abc import theta >>> params = x0, y0, sx, sy, theta, shearx, tx, ty = sympy.symbols( >>> 'x0, y0, sx, sy, theta, shearx, tx, ty') >>> # move the center to 0, 0 >>> tr1_ = np.array([[1, 0, -x0], >>> [0, 1, -y0], >>> [0, 0, 1]]) >>> # Define core components of the affine transform >>> S = np.array([ # scale >>> [sx, 0, 0], >>> [ 0, sy, 0], >>> [ 0, 0, 1]]) >>> E = np.array([ # x-shear >>> [1, shearx, 0], >>> [0, 1, 0], >>> [0, 0, 1]]) >>> R = np.array([ # rotation >>> [sympy.cos(theta), -sympy.sin(theta), 0], >>> [sympy.sin(theta), sympy.cos(theta), 0], >>> [ 0, 0, 1]]) >>> T = np.array([ # translation >>> [ 1, 0, tx], >>> [ 0, 1, ty], >>> [ 0, 0, 1]]) >>> # Contruct the affine 3x3 about the origin >>> aff0 = np.array(sympy.simplify(T @ R @ E @ S)) >>> # move 0, 0 back to the specified origin >>> tr2_ = np.array([[1, 0, x0], >>> [0, 1, y0], >>> [0, 0, 1]]) >>> # combine transformations >>> aff = tr2_ @ aff0 @ tr1_ >>> print('aff = {}'.format(ub.urepr(aff.tolist(), nl=1)))
- classmethod fit(pts1, pts2)[source]¶
Fit an affine transformation between a set of corresponding points
- Parameters:
pts1 (ndarray) – An Nx2 array of points in “space 1”.
pts2 (ndarray) – A corresponding Nx2 array of points in “space 2”
- Returns:
a transform that warps from “space1” to “space2”.
- Return type:
Note
An affine matrix has 6 degrees of freedom, so at least 3 non-colinear xy-point pairs are needed.
References
..[Lowe04] https://www.cs.ubc.ca/~lowe/papers/ijcv04.pdf page 22
Example
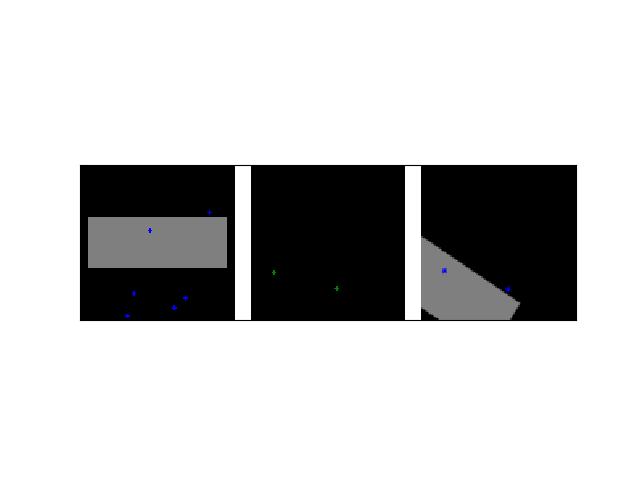
>>> # Create a set of points, warp them, then recover the warp >>> import kwimage >>> points = kwimage.Points.random(6).scale(64) >>> #A1 = kwimage.Affine.affine(scale=0.9, theta=-3.2, offset=(2, 3), about=(32, 32), skew=2.3) >>> #A2 = kwimage.Affine.affine(scale=0.8, theta=0.8, offset=(2, 0), about=(32, 32)) >>> A1 = kwimage.Affine.random() >>> A2 = kwimage.Affine.random() >>> A12_real = A2 @ A1.inv() >>> points1 = points.warp(A1) >>> points2 = points.warp(A2) >>> # Recover the warp >>> pts1, pts2 = points1.xy, points2.xy >>> A_recovered = kwimage.Affine.fit(pts1, pts2) >>> assert np.all(np.isclose(A_recovered.matrix, A12_real.matrix)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> base1 = np.zeros((96, 96, 3)) >>> base1[32:-32, 5:-5] = 0.5 >>> base2 = np.zeros((96, 96, 3)) >>> img1 = points1.draw_on(base1, radius=3, color='blue') >>> img2 = points2.draw_on(base2, radius=3, color='green') >>> img1_warp = kwimage.warp_affine(img1, A_recovered) >>> canvas = kwimage.stack_images([img1, img2, img1_warp], pad=10, axis=1, bg_value=(1., 1., 1.)) >>> kwplot.imshow(canvas)

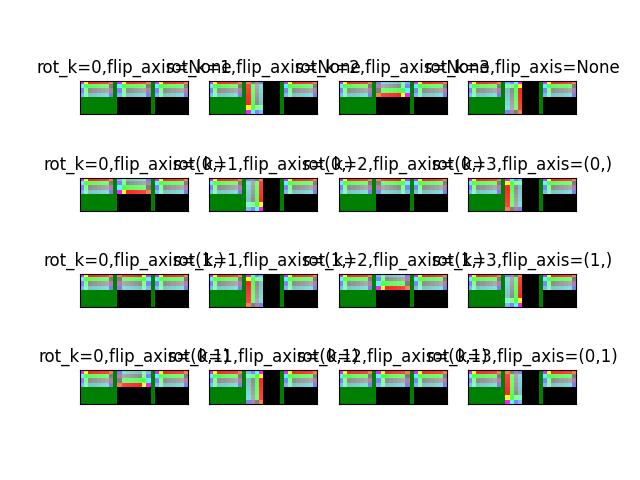
- classmethod fliprot(flip_axis=None, rot_k=0, axes=(0, 1), canvas_dsize=None)[source]¶
Creates a flip/rotation transform with respect to an image of a given size in the positive quadrent. (i.e. warped data within the specified canvas size will stay in the positive quadrant)
- Parameters:
flip_axis (int) – the axis dimension to flip. I.e. 0 flips the y-axis and 1-flips the x-axis.
rot_k (int) – number of counterclockwise 90 degree rotations that occur after the flips.
axes (Tuple[int, int]) – The axis ordering. Unhandled in this version. Dont change this.
canvas_dsize (Tuple[int, int]) – The width / height of the canvas the fliprot is applied in.
- Returns:
The affine matrix representing the canvas-aligned flip and rotation.
- Return type:
Note
Requiring that the image size is known makes this a place that errors could occur depending on your interpretation of pixels as points or areas. There is probably a better way to describe the issue, but the second doctest shows the issue when trying to use warp-affine’s auto-dsize feature. See [MR81] for details.
References
CommandLine
xdoctest -m kwimage.transform Affine.fliprot:0 --show xdoctest -m kwimage.transform Affine.fliprot:1 --show
Example
>>> import kwimage >>> H, W = 64, 128 >>> canvas_dsize = (W, H) >>> box1 = kwimage.Boxes.random(1).scale((W, H)).quantize() >>> ltrb = box1.data >>> rot_k = 4 >>> annot = box1 >>> annot = box1.to_polygons()[0] >>> annot1 = annot.copy() >>> # The first 8 are the cannonically unique group elements >>> fliprot_params = [ >>> {'rot_k': 0, 'flip_axis': None}, >>> {'rot_k': 1, 'flip_axis': None}, >>> {'rot_k': 2, 'flip_axis': None}, >>> {'rot_k': 3, 'flip_axis': None}, >>> {'rot_k': 0, 'flip_axis': (0,)}, >>> {'rot_k': 1, 'flip_axis': (0,)}, >>> {'rot_k': 2, 'flip_axis': (0,)}, >>> {'rot_k': 3, 'flip_axis': (0,)}, >>> # The rest of these dont result in any different data, but we need to test them >>> {'rot_k': 0, 'flip_axis': (1,)}, >>> {'rot_k': 1, 'flip_axis': (1,)}, >>> {'rot_k': 2, 'flip_axis': (1,)}, >>> {'rot_k': 3, 'flip_axis': (1,)}, >>> {'rot_k': 0, 'flip_axis': (0, 1)}, >>> {'rot_k': 1, 'flip_axis': (0, 1)}, >>> {'rot_k': 2, 'flip_axis': (0, 1)}, >>> {'rot_k': 3, 'flip_axis': (0, 1)}, >>> ] >>> results = [] >>> for params in fliprot_params: >>> tf = kwimage.Affine.fliprot(canvas_dsize=canvas_dsize, **params) >>> annot2 = annot.warp(tf) >>> annot3 = annot2.warp(tf.inv()) >>> #annot3 = inv_fliprot_annot(annot2, canvas_dsize=canvas_dsize, **params) >>> results.append({ >>> 'annot2': annot2, >>> 'annot3': annot3, >>> 'params': params, >>> 'tf': tf, >>> 'canvas_dsize': canvas_dsize, >>> }) >>> box = kwimage.Box.coerce([0, 0, W, H], format='xywh') >>> for result in results: >>> params = result['params'] >>> warped = box.warp(result['tf']) >>> print('---') >>> print('params = {}'.format(ub.urepr(params, nl=1))) >>> print('box = {}'.format(ub.urepr(box, nl=1))) >>> print('warped = {}'.format(ub.urepr(warped, nl=1))) >>> print(ub.hzcat(['tf = ', ub.urepr(result['tf'], nl=1)]))
>>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> S = max(W, H) >>> image1 = kwimage.grab_test_image('astro', dsize=(S, S))[:H, :W] >>> pnum_ = kwplot.PlotNums(nCols=4, nSubplots=len(results)) >>> for result in results: >>> #image2 = kwimage.warp_affine(image1.copy(), result['tf'], dsize=(S, S)) # fixme dsize=positive should work here >>> image2 = kwimage.warp_affine(image1.copy(), result['tf'], dsize='positive') # fixme dsize=positive should work here >>> #image3 = kwimage.warp_affine(image2.copy(), result['tf'].inv(), dsize=(S, S)) >>> image3 = kwimage.warp_affine(image2.copy(), result['tf'].inv(), dsize='positive') >>> annot2 = result['annot2'] >>> annot3 = result['annot3'] >>> canvas1 = annot1.draw_on(image1.copy(), edgecolor='kitware_blue', fill=False) >>> canvas2 = annot2.draw_on(image2.copy(), edgecolor='kitware_green', fill=False) >>> canvas3 = annot3.draw_on(image3.copy(), edgecolor='kitware_red', fill=False) >>> canvas = kwimage.stack_images([canvas1, canvas2, canvas3], axis=1, pad=10, bg_value='green') >>> kwplot.imshow(canvas, pnum=pnum_(), title=ub.urepr(result['params'], nl=0, compact=1, nobr=1)) >>> kwplot.show_if_requested()

Example
>>> # Second similar test with a very small image to catch small errors >>> import kwimage >>> H, W = 4, 8 >>> canvas_dsize = (W, H) >>> box1 = kwimage.Boxes.random(1).scale((W, H)).quantize() >>> ltrb = box1.data >>> rot_k = 4 >>> annot = box1 >>> annot = box1.to_polygons()[0] >>> annot1 = annot.copy() >>> # The first 8 are the cannonically unique group elements >>> fliprot_params = [ >>> {'rot_k': 0, 'flip_axis': None}, >>> {'rot_k': 1, 'flip_axis': None}, >>> {'rot_k': 2, 'flip_axis': None}, >>> {'rot_k': 3, 'flip_axis': None}, >>> {'rot_k': 0, 'flip_axis': (0,)}, >>> {'rot_k': 1, 'flip_axis': (0,)}, >>> {'rot_k': 2, 'flip_axis': (0,)}, >>> {'rot_k': 3, 'flip_axis': (0,)}, >>> # The rest of these dont result in any different data, but we need to test them >>> {'rot_k': 0, 'flip_axis': (1,)}, >>> {'rot_k': 1, 'flip_axis': (1,)}, >>> {'rot_k': 2, 'flip_axis': (1,)}, >>> {'rot_k': 3, 'flip_axis': (1,)}, >>> {'rot_k': 0, 'flip_axis': (0, 1)}, >>> {'rot_k': 1, 'flip_axis': (0, 1)}, >>> {'rot_k': 2, 'flip_axis': (0, 1)}, >>> {'rot_k': 3, 'flip_axis': (0, 1)}, >>> ] >>> results = [] >>> for params in fliprot_params: >>> tf = kwimage.Affine.fliprot(canvas_dsize=canvas_dsize, **params) >>> annot2 = annot.warp(tf) >>> annot3 = annot2.warp(tf.inv()) >>> #annot3 = inv_fliprot_annot(annot2, canvas_dsize=canvas_dsize, **params) >>> results.append({ >>> 'annot2': annot2, >>> 'annot3': annot3, >>> 'params': params, >>> 'tf': tf, >>> 'canvas_dsize': canvas_dsize, >>> }) >>> box = kwimage.Box.coerce([0, 0, W, H], format='xywh') >>> print('box = {}'.format(ub.urepr(box, nl=1))) >>> for result in results: >>> params = result['params'] >>> warped = box.warp(result['tf']) >>> print('---') >>> print('params = {}'.format(ub.urepr(params, nl=1))) >>> print('warped = {}'.format(ub.urepr(warped, nl=1))) >>> print(ub.hzcat(['tf = ', ub.urepr(result['tf'], nl=1)]))
>>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> S = max(W, H) >>> image1 = np.linspace(.1, .9, W * H).reshape((H, W)) >>> image1 = kwimage.atleast_3channels(image1) >>> image1[0, :, 0] = 1 >>> image1[:, 0, 2] = 1 >>> image1[1, :, 1] = 1 >>> image1[:, 1, 1] = 1 >>> image1[3, :, 0] = 0.5 >>> image1[:, 7, 1] = 0.5 >>> pnum_ = kwplot.PlotNums(nCols=4, nSubplots=len(results)) >>> # NOTE: setting new_dsize='positive' illustrates an issuew with >>> # the pixel interpretation. >>> new_dsize = (S, S) >>> #new_dsize = 'positive' >>> for result in results: >>> image2 = kwimage.warp_affine(image1.copy(), result['tf'], dsize=new_dsize) >>> image3 = kwimage.warp_affine(image2.copy(), result['tf'].inv(), dsize=new_dsize) >>> annot2 = result['annot2'] >>> annot3 = result['annot3'] >>> #canvas1 = annot1.draw_on(image1.copy(), edgecolor='kitware_blue', fill=False) >>> #canvas2 = annot2.draw_on(image2.copy(), edgecolor='kitware_green', fill=False) >>> #canvas3 = annot3.draw_on(image3.copy(), edgecolor='kitware_red', fill=False) >>> canvas = kwimage.stack_images([image1, image2, image3], axis=1, pad=1, bg_value='green') >>> kwplot.imshow(canvas, pnum=pnum_(), title=ub.urepr(result['params'], nl=0, compact=1, nobr=1)) >>> kwplot.show_if_requested()

- class kwimage.Box(boxes, _check: bool = False)[source]¶
Bases:
NiceReprRepresents a single Box.
This is a convinience class. For multiple boxes use kwimage.Boxes, which is more efficient.
Currently implemented by storing a Boxes object with one item and indexing into it. This implementation could be done more efficiently.
- SeeAlso:
Example
>>> from kwimage.structs.single_box import * # NOQA >>> box = Box.random() >>> print(f'box={box}') >>> # >>> box.scale(10).quantize().to_slice() >>> # >>> sl = (slice(0, 10), slice(0, 30)) >>> box = Box.from_slice(sl) >>> print(f'box={box}')
- property format¶
- property data¶
- classmethod from_slice(slice_, shape=None, clip=True, endpoint=True, wrap=False)[source]¶
Example
>>> import kwimage >>> slice_ = kwimage.Box.random().scale(10).quantize().to_slice() >>> new = kwimage.Box.from_slice(slice_)
- property dsize¶
- intersection(other)[source]¶
Example
>>> import kwimage >>> self = kwimage.Box.coerce([0, 0, 10, 10], 'xywh') >>> other = kwimage.Box.coerce([3, 3, 10, 10], 'xywh') >>> print(str(self.intersection(other))) <Box(ltrb, [3, 3, 10, 10])>
- union_hull(other)[source]¶
Example
>>> import kwimage >>> self = kwimage.Box.coerce([0, 0, 10, 10], 'xywh') >>> other = kwimage.Box.coerce([3, 3, 10, 10], 'xywh') >>> print(str(self.union_hull(other))) <Box(ltrb, [0, 0, 13, 13])>
- to_ltrb(*args, **kwargs)[source]¶
Example
>>> import kwimage >>> self = kwimage.Box.random().to_ltrb() >>> assert self.format == 'ltrb'
- to_xywh(*args, **kwargs)[source]¶
Example
>>> import kwimage >>> self = kwimage.Box.random().to_xywh() >>> assert self.format == 'xywh'
- to_cxywh(*args, **kwargs)[source]¶
Example
>>> import kwimage >>> self = kwimage.Box.random().to_cxywh() >>> assert self.format == 'cxywh'
- corners(*args, **kwargs)[source]¶
Example
>>> import kwimage >>> assert kwimage.Box.random().corners().shape == (4, 2)
- property aspect_ratio¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().aspect_ratio)
- property center¶
Example:
>>> import kwimage >>> assert len(kwimage.Box.random().center) == 2
- property center_x¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().center_x)
- property center_y¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().center_y)
- property width¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().width)
- property height¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().height)
- property tl_x¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().tl_x)
- property tl_y¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().tl_y)
- property br_x¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().br_y)
- property br_y¶
Example:
>>> import kwimage >>> assert not ub.iterable(kwimage.Box.random().br_y)
- property dtype¶
- property area¶
- to_slice(endpoint=True)[source]¶
Example
>>> import kwimage >>> kwimage.Box.random(rng=0).scale(10).quantize().to_slice() (slice(5, 8, None), slice(5, 7, None))
- draw_on(image=None, color='blue', alpha=None, label=None, copy=False, thickness=2, label_loc='top_left')[source]¶
Draws a box directly on an image using OpenCV
Example
>>> import kwimage >>> self = kwimage.Box.random(scale=256, rng=10, format='ltrb') >>> canvas = np.zeros((256, 256, 3), dtype=np.uint8) >>> image = self.draw_on(canvas) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.figure(fnum=2000, doclf=True) >>> kwplot.autompl() >>> kwplot.imshow(image) >>> kwplot.show_if_requested()

- draw(color='blue', alpha=None, label=None, centers=False, fill=False, lw=2, ax=None, setlim=False, **kwargs)[source]¶
Draws a box directly on an image using OpenCV
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwimage >>> self = kwimage.Box.random(scale=512.0, rng=0, format='ltrb') >>> self.translate((-128, -128), inplace=True) >>> #image = (np.random.rand(256, 256) * 255).astype(np.uint8) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> #kwplot.imshow(image) >>> # xdoctest: +REQUIRES(--show) >>> self.draw(color='blue', setlim=1.2) >>> # xdoctest: +REQUIRES(--show) >>> for o in fig.findobj(): # http://matplotlib.1069221.n5.nabble.com/How-to-turn-off-all-clipping-td1813.html >>> o.set_clip_on(False) >>> kwplot.show_if_requested()
- class kwimage.Boxes(data, format=None, check=True)[source]¶
Bases:
_BoxConversionMixins,_BoxPropertyMixins,_BoxTransformMixins,_BoxDrawMixins,NiceReprConverts boxes between different formats as long as the last dimension contains 4 coordinates and the format is specified.
This is a convinience class, and should not not store the data for very long. The general idiom should be create class, convert data, and then get the raw data and let the class be garbage collected. This will help ensure that your code is portable and understandable if this class is not available.
This class is meant to efficiently store and manipulate multiple boxes. In the case of a single box the
kwimage.structs.single_box.Boxclass can be used instead.Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> import kwimage >>> import numpy as np >>> # Given an array / tensor that represents one or more boxes >>> data = np.array([[ 0, 0, 10, 10], >>> [ 5, 5, 50, 50], >>> [20, 0, 30, 10]]) >>> # The kwimage.Boxes data structure is a thin fast wrapper >>> # that provides methods for operating on the boxes. >>> # It requires that the user explicitly provide a code that denotes >>> # the format of the boxes (i.e. what each column represents) >>> boxes = kwimage.Boxes(data, 'ltrb') >>> # This means that there is no ambiguity about box format >>> # The representation string of the Boxes object demonstrates this >>> print('boxes = {!r}'.format(boxes)) boxes = <Boxes(ltrb, array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]))> >>> # if you pass this data around. You can convert to other formats >>> # For docs on available format codes see :class:`BoxFormat`. >>> # In this example we will convert (left, top, right, bottom) >>> # to (left-x, top-y, width, height). >>> boxes.toformat('xywh') <Boxes(xywh, array([[ 0, 0, 10, 10], [ 5, 5, 45, 45], [20, 0, 10, 10]]))> >>> # In addition to format conversion there are other operations >>> # We can quickly (using a C-backend) find IoUs >>> ious = boxes.ious(boxes) >>> print('{}'.format(ub.urepr(ious, nl=1, precision=2, with_dtype=False))) np.array([[1. , 0.01, 0. ], [0.01, 1. , 0.02], [0. , 0.02, 1. ]]) >>> # We can ask for the area of each box >>> print('boxes.area = {}'.format(ub.urepr(boxes.area, nl=0, with_dtype=False))) boxes.area = np.array([[ 100],[2025],[ 100]]) >>> # We can ask for the center of each box >>> print('boxes.center = {}'.format(ub.urepr(boxes.center, nl=1, with_dtype=False))) boxes.center = ( np.array([[ 5. ],[27.5],[25. ]]), np.array([[ 5. ],[27.5],[ 5. ]]), ) >>> # We can translate / scale the boxes >>> boxes.translate((10, 10)).scale(100) <Boxes(ltrb, array([[1000., 1000., 2000., 2000.], [1500., 1500., 6000., 6000.], [3000., 1000., 4000., 2000.]]))> >>> # We can clip the bounding boxes >>> boxes.translate((10, 10)).scale(100).clip(1200, 1200, 1700, 1800) <Boxes(ltrb, array([[1200., 1200., 1700., 1800.], [1500., 1500., 1700., 1800.], [1700., 1200., 1700., 1800.]]))> >>> # We can perform arbitrary warping of the boxes >>> # (note that if the transform is not axis aligned, the axis aligned >>> # bounding box of the transform result will be returned) >>> transform = np.array([[-0.83907153, 0.54402111, 0. ], >>> [-0.54402111, -0.83907153, 0. ], >>> [ 0. , 0. , 1. ]]) >>> boxes.warp(transform) <Boxes(ltrb, array([[ -8.3907153 , -13.8309264 , 5.4402111 , 0. ], [-39.23347095, -69.154632 , 23.00569785, -6.9154632 ], [-25.1721459 , -24.7113486 , -11.3412195 , -10.8804222 ]]))> >>> # Note, that we can transform the box to a Polygon for more >>> # accurate warping. >>> transform = np.array([[-0.83907153, 0.54402111, 0. ], >>> [-0.54402111, -0.83907153, 0. ], >>> [ 0. , 0. , 1. ]]) >>> warped_polys = boxes.to_polygons().warp(transform) >>> print(ub.urepr(warped_polys.data, sv=1)) [ <Polygon({ 'exterior': <Coords(data= array([[ 0. , 0. ], [ 5.4402111, -8.3907153], [ -2.9505042, -13.8309264], [ -8.3907153, -5.4402111], [ 0. , 0. ]]))>, 'interiors': [], })>, <Polygon({ 'exterior': <Coords(data= array([[ -1.4752521 , -6.9154632 ], [ 23.00569785, -44.67368205], [-14.752521 , -69.154632 ], [-39.23347095, -31.39641315], [ -1.4752521 , -6.9154632 ]]))>, 'interiors': [], })>, <Polygon({ 'exterior': <Coords(data= array([[-16.7814306, -10.8804222], [-11.3412195, -19.2711375], [-19.7319348, -24.7113486], [-25.1721459, -16.3206333], [-16.7814306, -10.8804222]]))>, 'interiors': [], })>, ] >>> # The kwimage.Boxes data structure is also convertable to >>> # several alternative data structures, like shapely, coco, and imgaug. >>> print(ub.urepr(boxes.to_shapely(), sv=1)) [ POLYGON ((0 0, 0 10, 10 10, 10 0, 0 0)), POLYGON ((5 5, 5 50, 50 50, 50 5, 5 5)), POLYGON ((20 0, 20 10, 30 10, 30 0, 20 0)), ] >>> # xdoctest: +REQUIRES(module:imgaug) >>> print(ub.urepr(boxes[0:1].to_imgaug(shape=(100, 100)), sv=1)) BoundingBoxesOnImage([BoundingBox(x1=0.0000, y1=0.0000, x2=10.0000, y2=10.0000, label=None)], shape=(100, 100)) >>> # xdoctest: -REQUIRES(module:imgaug) >>> print(ub.urepr(list(boxes.to_coco()), sv=1)) [ [0, 0, 10, 10], [5, 5, 45, 45], [20, 0, 10, 10], ] >>> # Finally, when you are done with your boxes object, you can >>> # unwrap the raw data by using the ``.data`` attribute >>> # all operations are done on this data, which gives the >>> # kwiamge.Boxes data structure almost no overhead when >>> # inserted into existing code. >>> print('boxes.data =\n{}'.format(ub.urepr(boxes.data, nl=1))) boxes.data = np.array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]], dtype=...) >>> # xdoctest: +REQUIRES(module:torch) >>> # This data structure was designed for use with both torch >>> # and numpy, the underlying data can be either an array or tensor. >>> boxes.tensor() <Boxes(ltrb, tensor([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]...))> >>> boxes.numpy() <Boxes(ltrb, array([[ 0, 0, 10, 10], [ 5, 5, 50, 50], [20, 0, 30, 10]]))>
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> # Demo of conversion methods >>> import kwimage >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh') <Boxes(xywh, array([[25, 30, 15, 10]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_xywh() <Boxes(xywh, array([[25, 30, 15, 10]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_cxywh() <Boxes(cxywh, array([[32.5, 35. , 15. , 10. ]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').to_ltrb() <Boxes(ltrb, array([[25, 30, 40, 40]]))> >>> kwimage.Boxes([[25, 30, 15, 10]], 'xywh').scale(2).to_ltrb() <Boxes(ltrb, array([[50., 60., 80., 80.]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> kwimage.Boxes(torch.FloatTensor([[25, 30, 15, 20]]), 'xywh').scale(.1).to_ltrb() <Boxes(ltrb, tensor([[ 2.5000, 3.0000, 4.0000, 5.0000]]))>
Note
In the following examples we show cases where
Boxescan hold a single 1-dimensional box array. This is a holdover from an older codebase, and some functions may assume that the input is at least 2-D. Thus when representing a single bounding box it is best practice to view it as a list of 1 box. While many function will work in the 1-D case, not all functions have been tested and thus we cannot gaurentee correctness.Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes([25, 30, 15, 10], 'xywh') <Boxes(xywh, array([25, 30, 15, 10]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_xywh() <Boxes(xywh, array([25, 30, 15, 10]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_cxywh() <Boxes(cxywh, array([32.5, 35. , 15. , 10. ]))> >>> Boxes([25, 30, 15, 10], 'xywh').to_ltrb() <Boxes(ltrb, array([25, 30, 40, 40]))> >>> Boxes([25, 30, 15, 10], 'xywh').scale(2).to_ltrb() <Boxes(ltrb, array([50., 60., 80., 80.]))> >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> Boxes(torch.FloatTensor([[25, 30, 15, 20]]), 'xywh').scale(.1).to_ltrb() <Boxes(ltrb, tensor([[ 2.5000, 3.0000, 4.0000, 5.0000]]))>
Example
>>> datas = [ >>> [1, 2, 3, 4], >>> [[1, 2, 3, 4], [4, 5, 6, 7]], >>> [[[1, 2, 3, 4], [4, 5, 6, 7]]], >>> ] >>> formats = BoxFormat.cannonical >>> for format1 in formats: >>> for data in datas: >>> self = box1 = Boxes(data, format1) >>> for format2 in formats: >>> box2 = box1.toformat(format2) >>> back = box2.toformat(format1) >>> assert box1 == back
- Parameters:
data (ndarray | Tensor | Boxes) – Either an ndarray or Tensor with trailing shape of 4, or an existing Boxes object.
format (str) – format code indicating which coordinates are represented by data. If data is a Boxes object then this is not necessary.
check (bool) – if True runs input checks on raw data.
- Raises:
ValueError – if data is specified without a format
- classmethod random(num=1, scale=1.0, format='xywh', anchors=None, anchor_std=0.16666666666666666, tensor=False, rng=None)[source]¶
Makes random boxes; typically for testing purposes
- Parameters:
num (int) – number of boxes to generate
scale (float | Tuple[float, float]) – size of imgdims
format (str) – format of boxes to be created (e.g. ltrb, xywh)
anchors (ndarray) – normalized width / heights of anchor boxes to perterb and randomly place. (must be in range 0-1)
anchor_std (float) – magnitude of noise applied to anchor shapes
tensor (bool) – if True, returns boxes in tensor format
rng (None | int | RandomState) – initial random seed
- Returns:
random boxes
- Return type:
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes.random(3, rng=0, scale=100) <Boxes(xywh, array([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]]...))> >>> # xdoctest: +REQUIRES(module:torch) >>> Boxes.random(3, rng=0, scale=100).tensor() <Boxes(xywh, tensor([[ 54, 54, 6, 17], [ 42, 64, 1, 25], [ 79, 38, 17, 14]]...))> >>> anchors = np.array([[.5, .5], [.3, .3]]) >>> Boxes.random(3, rng=0, scale=100, anchors=anchors) <Boxes(xywh, array([[ 2, 13, 51, 51], [32, 51, 32, 36], [36, 28, 23, 26]]...))>
Example
>>> # Boxes position/shape within 0-1 space should be uniform. >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> fig.gca().set_xlim(0, 128) >>> fig.gca().set_ylim(0, 128) >>> import kwimage >>> kwimage.Boxes.random(num=10).scale(128).draw()
- classmethod concatenate(boxes, axis=0)[source]¶
Concatenates multiple boxes together
- Parameters:
boxes (Sequence[Boxes]) – list of boxes to concatenate
axis (int) – axis to stack on. Defaults to 0.
- Returns:
stacked boxes
- Return type:
Example
>>> boxes = [Boxes.random(3) for _ in range(3)] >>> new = Boxes.concatenate(boxes) >>> assert len(new) == 9 >>> assert np.all(new.data[3:6] == boxes[1].data)
Example
>>> boxes = [Boxes.random(3) for _ in range(3)] >>> boxes[0].data = boxes[0].data[0] >>> boxes[1].data = boxes[0].data[0:0] >>> new = Boxes.concatenate(boxes) >>> assert len(new) == 4 >>> # xdoctest: +REQUIRES(module:torch) >>> new = Boxes.concatenate([b.tensor() for b in boxes]) >>> assert len(new) == 4
- compress(flags, axis=0, inplace=False)[source]¶
Filters boxes based on a boolean criterion
- Parameters:
flags (ArrayLike) – true for items to be kept. Extended type: ArrayLike[bool]
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
the boxes corresponding to where flags were true
- Return type:
Example
>>> self = Boxes([[25, 30, 15, 10]], 'ltrb') >>> self.compress([True]) <Boxes(ltrb, array([[25, 30, 15, 10]]))> >>> self.compress([False]) <Boxes(ltrb, array([], shape=(0, 4), dtype=...))>
- take(idxs, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
- Parameters:
indices (ArrayLike) – Indexes of items to take. Extended type ArrayLike[int].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
the boxes corresponding to the specified indices
- Return type:
Example
>>> self = Boxes([[25, 30, 15, 10]], 'ltrb') >>> self.take([0]) <Boxes(ltrb, array([[25, 30, 15, 10]]))> >>> self.take([]) <Boxes(ltrb, array([], shape=(0, 4), dtype=...))>
- is_tensor()[source]¶
is the backend fueled by torch?
- Returns:
True if the Boxes are torch tensors
- Return type:
- is_numpy()[source]¶
is the backend fueled by numpy?
- Returns:
True if the Boxes are numpy arrays
- Return type:
- property _impl¶
returns the kwarray.ArrayAPI implementation for the data
- Returns:
the array API for the box backend
- Return type:
Example
>>> assert Boxes.random().numpy()._impl.is_numpy >>> # xdoctest: +REQUIRES(module:torch) >>> assert Boxes.random().tensor()._impl.is_tensor
- property device¶
If the backend is torch returns the data device, otherwise None
- astype(dtype)[source]¶
Changes the type of the internal array used to represent the boxes
Note
this operation is not inplace
- Returns:
the boxes with the chosen type
- Return type:
Example
>>> # xdoctest: +IGNORE_WHITESPACE >>> # xdoctest: +REQUIRES(module:torch) >>> Boxes.random(3, 100, rng=0).tensor().astype('int16') <Boxes(xywh, tensor([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]], dtype=torch.int16))> >>> Boxes.random(3, 100, rng=0).numpy().astype('int16') <Boxes(xywh, array([[54, 54, 6, 17], [42, 64, 1, 25], [79, 38, 17, 14]], dtype=int16))> >>> Boxes.random(3, 100, rng=0).tensor().astype('float32') >>> Boxes.random(3, 100, rng=0).numpy().astype('float32')
- round(inplace=False)[source]¶
Rounds data coordinates to the nearest integer.
This operation is applied directly to the box coordinates, so its output will depend on the format the boxes are stored in.
- Parameters:
inplace (bool) – if True, modifies this object. Defaults to False.
- Returns:
the boxes with rounded coordinates
- Return type:
- SeeAlso:
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0).scale(10) >>> new = self.round() >>> print('self = {!r}'.format(self)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[5.48813522, 5.44883192, 0.53949833, 1.70306146], [4.23654795, 6.4589411 , 0.13932407, 2.45878875], [7.91725039, 3.83441508, 1.71937704, 1.45453393]]))> new = <Boxes(xywh, array([[5., 5., 1., 2.], [4., 6., 0., 2.], [8., 4., 2., 1.]]))>
- quantize(inplace=False, dtype=<class 'numpy.int32'>)[source]¶
Converts the box to integer coordinates.
This operation takes the floor of the left side and the ceil of the right side. Thus the area of the box will never decreases. But this will often increase the width / height of the box by a pixel.
- Parameters:
inplace (bool) – if True, modifies this object
dtype (type) – type to cast as
- Returns:
the boxes with quantized coordinates
- Return type:
- SeeAlso:
Boxes.round()Boxes.resize()if you need to ensure the size does not change
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0).scale(10) >>> new = self.quantize() >>> print('self = {!r}'.format(self)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[5.48813522, 5.44883192, 0.53949833, 1.70306146], [4.23654795, 6.4589411 , 0.13932407, 2.45878875], [7.91725039, 3.83441508, 1.71937704, 1.45453393]]))> new = <Boxes(xywh, array([[5, 5, 2, 3], [4, 6, 1, 3], [7, 3, 3, 3]]...))>
Example
>>> import kwimage >>> # Be careful if it is important to preserve the width/height >>> self = kwimage.Boxes([[0, 0, 10, 10]], 'xywh') >>> aff = kwimage.Affine.coerce(offset=(0.5, 0.0)) >>> warped = self.warp(aff) >>> new = warped.quantize(dtype=int) >>> print('self = {!r}'.format(self)) >>> print('warped = {!r}'.format(warped)) >>> print('new = {!r}'.format(new)) self = <Boxes(xywh, array([[ 0, 0, 10, 10]]))> warped = <Boxes(xywh, array([[ 0.5, 0. , 10. , 10. ]]))> new = <Boxes(xywh, array([[ 0, 0, 11, 10]]))>
Example
>>> import kwimage >>> self = kwimage.Boxes.random(3, rng=0) >>> orig = self.copy() >>> self.quantize(inplace=True) >>> assert np.any(self.data != orig.data)
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
- Returns:
the boxes with a numpy backend
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(3).tensor() >>> newself = self.numpy() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
- Parameters:
device (int | None | torch.device) – The torch device to put the backend tensors on
- Returns:
the boxes with a torch backend
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(3) >>> # xdoctest: +REQUIRES(module:torch) >>> newself = self.tensor() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- ious(other, bias=0, impl='auto', mode=None)[source]¶
Intersection over union.
Compute IOUs (intersection area over union area) between these boxes and another set of boxes. This is a symmetric measure of similarity between boxes.
Todo
- [ ] Add pairwise flag to toggle between one-vs-one and all-vs-all
computation. I.E. Add option for componentwise calculation.
- Parameters:
other (Boxes) – boxes to compare IoUs against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
impl (str) – code to specify implementation used to ious. Can be either torch, py, c, or auto. Efficiency and the exact result will vary by implementation, but they will always be close. Some implementations only accept certain data types (e.g. impl=’c’, only accepts float32 numpy arrays). See ~/code/kwimage/dev/bench_bbox.py for benchmark details. On my system the torch impl was fastest (when the data was on the GPU). Defaults to ‘auto’
mode (str) – depricated, use impl
- Returns:
the ious
- Return type:
ndarray
- SeeAlso:
iooas - for a measure of coverage between boxes
Examples
>>> import kwimage >>> self = kwimage.Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = kwimage.Boxes(np.array([6, 2, 20, 10]), 'ltrb') >>> overlaps = self.ious(other, bias=1).round(2) >>> assert np.all(np.isclose(overlaps, [0.21, 0.63, 0.04])), repr(overlaps)
Examples
>>> import kwimage >>> boxes1 = kwimage.Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = kwimage.Boxes(np.array([[6, 2, 20, 10], >>> [100, 200, 300, 300]]), 'ltrb') >>> overlaps = boxes1.ious(other) >>> print('{}'.format(ub.urepr(overlaps, precision=2, nl=1))) np.array([[0.18, 0. ], [0.61, 0. ], [0. , 0. ]]...)
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> Boxes(np.empty(0), 'xywh').ious(Boxes(np.empty(4), 'xywh')).shape (0,) >>> #Boxes(np.empty(4), 'xywh').ious(Boxes(np.empty(0), 'xywh')).shape >>> Boxes(np.empty((0, 4)), 'xywh').ious(Boxes(np.empty((0, 4)), 'xywh')).shape (0, 0) >>> Boxes(np.empty((1, 4)), 'xywh').ious(Boxes(np.empty((0, 4)), 'xywh')).shape (1, 0) >>> Boxes(np.empty((0, 4)), 'xywh').ious(Boxes(np.empty((1, 4)), 'xywh')).shape (0, 1)
Examples
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> formats = BoxFormat.cannonical >>> istensors = [False, True] >>> results = {} >>> for format in formats: >>> for tensor in istensors: >>> boxes1 = Boxes.random(5, scale=10.0, rng=0, format=format, tensor=tensor) >>> boxes2 = Boxes.random(7, scale=10.0, rng=1, format=format, tensor=tensor) >>> ious = boxes1.ious(boxes2) >>> results[(format, tensor)] = ious >>> results = {k: v.numpy() if torch.is_tensor(v) else v for k, v in results.items() } >>> results = {k: v.tolist() for k, v in results.items()} >>> print(ub.urepr(results, sk=True, precision=3, nl=2)) >>> from functools import partial >>> assert ub.allsame(results.values(), partial(np.allclose, atol=1e-07))
- iooas(other, bias=0)[source]¶
Intersection over other area.
This is an asymetric measure of coverage. How much of the “other” boxes are covered by these boxes. It is the area of intersection between each pair of boxes and the area of the “other” boxes.
- SeeAlso:
ious - for a measure of similarity between boxes
- Parameters:
other (Boxes) – boxes to compare IoOA against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
- Returns:
the iooas
- Return type:
ndarray
Examples
>>> self = Boxes(np.array([[ 0, 0, 10, 10], >>> [10, 0, 20, 10], >>> [20, 0, 30, 10]]), 'ltrb') >>> other = Boxes(np.array([[6, 2, 20, 10], [0, 0, 0, 3]]), 'xywh') >>> coverage = self.iooas(other, bias=0).round(2) >>> print('coverage = {!r}'.format(coverage))
- isect_area(other, bias=0)[source]¶
Intersection part of intersection over union computation
- Parameters:
other (Boxes) – boxes to compare IoOA against
bias (int) – either 0 or 1, does TL=BR have area of 0 or 1? Defaults to 0.
- Returns:
the iooas
- Return type:
ndarray
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> self = Boxes.random(5, scale=10.0, rng=0, format='ltrb') >>> other = Boxes.random(3, scale=10.0, rng=1, format='ltrb') >>> isect = self.isect_area(other, bias=0) >>> ious_v1 = isect / ((self.area + other.area.T) - isect) >>> ious_v2 = self.ious(other, bias=0) >>> assert np.allclose(ious_v1, ious_v2)
- intersection(other)[source]¶
Componentwise intersection between two sets of Boxes
intersections of boxes are always boxes, so this works
- Parameters:
other (Boxes) – boxes to intersect with this object. (must be of same length)
- Returns:
the component-wise intersection geometry
- Return type:
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.intersection(other) >>> new_area = np.nan_to_num(new.area).ravel() >>> alt_area = np.diag(self.isect_area(other)) >>> close = np.isclose(new_area, alt_area) >>> assert np.all(close)
- union_hull(other)[source]¶
Componentwise bounds of the union between two sets of Boxes
NOTE: convert to polygon to do a real union.
Note
This is not a real hull. A better name for this might be union_bounds because we are returning the axis-aligned bounds of the convex hull of the union.
- Parameters:
other (Boxes) – boxes to union with this object. (must be of same length)
- Returns:
bounding box of the unioned boxes
- Return type:
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.union_hull(other) >>> new_area = np.nan_to_num(new.area).ravel()
- bounding_box()[source]¶
Returns the box that bounds all of the contained boxes
- Returns:
a single box
- Return type:
Examples
>>> # xdoctest: +IGNORE_WHITESPACE >>> from kwimage.structs.boxes import * # NOQA >>> self = Boxes.random(5, rng=0).scale(10.) >>> other = self.translate(1) >>> new = self.union_hull(other) >>> new_area = np.nan_to_num(new.area).ravel()
- contains(other)[source]¶
Determine of points are completely contained by these boxes
- Parameters:
other (kwimage.Points) – points to test for containment. TODO: support generic data types
- Returns:
- flags - N x M boolean matrix indicating which box
contains which points, where N is the number of boxes and M is the number of points.
- Return type:
ArrayLike
Examples
>>> import kwimage >>> self = kwimage.Boxes.random(10).scale(10).round() >>> other = kwimage.Points.random(10).scale(10).round() >>> flags = self.contains(other) >>> flags = self.contains(self.xy_center) >>> assert np.all(np.diag(flags))
- view(*shape)[source]¶
Passthrough method to view or reshape
- Parameters:
*shape (Tuple[int, …]) – new shape
- Returns:
data with a different view
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Boxes.random(6, scale=10.0, rng=0, format='xywh').tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4] >>> self = Boxes.random(6, scale=10.0, rng=0, format='ltrb').tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4]
- _ensure_nonnegative_extent(inplace=False)[source]¶
Experimental. If the box has a negative width / height make them positive and adjust the tlxy point.
Need a better name for this function.
- FIXME:
- [ ] Slice semantics (i.e. start/stop) of boxes break under
rotations and reflections. This function needs to be thought out a bit more before becoming non-experimental.
- Returns:
Boxes
Example
>>> import kwimage >>> self = kwimage.Boxes(np.array([ >>> [20, 30, -10, -20], >>> [0, 0, 10, 20], >>> [0, 0, -10, 20], >>> [0, 0, 10, -20], >>> ]), 'xywh') >>> new = self._ensure_nonnegative_extent(inplace=0) >>> assert np.any(self.width < 0) >>> assert not np.any(new.width < 0) >>> assert np.any(self.height < 0) >>> assert not np.any(new.height < 0)
>>> import kwimage >>> self = kwimage.Boxes(np.array([ >>> [0, 3, 8, -4], >>> ]), 'xywh') >>> new = self._ensure_nonnegative_extent(inplace=0) >>> print('self = {}'.format(ub.urepr(self, nl=1))) >>> print('new = {}'.format(ub.urepr(new, nl=1))) >>> assert not np.any(self.width < 0) >>> assert not np.any(new.width < 0) >>> assert np.any(self.height < 0) >>> assert not np.any(new.height < 0)
- class kwimage.Color(color, alpha=None, space=None, coerce=True)[source]¶
Bases:
NiceReprUsed for converting a single color between spaces and encodings. This should only be used when handling small numbers of colors(e.g. 1), don’t use this to represent an image.
- Parameters:
space (str) – colorspace of wrapped color. Assume RGB if not specified and it cannot be inferred
CommandLine
xdoctest -m ~/code/kwimage/kwimage/im_color.py Color
Example
>>> print(Color('g')) >>> print(Color('orangered')) >>> print(Color('#AAAAAA').as255()) >>> print(Color([0, 255, 0])) >>> print(Color([1, 1, 1.])) >>> print(Color([1, 1, 1])) >>> print(Color(Color([1, 1, 1])).as255()) >>> print(Color(Color([1., 0, 1, 0])).ashex()) >>> print(Color([1, 1, 1], alpha=255)) >>> print(Color([1, 1, 1], alpha=255, space='lab'))
- Parameters:
color (Color | Iterable[int | float] | str) – something coercable into a color
alpha (float | None) – if psecified adds an alpha value
space (str) – The colorspace to interpret this color as. Defaults to rgb.
coerce (bool) – The exsting init is not lightweight. This is a design problem that will need to be fixed in future versions. Setting coerce=False will disable all magic and use imputed color and space args directly. Alpha will be ignored.
- forimage(image, space='auto')[source]¶
Return a numeric value for this color that can be used in the given image.
Create a numeric color tuple that agrees with the format of the input image (i.e. float or int, with 3 or 4 channels).
- Parameters:
image (ndarray) – image to return color for
space (str) – colorspace of the input image. Defaults to ‘auto’, which will choose rgb or rgba
- Returns:
the color value
- Return type:
Tuple[Number, …]
Example
>>> import kwimage >>> img_f3 = np.zeros([8, 8, 3], dtype=np.float32) >>> img_u3 = np.zeros([8, 8, 3], dtype=np.uint8) >>> img_f4 = np.zeros([8, 8, 4], dtype=np.float32) >>> img_u4 = np.zeros([8, 8, 4], dtype=np.uint8) >>> kwimage.Color('red').forimage(img_f3) (1.0, 0.0, 0.0) >>> kwimage.Color('red').forimage(img_f4) (1.0, 0.0, 0.0, 1.0) >>> kwimage.Color('red').forimage(img_u3) (255, 0, 0) >>> kwimage.Color('red').forimage(img_u4) (255, 0, 0, 255) >>> kwimage.Color('red', alpha=0.5).forimage(img_f4) (1.0, 0.0, 0.0, 0.5) >>> kwimage.Color('red', alpha=0.5).forimage(img_u4) (255, 0, 0, 127) >>> kwimage.Color('red').forimage(np.uint8) (255, 0, 0)
- ashex(space=None)[source]¶
Convert to hex values
- Parameters:
space (None | str) – if specified convert to this colorspace before returning
- Returns:
the hex representation
- Return type:
- as01(space=None)[source]¶
Convert to float values
- Parameters:
space (None | str) – if specified convert to this colorspace before returning
- Returns:
The float tuple of color values between 0 and 1
- Return type:
Tuple[float, float, float] | Tuple[float, float, float, float]
Note
This function is only guarenteed to return 0-1 values for rgb values. For HSV and LAB, the native spaces are used. This is not ideal, and we may create a new function that fixes this - at least conceptually - and deprate this for that in the future.
For HSV, H is between 0 and 360. S, and V are in [0, 1]
- classmethod _is_base255(channels)[source]¶
there is a one corner case where all pixels are 1 or less
- classmethod named_colors()[source]¶
- Returns:
names of colors that Color accepts
- Return type:
List[str]
Example
>>> import kwimage >>> named_colors = kwimage.Color.named_colors() >>> color_lut = {name: kwimage.Color(name).as01() for name in named_colors} >>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> # This is a very big table if we let it be, reduce it >>> color_lut =dict(list(color_lut.items())[0:10]) >>> canvas = kwplot.make_legend_img(color_lut) >>> kwplot.imshow(canvas)

- classmethod distinct(num, existing=None, space='rgb', legacy='auto', exclude_black=True, exclude_white=True)[source]¶
Make multiple distinct colors.
The legacy variant is based on a stack overflow post [HowToDistinct], but the modern variant is based on the
distinctipypackage.References
[HowToDistinct]https://stackoverflow.com/questions/470690/how-to-automatically-generate-n-distinct-colors
Todo
- [ ] If num is more than a threshold we should switch to
a different strategy to generating colors that just samples uniformly from some colormap and then shuffles. We have no hope of making things distinguishable when num starts going over 10 or so. See [ColorLimits] [WikiDistinguish] [Disinct2] for more ideas.
- Returns:
list of distinct float color values
- Return type:
List[Tuple]
Example
>>> # xdoctest: +REQUIRES(module:matplotlib) >>> from kwimage.im_color import * # NOQA >>> import kwimage >>> colors1 = kwimage.Color.distinct(5, legacy=False) >>> colors2 = kwimage.Color.distinct(3, existing=colors1) >>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> from kwimage.im_color import _draw_color_swatch >>> swatch1 = _draw_color_swatch(colors1, cellshape=9) >>> swatch2 = _draw_color_swatch(colors1 + colors2, cellshape=9) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(swatch1, pnum=(1, 2, 1), fnum=1) >>> kwplot.imshow(swatch2, pnum=(1, 2, 2), fnum=1) >>> kwplot.show_if_requested()
- distance(other, space='lab')[source]¶
Distance between self an another color
- Parameters:
other (Color) – the color to compare
space (str) – the colorspace to comapre in
- Returns:
float
- interpolate(other, alpha=0.5, ispace=None, ospace=None)[source]¶
Interpolate between colors
- Parameters:
other (Color) – A coercable Color
alpha (float | List[float]) – one or more interpolation values
ispace (str | None) – colorspace to interpolate in
ospace (str | None) – colorspace of returned color
- Returns:
Color | List[Color]
Example
>>> import kwimage >>> color1 = self = kwimage.Color.coerce('orangered') >>> color2 = other = kwimage.Color.coerce('dodgerblue') >>> alpha = np.linspace(0, 1, 6) >>> ispace = 'rgb' >>> ospace = 'rgb' >>> colorBs = self.interpolate(other, alpha, ispace=ispace, ospace=ospace) >>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> from kwimage.im_color import _draw_color_swatch >>> swatch_colors = [color1] + colorBs + [color2] >>> print('swatch_colors = {}'.format(ub.urepr(swatch_colors, nl=1))) >>> swatch1 = _draw_color_swatch(swatch_colors, cellshape=(8, 8)) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(swatch1, pnum=(1, 1, 1), fnum=1) >>> kwplot.show_if_requested()
- to_image(dsize=(8, 8))[source]¶
Create an solid-color image with this color
- Parameters:
dsize (Tuple[int, int]) – the desired width / height of the image (defaults to 8x8)
- adjust(saturate=0, lighten=0)[source]¶
Adjust the saturation or value of a color.
Requires that
colormathis installed.- Parameters:
saturate (float) – between +1 and -1, when positive saturates the color, when negative desaturates the color.
lighten (float) – between +1 and -1, when positive lightens the color, when negative darkens the color.
Example
>>> # xdoctest: +REQUIRES(module:colormath) >>> import kwimage >>> self = kwimage.Color.coerce('salmon') >>> new = self.adjust(saturate=+0.2) >>> cell1 = self.to_image() >>> cell2 = new.to_image() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> canvas = kwimage.stack_images([cell1, cell2], axis=1) >>> kwplot.imshow(canvas)
Example
>>> # xdoctest: +REQUIRES(module:colormath) >>> import kwimage >>> self = kwimage.Color.coerce('salmon', alpha=0.5) >>> new = self.adjust(saturate=+0.2) >>> cell1 = self.to_image() >>> cell2 = new.to_image() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> canvas = kwimage.stack_images([cell1, cell2], axis=1) >>> kwplot.imshow(canvas)
Example
>>> # xdoctest: +REQUIRES(module:colormath) >>> import kwimage >>> adjustments = [ >>> {'saturate': -0.2}, >>> {'saturate': +0.2}, >>> {'lighten': +0.2}, >>> {'lighten': -0.2}, >>> {'saturate': -0.9}, >>> {'saturate': +0.9}, >>> {'lighten': +0.9}, >>> {'lighten': -0.9}, >>> ] >>> self = kwimage.Color.coerce('kitware_green') >>> dsize = (256, 64) >>> to_show = [] >>> to_show.append(self.to_image(dsize)) >>> for kwargs in adjustments: >>> new = self.adjust(**kwargs) >>> cell = new.to_image(dsize=dsize) >>> text = ub.urepr(kwargs, compact=1, nobr=1) >>> cell, info = kwimage.draw_text_on_image(cell, text, return_info=1, border={'thickness': 2}, color='white', fontScale=1.0) >>> to_show.append(cell) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> canvas = kwimage.stack_images_grid(to_show) >>> canvas = kwimage.draw_header_text(canvas, 'kwimage.Color.adjust') >>> kwplot.imshow(canvas)

- class kwimage.Coords(data=None, meta=None)[source]¶
-
A data structure to store n-dimensional coordinate geometry.
Currently it is up to the user to maintain what coordinate system this geometry belongs to.
Note
This class was designed to hold coordinates in r/c format, but in general this class is anostic to dimension ordering as long as you are consistent. However, there are two places where this matters: (1) drawing and (2) gdal/imgaug-warping. In these places we will assume x/y for legacy reasons. This may change in the future.
The term axes with resepct to
Coordsalways refers to the final numpy axis. In other words the final numpy-axis represents ALL of the coordinate-axes.CommandLine
xdoctest -m kwimage.structs.coords Coords
Example
>>> from kwimage.structs.coords import * # NOQA >>> import kwarray >>> rng = kwarray.ensure_rng(0) >>> self = Coords.random(num=4, dim=3, rng=rng) >>> print('self = {}'.format(self)) self = <Coords(data= array([[0.5488135 , 0.71518937, 0.60276338], [0.54488318, 0.4236548 , 0.64589411], [0.43758721, 0.891773 , 0.96366276], [0.38344152, 0.79172504, 0.52889492]]))> >>> matrix = rng.rand(4, 4) >>> self.warp(matrix) <Coords(data= array([[0.71037426, 1.25229659, 1.39498435], [0.60799503, 1.26483447, 1.42073131], [0.72106004, 1.39057144, 1.38757508], [0.68384299, 1.23914654, 1.29258196]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[3.5488135 , 3.71518937, 3.60276338], [3.54488318, 3.4236548 , 3.64589411], [3.43758721, 3.891773 , 3.96366276], [3.38344152, 3.79172504, 3.52889492]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[6.5488135 , 6.71518937, 6.60276338], [6.54488318, 6.4236548 , 6.64589411], [6.43758721, 6.891773 , 6.96366276], [6.38344152, 6.79172504, 6.52889492]]))> >>> self.scale(2) <Coords(data= array([[13.09762701, 13.43037873, 13.20552675], [13.08976637, 12.8473096 , 13.29178823], [12.87517442, 13.783546 , 13.92732552], [12.76688304, 13.58345008, 13.05778984]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> self.tensor() >>> self.tensor().tensor().numpy().numpy() >>> self.numpy() >>> #self.draw_on()
- property dtype¶
- property dim¶
- property shape¶
- classmethod random(num=1, dim=2, rng=None, meta=None)[source]¶
Makes random coordinates; typically for testing purposes
- compress(flags, axis=0, inplace=False)[source]¶
Filters items based on a boolean criterion
- Parameters:
flags (ArrayLike) – true for items to be kept. Extended type: ArrayLike[bool].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
filtered coords
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords.random(10, rng=0) >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self)) <Coords(data=array([], shape=(0, 2), dtype=float64))> >>> # xdoctest: +REQUIRES(module:torch) >>> self = self.tensor() >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self))
- take(indices, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
- Parameters:
indices (ArrayLike) – indexes of items to take. Extended type ArrayLike[int].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
filtered coords
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords(np.array([[25, 30, 15, 10]])) >>> self.take([0]) <Coords(data=array([[25, 30, 15, 10]]))> >>> self.take([]) <Coords(data=array([], shape=(0, 4), dtype=...))>
- astype(dtype, inplace=False)[source]¶
Changes the data type
- Parameters:
dtype – new type
inplace (bool) – if True, modifies this object
- Returns:
modified coordinates
- Return type:
- round(decimals=0, inplace=False)[source]¶
Rounds data to the specified decimal place
- Parameters:
inplace (bool) – if True, modifies this object
decimals (int) – number of decimal places to round to
- Returns:
modified coordinates
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords.random(3).scale(10) >>> self.round()
- view(*shape)[source]¶
Passthrough method to view or reshape
- Parameters:
*shape – new shape of the data
- Returns:
modified coordinates
- Return type:
Example
>>> self = Coords.random(6, dim=4).numpy() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4] >>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(6, dim=4).tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4]
- classmethod concatenate(coords, axis=0)[source]¶
Concatenates lists of coordinates together
- Parameters:
coords (Sequence[Coords]) – list of coords to concatenate
axis (int) – axis to stack on. Defaults to 0.
- Returns:
stacked coords
- Return type:
CommandLine
xdoctest -m kwimage.structs.coords Coords.concatenate
Example
>>> coords = [Coords.random(3) for _ in range(3)] >>> new = Coords.concatenate(coords) >>> assert len(new) == 9 >>> assert np.all(new.data[3:6] == coords[1].data)
- property device¶
If the backend is torch returns the data device, otherwise None
- property _impl¶
Returns the internal tensor/numpy ArrayAPI implementation
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
- Returns:
modified coordinates
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).numpy() >>> newself = self.tensor() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
- Returns:
modified coordinates
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).tensor() >>> newself = self.numpy() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- reorder_axes(new_order, inplace=False)[source]¶
Change the ordering of the coordinate axes.
- Parameters:
new_order (Tuple[int]) –
new_order[i]should specify which axes in the original coordinates should be mapped to thei-thposition in the returned axes.inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Note
This is the ordering of the “columns” in final numpy axis, not the numpy axes themselves.
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords(data=np.array([ >>> [7, 11], >>> [13, 17], >>> [21, 23], >>> ])) >>> new = self.reorder_axes((1, 0)) >>> print('new = {!r}'.format(new)) new = <Coords(data= array([[11, 7], [17, 13], [23, 21]]))>
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.reorder_axes((1, 0)) >>> # Remapping using 1, 0 reverses the axes >>> assert np.all(new.data[:, 0] == self.data[:, 1]) >>> assert np.all(new.data[:, 1] == self.data[:, 0]) >>> # Remapping using 0, 1 does nothing >>> eye = self.reorder_axes((0, 1)) >>> assert np.all(eye.data == self.data) >>> # Remapping using 0, 0, destroys the 1-th column >>> bad = self.reorder_axes((0, 0)) >>> assert np.all(bad.data[:, 0] == self.data[:, 0]) >>> assert np.all(bad.data[:, 1] == self.data[:, 0])
- warp(transform, input_dims=None, output_dims=None, inplace=False)[source]¶
Generalized coordinate transform.
- Parameters:
transform (SKImageGeometricTransform | ArrayLike | Augmenter | Callable) – scikit-image tranform, a 3x3 transformation matrix, an imgaug Augmenter, or generic callable which transforms an NxD ndarray.
input_dims (Tuple) – shape of the image these objects correspond to (only needed / used when transform is an imgaug augmenter)
output_dims (Tuple) – unused in non-raster structures, only exists for compatibility.
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Note
Let D = self.dims
- transformation matrices can be either:
(D + 1) x (D + 1) # for homog
D x D # for scale / rotate
D x (D + 1) # for affine
Example
>>> from kwimage.structs.coords import * # NOQA >>> import skimage >>> self = Coords.random(10, rng=0) >>> transform = skimage.transform.AffineTransform(scale=(2, 2)) >>> new = self.warp(transform) >>> assert np.all(new.data == self.scale(2).data)
Doctest
>>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(np.eye(3)).data == self.data) >>> assert np.all(self.warp(np.eye(2)).data == self.data)
Doctest
>>> # xdoctest: +REQUIRES(module:osgeo) >>> from osgeo import osr >>> wgs84_crs = osr.SpatialReference() >>> wgs84_crs.ImportFromEPSG(4326) >>> dst_crs = osr.SpatialReference() >>> dst_crs.ImportFromEPSG(2927) >>> transform = osr.CoordinateTransformation(wgs84_crs, dst_crs) >>> self = Coords.random(10, rng=0) >>> new = self.warp(transform) >>> assert np.all(new.data != self.data)
>>> # Alternative using generic func >>> def _gdal_coord_tranform(pts): ... return np.array([transform.TransformPoint(x, y, 0)[0:2] ... for x, y in pts]) >>> alt = self.warp(_gdal_coord_tranform) >>> assert np.all(alt.data != self.data) >>> assert np.all(alt.data == new.data)
Doctest
>>> # can use a generic function >>> def func(xy): ... return np.zeros_like(xy) >>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(func).data == 0)
- _warp_imgaug(augmenter, input_dims, inplace=False)[source]¶
Warps by applying an augmenter from the imgaug library
Note
We are assuming you are using X/Y coordinates here.
- Parameters:
augmenter (imgaug.augmenters.Augmenter)
input_dims (Tuple) – h/w of the input image
inplace (bool) – if True, modifies data inplace
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/coords.py Coords._warp_imgaug
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> from kwimage.structs.coords import * # NOQA >>> import imgaug >>> input_dims = (10, 10) >>> self = Coords.random(10).scale(input_dims) >>> augmenter = imgaug.augmenters.Fliplr(p=1) >>> new = self._warp_imgaug(augmenter, input_dims) >>> # y coordinate should not change >>> assert np.allclose(self.data[:, 1], new.data[:, 1]) >>> assert np.allclose(input_dims[0] - self.data[:, 0], new.data[:, 0])
>>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> from matplotlib import pyplot as pl >>> ax = plt.gca() >>> ax.set_xlim(0, input_dims[0]) >>> ax.set_ylim(0, input_dims[1]) >>> self.draw(color='red', alpha=.4, radius=0.1) >>> new.draw(color='blue', alpha=.4, radius=0.1)
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> from kwimage.structs.coords import * # NOQA >>> import imgaug >>> input_dims = (32, 32) >>> inplace = 0 >>> self = Coords.random(1000, rng=142).scale(input_dims).scale(.8) >>> self.data = self.data.astype(np.int32).astype(np.float32) >>> augmenter = imgaug.augmenters.CropAndPad(px=(-4, 4), keep_size=1).to_deterministic() >>> new = self._warp_imgaug(augmenter, input_dims) >>> # Change should be linear >>> norm1 = (self.data - self.data.min(axis=0)) / (self.data.max(axis=0) - self.data.min(axis=0)) >>> norm2 = (new.data - new.data.min(axis=0)) / (new.data.max(axis=0) - new.data.min(axis=0)) >>> diff = norm1 - norm2 >>> assert np.allclose(diff, 0, atol=1e-6, rtol=1e-4) >>> #assert np.allclose(self.data[:, 1], new.data[:, 1]) >>> #assert np.allclose(input_dims[0] - self.data[:, 0], new.data[:, 0]) >>> # xdoctest: +REQUIRES(--show) >>> import kwimage >>> im = kwimage.imresize(kwimage.grab_test_image(), dsize=input_dims[::-1]) >>> new_im = augmenter.augment_image(im) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(im, pnum=(1, 2, 1), fnum=1) >>> self.draw(color='red', alpha=.8, radius=0.5) >>> kwplot.imshow(new_im, pnum=(1, 2, 2), fnum=1) >>> new.draw(color='blue', alpha=.8, radius=0.5, coord_axes=[1, 0])
- to_imgaug(input_dims)[source]¶
Translate to an imgaug object
- Returns:
imgaug data structure
- Return type:
imgaug.KeypointsOnImage
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> import kwimage >>> import numpy as np >>> self = kwimage.Coords.random(10) >>> input_dims = (10, 10) >>> kpoi = self.to_imgaug(input_dims) >>> new = kwimage.Coords.from_imgaug(kpoi) >>> assert np.allclose(new.data, self.data)
- scale(factor, about=None, output_dims=None, inplace=False)[source]¶
Scale coordinates by a factor
- Parameters:
factor (float | Tuple[float, float]) – scale factor as either a scalar or per-dimension tuple.
about (Tuple | None) – if unspecified scales about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.scale(10) >>> assert new.data.max() <= 10
>>> self = Coords.random(10, rng=0) >>> self.data = (self.data * 10).astype(int) >>> new = self.scale(10) >>> assert new.data.dtype.kind == 'i' >>> new = self.scale(10.0) >>> assert new.data.dtype.kind == 'f'
- translate(offset, output_dims=None, inplace=False)[source]¶
Shift the coordinates
- Parameters:
offset (float | Tuple[float, float]) – transation offset as either a scalar or a per-dimension tuple.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=3, rng=0) >>> new = self.translate(10) >>> assert new.data.min() >= 10 >>> assert new.data.max() <= 11 >>> Coords.random(3, dim=3, rng=0) >>> Coords.random(3, dim=3, rng=0).translate((1, 2, 3))
- rotate(theta, about=None, output_dims=None, inplace=False)[source]¶
Rotate the coordinates about a point.
- Parameters:
theta (float) – rotation angle in radians
about (Tuple | None) – if unspecified rotates about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Todo
[ ] Generalized ND Rotations?
References
https://math.stackexchange.com/questions/197772/gen-rot-matrix
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=2, rng=0) >>> theta = np.pi / 2 >>> new = self.rotate(theta)
>>> # Test rotate agrees with warp >>> sin_ = np.sin(theta) >>> cos_ = np.cos(theta) >>> rot_ = np.array([[cos_, -sin_], [sin_, cos_]]) >>> new2 = self.warp(rot_) >>> assert np.allclose(new.data, new2.data)
>>> # >>> # Rotate about a custom point >>> theta = np.pi / 2 >>> new3 = self.rotate(theta, about=(0.5, 0.5)) >>> # >>> # Rotate about the center of mass >>> about = self.data.mean(axis=0) >>> new4 = self.rotate(theta, about=about) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> plt = kwplot.autoplt() >>> self.draw(radius=0.01, color='blue', alpha=.5, coord_axes=[1, 0], setlim='grow') >>> plt.gca().set_aspect('equal') >>> new3.draw(radius=0.01, color='red', alpha=.5, coord_axes=[1, 0], setlim='grow')
- _rectify_about(about)[source]¶
Ensures that about returns a specified point. Allows for special keys like center to be used.
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=2, rng=0)
- fill(image, value, coord_axes=None, interp='bilinear')[source]¶
Sets sub-coordinate locations in a grid to a particular value
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns:
image with coordinates rasterized on it
- Return type:
ndarray
- soft_fill(image, coord_axes=None, radius=5)[source]¶
Used for drawing keypoint truth in heatmaps
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns:
image with coordinates rasterized on it
- Return type:
ndarray
References
https://stackoverflow.com/questions/54726703/generating-keypoint-heatmaps-in-tensorflow
Example
>>> from kwimage.structs.coords import * # NOQA >>> s = 64 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> # Put points on edges to to verify "edge cases" >>> self.data[1] = [0, 0] # top left >>> self.data[2] = [s, s] # bottom right >>> self.data[3] = [0, s + 10] # bottom left >>> self.data[4] = [-3, s // 2] # middle left >>> self.data[5] = [s + 1, -1] # top right >>> # Put points in the middle to verify overlap blending >>> self.data[6] = [32.5, 32.5] # middle >>> self.data[7] = [34.5, 34.5] # middle >>> fill_value = 1 >>> coord_axes = [1, 0] >>> radius = 10 >>> image1 = np.zeros((s, s)) >>> self.soft_fill(image1, coord_axes=coord_axes, radius=radius) >>> radius = 3.0 >>> image2 = np.zeros((s, s)) >>> self.soft_fill(image2, coord_axes=coord_axes, radius=radius) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image1, pnum=(1, 2, 1)) >>> kwplot.imshow(image2, pnum=(1, 2, 2))

- draw_on(image=None, fill_value=1, coord_axes=[1, 0], interp='bilinear')[source]¶
Note
unlike other methods, the defaults assume x/y internal data
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns:
image with coordinates drawn on it
- Return type:
ndarray
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> s = 256 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> self.data[0] = [10, 10] >>> self.data[1] = [20, 40] >>> image = np.zeros((s, s)) >>> fill_value = 1 >>> image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[0, 1], interp='nearest') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='nearest') >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(image) >>> self.draw(radius=3, alpha=.5, coord_axes=[1, 0])

- draw(color='blue', ax=None, alpha=None, coord_axes=[1, 0], radius=1, setlim=False)[source]¶
Draw these coordinates via matplotlib
Note
unlike other methods, the defaults assume x/y internal data
- Parameters:
setlim (bool) – if True ensures the limits of the axes contains the polygon
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns:
drawn matplotlib objects
- Return type:
List[mpl.collections.PatchCollection]
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> self.draw(radius=0.05, alpha=0.8) >>> plt.gca().set_xlim(0, 1) >>> plt.gca().set_ylim(0, 1) >>> plt.gca().set_aspect('equal')
- class kwimage.Detections(data=None, meta=None, datakeys=None, metakeys=None, checks=True, **kwargs)[source]¶
Bases:
NiceRepr,_DetAlgoMixin,_DetDrawMixinContainer for holding and manipulating multiple detections.
- Variables:
data (Dict) –
dictionary containing corresponding lists. The length of each list is the number of detections. This contains the bounding boxes, confidence scores, and class indices. Details of the most common keys and types are as follows:
boxes (kwimage.Boxes[ArrayLike]): multiple bounding boxes scores (ArrayLike): associated scores class_idxs (ArrayLike): associated class indices segmentations (ArrayLike): segmentations masks for each box, members can be
MaskorMultiPolygon. keypoints (ArrayLike): keypoints for each box. Members should bePoints.Additional custom keys may be specified as long as (a) the values are array-like and the first axis corresponds to the standard data values and (b) are custom keys are listed in the datakeys kwargs when constructing the Detections.
meta (Dict) – This contains contextual information about the detections. This includes the class names, which can be indexed into via the class indexes.
Example
>>> import kwimage >>> dets = kwimage.Detections( >>> # there are expected keys that do not need registration >>> boxes=kwimage.Boxes.random(3), >>> class_idxs=[0, 1, 1], >>> classes=['a', 'b'], >>> # custom data attrs must align with boxes >>> myattr1=np.random.rand(3), >>> myattr2=np.random.rand(3, 2, 8), >>> # there are no restrictions on metadata >>> mymeta='a custom metadata string', >>> # Note that any key not in kwimage.Detections.__datakeys__ or >>> # kwimage.Detections.__metakeys__ must be registered at the >>> # time of construction. >>> datakeys=['myattr1', 'myattr2'], >>> metakeys=['mymeta'], >>> checks=True, >>> ) >>> print('dets = {}'.format(dets)) dets = <Detections(3)>
Construct a Detections object by either explicitly specifying the internal data and meta dictionary structures or by passing expected attribute names as kwargs.
- Parameters:
data (Dict[str, ArrayLike]) – explicitly specify the data dictionary
meta (Dict[str, object]) – explicitly specify the meta dictionary
datakeys (List[str]) – a list of custom attributes that should be considered as data (i.e. must be an array aligned with boxes).
metakeys (List[str]) – a list of custom attributes that should be considered as metadata (i.e. can be arbitrary).
checks (bool) – if True and arguments are passed by kwargs, then check / ensure that all types are compatible. Defaults to True.
**kwargs – specify any key for the data or meta dictionaries.
Note
Custom data and metadata can be specified as long as you pass the names of these keys in the datakeys and/or metakeys kwargs.
In the case where you specify a custom attribute as a list, it will “currently” (we may change this behavior in the future) be coerced into a numpy or torch array. If you want to store a generic Python list, wrap the custom list in a
_generic.ObjectList.Example
>>> # Coerce to numpy >>> import kwimage >>> dets = Detections( >>> boxes=kwimage.Boxes.random(3).numpy(), >>> class_idxs=[0, 1, 1], >>> checks=True, >>> ) >>> # xdoctest: +REQUIRES(module:torch) >>> # Coerce to tensor >>> dets = Detections( >>> boxes=kwimage.Boxes.random(3).tensor(), >>> class_idxs=[0, 1, 1], >>> checks=True, >>> ) >>> # Error on incompatible types >>> import pytest >>> with pytest.raises(TypeError): >>> dets = Detections( >>> boxes=kwimage.Boxes.random(3).tensor(), >>> scores=np.random.rand(3), >>> class_idxs=[0, 1, 1], >>> checks=True, >>> )
Example
>>> self = Detections.random(10) >>> other = Detections(self) >>> assert other.data == self.data >>> assert other.data is self.data, 'try not to copy unless necessary'
- classmethod coerce(data=None, **kwargs)[source]¶
The “try-anything to get what I want” constructor
- Parameters:
data
**kwargs – currently boxes and cnames
Example
>>> from kwimage.structs.detections import * # NOQA >>> import kwimage >>> kwargs = dict( >>> boxes=kwimage.Boxes.random(4), >>> cnames=['a', 'b', 'c', 'c'], >>> ) >>> data = {} >>> self = kwimage.Detections.coerce(data, **kwargs)
- classmethod from_coco_annots(anns, cats=None, classes=None, kp_classes=None, shape=None, dset=None)[source]¶
Create a Detections object from a list of coco-like annotations.
- Parameters:
anns (List[Dict]) – list of coco-like annotation objects
dset (kwcoco.CocoDataset) – if specified, cats, classes, and kp_classes can are ignored.
cats (List[Dict]) – coco-format category information. Used only if dset is not specified.
classes (kwcoco.CategoryTree) – category tree with coco class info. Used only if dset is not specified.
kp_classes (kwcoco.CategoryTree) – keypoint category tree with coco keypoint class info. Used only if dset is not specified.
shape (tuple) – shape of parent image
- Returns:
a detections object
- Return type:
Example
>>> from kwimage.structs.detections import * # NOQA >>> # xdoctest: +REQUIRES(--module:ndsampler) >>> anns = [{ >>> 'id': 0, >>> 'image_id': 1, >>> 'category_id': 2, >>> 'bbox': [2, 3, 10, 10], >>> 'keypoints': [4.5, 4.5, 2], >>> 'segmentation': { >>> 'counts': '_11a04M2O0O20N101N3L_5', >>> 'size': [20, 20], >>> }, >>> }] >>> dataset = { >>> 'images': [], >>> 'annotations': [], >>> 'categories': [ >>> {'id': 0, 'name': 'background'}, >>> {'id': 2, 'name': 'class1', 'keypoints': ['spot']} >>> ] >>> } >>> #import ndsampler >>> #dset = ndsampler.CocoDataset(dataset) >>> cats = dataset['categories'] >>> dets = Detections.from_coco_annots(anns, cats)
Example
>>> # xdoctest: +REQUIRES(--module:ndsampler) >>> # Test case with no category information >>> from kwimage.structs.detections import * # NOQA >>> anns = [{ >>> 'id': 0, >>> 'image_id': 1, >>> 'category_id': None, >>> 'bbox': [2, 3, 10, 10], >>> 'prob': [.1, .9], >>> }] >>> cats = [ >>> {'id': 0, 'name': 'background'}, >>> {'id': 2, 'name': 'class1'} >>> ] >>> dets = Detections.from_coco_annots(anns, cats)
Example
>>> import kwimage >>> # xdoctest: +REQUIRES(--module:ndsampler) >>> import ndsampler >>> sampler = ndsampler.CocoSampler.demo('photos') >>> iminfo, anns = sampler.load_image_with_annots(1) >>> shape = iminfo['imdata'].shape[0:2] >>> kp_classes = sampler.dset.keypoint_categories() >>> dets = kwimage.Detections.from_coco_annots( >>> anns, sampler.dset.dataset['categories'], sampler.catgraph, >>> kp_classes, shape=shape)
- to_coco(cname_to_cat=None, style='orig', image_id=None, dset=None)[source]¶
Converts this set of detections into coco-like annotation dictionaries.
Note
Not all aspects of the MS-COCO format can be accurately represented, so some liberties are taken. The MS-COCO standard defines that annotations should specifiy a category_id field, but in some cases this information is not available so we will populate a ‘category_name’ field if possible and in the worst case fall back to ‘category_index’.
Additionally, detections may contain additional information beyond the MS-COCO standard, and this information (e.g. weight, prob, score) is added as forign fields.
- Parameters:
cname_to_cat – currently ignored.
style (str) – either ‘orig’ (for the original coco format) or ‘new’ for the more general kwcoco-style coco format. Defaults to ‘orig’
image_id (int) – if specified, populates the image_id field of each image.
dset (kwcoco.CocoDataset | None) – if specified, attempts to populate the category_id field to be compatible with this coco dataset.
- Yields:
dict – coco-like annotation structures
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> from kwimage.structs.detections import * >>> self = Detections.demo()[0] >>> cname_to_cat = None >>> list(self.to_coco())
- property boxes¶
- property class_idxs¶
- property scores¶
typically only populated for predicted detections
- property probs¶
typically only populated for predicted detections
- property weights¶
typically only populated for groundtruth detections
- property classes¶
- warp(transform, input_dims=None, output_dims=None, inplace=False)[source]¶
Spatially warp the detections.
- Parameters:
transform (kwimage.Affine | ndarray | Callable | Any) – Something coercable to a transform. Usually a kwimage.Affine object
input_dims (Tuple[int, int]) – shape of the expected input canvas
output_dims (Tuple[int, int]) – shape of the expected output canvas
inplace (bool) – if true operate inplace
- Returns:
the warped detections object
- Return type:
Example
>>> import skimage >>> transform = skimage.transform.AffineTransform(scale=(2, 3), translation=(4, 5)) >>> self = Detections.random(2) >>> new = self.warp(transform) >>> assert new.boxes == self.boxes.warp(transform) >>> assert new != self
- scale(factor, output_dims=None, inplace=False)[source]¶
Spatially scale the detections.
Example
>>> import skimage >>> transform = skimage.transform.AffineTransform(scale=(2, 3), translation=(4, 5)) >>> self = Detections.random(2) >>> new = self.warp(transform) >>> assert new.boxes == self.boxes.warp(transform) >>> assert new != self
- translate(offset, output_dims=None, inplace=False)[source]¶
Spatially translate the detections.
Example
>>> import skimage >>> self = Detections.random(2) >>> new = self.translate(10)
- classmethod concatenate(dets)[source]¶
- Parameters:
boxes (Sequence[Detections]) – list of detections to concatenate
- Returns:
stacked detections
- Return type:
Example
>>> self = Detections.random(2) >>> other = Detections.random(3) >>> dets = [self, other] >>> new = Detections.concatenate(dets) >>> assert new.num_boxes() == 5
>>> self = Detections.random(2, segmentations=True) >>> other = Detections.random(3, segmentations=True) >>> dets = [self, other] >>> new = Detections.concatenate(dets) >>> assert new.num_boxes() == 5
- argsort(reverse=True)[source]¶
Sorts detection indices by descending (or ascending) scores
- Returns:
sorted indices torch.Tensor: sorted indices if using torch backends
- Return type:
ndarray[Shape[‘*’], Integer]
- sort(reverse=True)[source]¶
Sorts detections by descending (or ascending) scores
- Returns:
sorted copy of self
- Return type:
- compress(flags, axis=0)[source]¶
Returns a subset where corresponding locations are True.
- Parameters:
flags (ndarray[Any, Bool] | torch.Tensor) – mask marking selected items
- Returns:
subset of self
- Return type:
CommandLine
xdoctest -m kwimage.structs.detections Detections.compress
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> dets = kwimage.Detections.random(keypoints='dense') >>> flags = np.random.rand(len(dets)) > 0.5 >>> subset = dets.compress(flags) >>> assert len(subset) == flags.sum() >>> subset = dets.tensor().compress(flags) >>> assert len(subset) == flags.sum()
- take(indices, axis=0)[source]¶
Returns a subset specified by indices
- Parameters:
indices (ndarray[Any, Integer]) – indices to select
- Returns:
subset of self
- Return type:
Example
>>> import kwimage >>> dets = kwimage.Detections(boxes=kwimage.Boxes.random(10)) >>> subset = dets.take([2, 3, 5, 7]) >>> assert len(subset) == 4 >>> # xdoctest: +REQUIRES(module:torch) >>> subset = dets.tensor().take([2, 3, 5, 7]) >>> assert len(subset) == 4
- property device¶
If the backend is torch returns the data device, otherwise None
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Detections.random(3).tensor() >>> newself = self.numpy() >>> self.scores[0] = 0 >>> assert newself.scores[0] == 0 >>> self.scores[0] = 1 >>> assert self.scores[0] == 1 >>> self.numpy().numpy()
- property dtype¶
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.detections import * >>> self = Detections.random(3) >>> newself = self.tensor() >>> self.scores[0] = 0 >>> assert newself.scores[0] == 0 >>> self.scores[0] = 1 >>> assert self.scores[0] == 1 >>> self.tensor().tensor()
- classmethod random(num=10, scale=1.0, classes=3, keypoints=False, segmentations=False, tensor=False, rng=None)[source]¶
Creates dummy data, suitable for use in tests and benchmarks
- Parameters:
num (int) – number of boxes
scale (float | tuple) – bounding image size. Defaults to 1.0
classes (int | Sequence) – list of class labels or number of classes
keypoints (bool) – if True include random keypoints for each box. Defaults to False.
segmentations (bool) – if True include random segmentations for each box. Defaults to False.
tensor (bool) – determines backend. DEPRECATED. Call
.tensor()on resulting object instead.rng (int | RandomState | None) – random state or seed
- Returns:
random detections
- Return type:
Example
>>> import kwimage >>> dets = kwimage.Detections.random(keypoints='jagged') >>> dets.data['keypoints'].data[0].data >>> dets.data['keypoints'].meta >>> dets = kwimage.Detections.random(keypoints='dense') >>> dets = kwimage.Detections.random(keypoints='dense', segmentations=True).scale(1000) >>> # xdoctest:+REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets.draw(setlim=True)
Example
>>> import kwimage >>> dets = kwimage.Detections.random( >>> keypoints='jagged', segmentations=True, rng=0).scale(1000) >>> print('dets = {}'.format(dets)) dets = <Detections(10)> >>> dets.data['boxes'].quantize(inplace=True) >>> print('dets.data = {}'.format(ub.urepr( >>> dets.data, nl=1, with_dtype=False, strvals=True, sort=1))) dets.data = { 'boxes': <Boxes(xywh, array([[548, 544, 55, 172], [423, 645, 15, 247], [791, 383, 173, 146], [ 71, 87, 498, 839], [ 20, 832, 759, 39], [461, 780, 518, 20], [118, 639, 26, 306], [264, 414, 258, 361], [ 18, 568, 439, 50], [612, 616, 332, 66]]...))>, 'class_idxs': [1, 2, 0, 0, 2, 0, 0, 0, 0, 0], 'keypoints': <PointsList(n=10)>, 'scores': [0.3595079 , 0.43703195, 0.6976312 , 0.06022547, 0.66676672, 0.67063787,0.21038256, 0.1289263 , 0.31542835, 0.36371077], 'segmentations': <SegmentationList(n=10)>, } >>> # xdoctest:+REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets.draw(setlim=True)
Example
>>> # Boxes position/shape within 0-1 space should be uniform. >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> fig.gca().set_xlim(0, 128) >>> fig.gca().set_ylim(0, 128) >>> import kwimage >>> kwimage.Detections.random(num=10, segmentations=True).scale(128).draw()
- class kwimage.Heatmap(data=None, meta=None, **kwargs)[source]¶
Bases:
Spatial,_HeatmapDrawMixin,_HeatmapWarpMixin,_HeatmapAlgoMixinKeeps track of a downscaled heatmap and how to transform it to overlay the original input image. Heatmaps generally are used to estimate class probabilites at each pixel. This data struction additionally contains logic to augment pixel with offset (dydx) and scale (diamter) information.
- Variables:
data (Dict[str, ArrayLike]) –
dictionary containing spatially aligned heatmap data. Valid keys are as follows.
- class_probs (ArrayLike[C, H, W] | ArrayLike[C, D, H, W]):
A probability map for each class. C is the number of classes.
- offset (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object center position offset in y,x / t,y,x coordinates
- diamter (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object bounding box sizes in h,w / d,h,w coordinates
- keypoints (ArrayLike[2, K, H, W] | ArrayLike[3, K, D, H, W], optional):
y/x offsets for K different keypoint classes
dictionary containing miscellanious metadata about the heatmap data. Valid keys are as follows.
- img_dims (Tuple[H, W] | Tuple[D, H, W]):
original image dimension
- tf_data_to_image (SKImageGeometricTransform):
transformation matrix (typically similarity or affine) that projects the given, heatmap onto the image dimensions such that the image and heatmap are spatially aligned.
- classes (List[str] | ndsampler.CategoryTree):
information about which index in
data['class_probs']corresponds to which semantic class.
dims (Tuple) – dimensions of the heatmap (See
image_dims) for the original image dimensions.**kwargs – any key that is accepted by the data or meta dictionaries can be specified as a keyword argument to this class and it will be properly placed in the appropriate internal dictionary.
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/heatmap.py Heatmap --show
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> import skimage >>> import kwimage >>> class_probs = kwimage.grab_test_image(dsize=(32, 32), space='gray')[None, ..., 0] / 255.0 >>> img_dims = (220, 220) >>> tf_data_to_img = skimage.transform.AffineTransform(translation=(-18, -18), scale=(8, 8)) >>> self = Heatmap(class_probs=class_probs, img_dims=img_dims, >>> tf_data_to_img=tf_data_to_img) >>> aligned = self.upscale() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(aligned[0]) >>> kwplot.show_if_requested()

Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> self = Heatmap.random() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw()
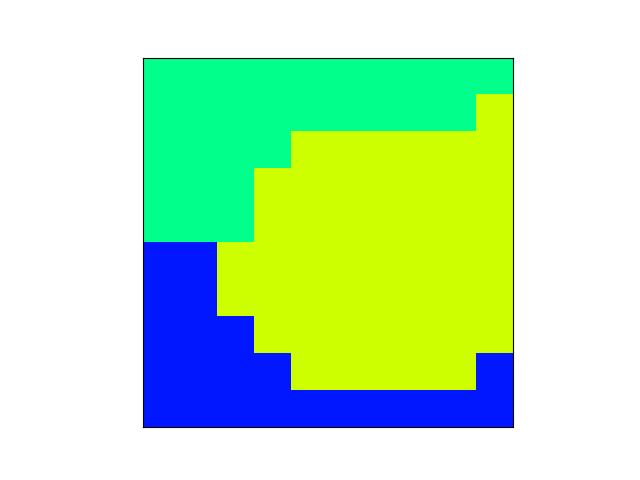

- property shape¶
- property bounds¶
- property dims¶
space-time dimensions of this heatmap
- property _impl¶
Returns the internal tensor/numpy ArrayAPI implementation
- Returns:
kwarray.ArrayAPI
- classmethod random(dims=(10, 10), classes=3, diameter=True, offset=True, keypoints=False, img_dims=None, dets=None, nblips=10, noise=0.0, smooth_k=3, rng=None, ensure_background=True)[source]¶
Creates dummy data, suitable for use in tests and benchmarks
- Parameters:
dims (Tuple[int, int]) – dimensions of the heatmap
classes (int | List[str] | kwcoco.CategoryTree) – foreground classes
diameter (bool) – if True, include a “diameter” heatmap
offset (bool) – if True, include an “offset” heatmap
keypoints (bool)
smooth_k (int) – kernel size for gaussian blur to smooth out the heatmaps.
img_dims (Tuple) – dimensions of an upscaled image the heatmap corresponds to. (This should be removed and simply handled with a transform
in the future).
- Returns:
Heatmap
Example
>>> from kwimage.structs.heatmap import * # NOQA >>> self = Heatmap.random((128, 128), img_dims=(200, 200), >>> classes=3, nblips=10, rng=0, noise=0.1) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(self.colorize(0, imgspace=0), fnum=1, pnum=(1, 4, 1), doclf=1) >>> kwplot.imshow(self.colorize(1, imgspace=0), fnum=1, pnum=(1, 4, 2)) >>> kwplot.imshow(self.colorize(2, imgspace=0), fnum=1, pnum=(1, 4, 3)) >>> kwplot.imshow(self.colorize(3, imgspace=0), fnum=1, pnum=(1, 4, 4))
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> self = kwimage.Heatmap.random(dims=(50, 200), dets='coco', >>> keypoints=True) >>> image = np.zeros(self.img_dims) >>> # xdoctest: +REQUIRES(module:kwplot) >>> toshow = self.draw_on(image, 1, vecs=True, kpts=0, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(toshow)
- property class_probs¶
- property offset¶
- property diameter¶
- property img_dims¶
- property tf_data_to_img¶
- property classes¶
- class kwimage.Mask(data=None, format=None)[source]¶
Bases:
NiceRepr,_MaskConversionMixin,_MaskConstructorMixin,_MaskTransformMixin,_MaskDrawMixinManages a single segmentation mask and can convert to and from multiple formats including:
bytes_rle - byte encoded run length encoding
array_rle - raw run length encoding
c_mask - c-style binary mask
f_mask - fortran-style binary mask
Example
>>> # xdoctest: +REQUIRES(--mask) >>> # a ms-coco style compressed bytes rle segmentation >>> segmentation = {'size': [5, 9], 'counts': ';?1B10O30O4'} >>> mask = Mask(segmentation, 'bytes_rle') >>> # convert to binary numpy representation >>> binary_mask = mask.to_c_mask().data >>> print(ub.urepr(binary_mask.tolist(), nl=1, nobr=1)) [0, 0, 0, 1, 1, 1, 1, 1, 0], [0, 0, 1, 1, 1, 0, 0, 0, 0], [0, 0, 1, 1, 1, 1, 1, 1, 0], [0, 0, 1, 1, 1, 0, 1, 1, 0], [0, 0, 1, 1, 1, 0, 1, 1, 0],
- property dtype¶
- classmethod random(rng=None, shape=(32, 32))[source]¶
Create a random binary mask object
- Parameters:
rng (int | RandomState | None) – the random seed
shape (Tuple[int, int]) – the height / width of the returned mask
- Returns:
the random mask
- Return type:
Example
>>> import kwimage >>> mask = kwimage.Mask.random() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> mask.draw() >>> kwplot.show_if_requested()
- classmethod demo()[source]¶
Demo mask with holes and disjoint shapes
- Returns:
the demo mask
- Return type:
- classmethod from_text(text, zero_chr='.', shape=None, has_border=False)[source]¶
Construct a mask from a text art representation
- Parameters:
text (str) – the text representing a mask
zero_chr (str) – the character that represents a zero
shape (None | Tuple[int, int]) – if specified force a specific height / width, otherwise the character extent determines this.
has_border (bool) – if True, assume the characters at the edge are representing a border and remove them.
Example
>>> import kwimage >>> import ubelt as ub >>> text = ub.indent(ub.codeblock( >>> ''' >>> ooo >>> ooo >>> ooooo >>> o >>> ''')) >>> mask = kwimage.Mask.from_text(text, zero_chr=' ') >>> print(mask.data) [[0 0 0 0 1 1 1 0 0] [0 0 0 0 1 1 1 0 0] [0 0 0 0 1 1 1 1 1] [0 0 0 0 0 0 0 0 1]]
Example
>>> import kwimage >>> import ubelt as ub >>> text = ub.codeblock( >>> ''' >>> +------------+ >>> | | >>> | ooo | >>> | ooo | >>> | ooooo | >>> | o | >>> | | >>> +------------+ >>> ''') >>> mask = kwimage.Mask.from_text(text, has_border=True, zero_chr=' ') >>> print(mask.data) [[0 0 0 0 0 0 0 0 0 0 0 0] [0 0 0 0 1 1 1 0 0 0 0 0] [0 0 0 0 1 1 1 0 0 0 0 0] [0 0 0 0 1 1 1 1 1 0 0 0] [0 0 0 0 0 0 0 0 1 0 0 0] [0 0 0 0 0 0 0 0 0 0 0 0]]
- copy()[source]¶
Performs a deep copy of the mask data
- Returns:
the copied mask
- Return type:
Example
>>> self = Mask.random(shape=(8, 8), rng=0) >>> other = self.copy() >>> assert other.data is not self.data
- union(*others)[source]¶
This can be used as a staticmethod or an instancemethod
- Parameters:
*others – multiple input masks to union
- Returns:
the unioned mask
- Return type:
Example
>>> # xdoctest: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> masks = [Mask.random(shape=(8, 8), rng=i) for i in range(2)] >>> mask = Mask.union(*masks) >>> print(mask.area) >>> masks = [m.to_c_mask() for m in masks] >>> mask = Mask.union(*masks) >>> print(mask.area)
>>> masks = [m.to_bytes_rle() for m in masks] >>> mask = Mask.union(*masks) >>> print(mask.area)
- intersection(*others)[source]¶
This can be used as a staticmethod or an instancemethod
- Parameters:
*others – multiple input masks to intersect
- Returns:
the intersection of the masks
- Return type:
Example
>>> n = 3 >>> masks = [Mask.random(shape=(8, 8), rng=i) for i in range(n)] >>> items = masks >>> mask = Mask.intersection(*masks) >>> areas = [item.area for item in items] >>> print('areas = {!r}'.format(areas)) >>> print(mask.area) >>> print(Mask.intersection(*masks).area / Mask.union(*masks).area)
- property shape¶
- property area¶
Returns the number of non-zero pixels
- Returns:
the number of non-zero pixels
- Return type:
Example
>>> self = Mask.demo() >>> self.area 150
- get_patch()[source]¶
Extract the patch with non-zero data
Example
>>> # xdoctest: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.random(shape=(8, 8), rng=0) >>> self.get_patch()
- get_xywh()[source]¶
Gets the bounding xywh box coordinates of this mask
- Returns:
- x, y, w, h: Note we dont use a Boxes object because
a general singular version does not yet exist.
- Return type:
ndarray
Example
>>> # xdoctest: +REQUIRES(--mask) >>> self = Mask.random(shape=(8, 8), rng=0) >>> self.get_xywh().tolist() >>> self = Mask.random(rng=0).translate((10, 10)) >>> self.get_xywh().tolist()
Example
>>> # test empty case >>> import kwimage >>> self = kwimage.Mask(np.empty((0, 0), dtype=np.uint8), format='c_mask') >>> assert self.get_xywh().tolist() == [0, 0, 0, 0]
- get_polygon()[source]¶
DEPRECATED: USE to_multi_polygon
Returns a list of (x,y)-coordinate lists. The length of the list is equal to the number of disjoint regions in the mask.
- Returns:
- polygon around each connected component of the
mask. Each ndarray is an Nx2 array of xy points.
- Return type:
List[ndarray]
Note
The returned polygon may not surround points that are only one pixel thick.
Example
>>> # xdoctest: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.random(shape=(8, 8), rng=0) >>> polygons = self.get_polygon() >>> print('polygons = ' + ub.urepr(polygons)) >>> polygons = self.get_polygon() >>> self = self.to_bytes_rle() >>> other = Mask.from_polygons(polygons, self.shape) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> image = np.ones(self.shape) >>> image = self.draw_on(image, color='blue') >>> image = other.draw_on(image, color='red') >>> kwplot.imshow(image)
- to_mask(dims=None, pixels_are='points')[source]¶
Converts to a mask object (which does nothing because this already is mask object!)
- Returns:
kwimage.Mask
- to_multi_polygon(pixels_are='points')[source]¶
Returns a MultiPolygon object fit around this raster including disjoint pieces and holes.
- Parameters:
pixel_are (str) – Can either be “points” or “areas”.
If pixels are “points”, the we treat each pixel (i, j) as a single infinitely small point at (i, j). As such, some polygons may have zero area.
If pixels are “areas”, then each pixel (i, j) represents a square with coordinates ([i - 0.5, j - 0.5], [i + 0.5, j - 0.5], [i + 0.5, j + 0.5], and [i - 0.5, j + 0.5]). Must have rasterio installed to use this method.
- Returns:
vectorized representation
- Return type:
Note
The OpenCV (and thus this function) coordinate system places coordinates at the center of pixels, and the polygon is traced tightly around these coordinates. A single pixel is not considered to have any width, so polygon edges will directly trace through the centers of pixels, and in the case where an object is only 1 pixel thick, this will produce a polygon that is not a valid shapely polygon.
Todo
[x] add a flag where polygons consider pixels to have width and the resulting polygon is traced around the pixel edges, not the pixel centers.
[ ] Polygons and Masks should keep track of what “pixels_are”
Example
>>> # xdoctest: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.demo() >>> self = self.scale(5) >>> multi_poly = self.to_multi_polygon() >>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> self.draw(color='red') >>> multi_poly.scale(1.1).draw(color='blue')
>>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> image = np.ones(self.shape) >>> image = self.draw_on(image, color='blue') >>> #image = other.draw_on(image, color='red') >>> kwplot.imshow(image) >>> multi_poly.draw()
Example
>>> # Test empty cases >>> import kwimage >>> mask0 = kwimage.Mask(np.zeros((0, 0), dtype=np.uint8), format='c_mask') >>> mask1 = kwimage.Mask(np.zeros((1, 1), dtype=np.uint8), format='c_mask') >>> mask2 = kwimage.Mask(np.zeros((2, 2), dtype=np.uint8), format='c_mask') >>> mask3 = kwimage.Mask(np.zeros((3, 3), dtype=np.uint8), format='c_mask') >>> pixels_are = 'points' >>> poly0 = mask0.to_multi_polygon(pixels_are=pixels_are) >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert len(poly0) == 0 >>> assert len(poly1) == 0 >>> assert len(poly2) == 0 >>> assert len(poly3) == 0 >>> # xdoctest: +REQUIRES(module:rasterio) >>> pixels_are = 'areas' >>> poly0 = mask0.to_multi_polygon(pixels_are=pixels_are) >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert len(poly0) == 0 >>> assert len(poly1) == 0 >>> assert len(poly2) == 0 >>> assert len(poly3) == 0
Example
>>> # Test full ones cases >>> import kwimage >>> mask1 = kwimage.Mask(np.ones((1, 1), dtype=np.uint8), format='c_mask') >>> mask2 = kwimage.Mask(np.ones((2, 2), dtype=np.uint8), format='c_mask') >>> mask3 = kwimage.Mask(np.ones((3, 3), dtype=np.uint8), format='c_mask') >>> pixels_are = 'points' >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert np.all(poly1.to_mask(mask1.shape).data == 1) >>> assert np.all(poly2.to_mask(mask2.shape).data == 1) >>> assert np.all(poly3.to_mask(mask3.shape).data == 1) >>> # xdoctest: +REQUIRES(module:rasterio) >>> pixels_are = 'areas' >>> poly1 = mask1.to_multi_polygon(pixels_are=pixels_are) >>> poly2 = mask2.to_multi_polygon(pixels_are=pixels_are) >>> poly3 = mask3.to_multi_polygon(pixels_are=pixels_are) >>> assert np.all(poly1.to_mask(mask1.shape).data == 1) >>> assert np.all(poly2.to_mask(mask2.shape).data == 1) >>> assert np.all(poly3.to_mask(mask3.shape).data == 1)
Example
>>> # Corner case, only two pixels are on >>> import kwimage >>> self = kwimage.Mask(np.zeros((768, 768), dtype=np.uint8), format='c_mask') >>> x_coords = np.array([621, 752]) >>> y_coords = np.array([366, 292]) >>> self.data[y_coords, x_coords] = 1 >>> poly = self.to_multi_polygon()
Example
>>> # xdoctest: +REQUIRES(module:rasterio) >>> import kwimage >>> dims = (10, 10) >>> data = np.zeros(dims, dtype=np.uint8) >>> data[0, 3:5] = 1 >>> data[9, 1:3] = 1 >>> data[3:5, 0:2] = 1 >>> data[1, 1] = 1 >>> # 1 pixel L shape >>> data[3, 5] = 1 >>> data[4, 5] = 1 >>> data[4, 6] = 1 >>> data[1, 5] = 1 >>> data[2, 6] = 1 >>> data[3, 7] = 1 >>> data[6, 1] = 1 >>> data[7, 1] = 1 >>> data[7, 2] = 1 >>> data[6:10, 5] = 1 >>> data[6:10, 8] = 1 >>> data[9, 5:9] = 1 >>> data[6, 5:9] = 1 >>> #data = kwimage.imresize(data, scale=2.0, interpolation='nearest') >>> self = kwimage.Mask.coerce(data) >>> #self = self.translate((0, 0), output_dims=(10, 9)) >>> self = self.translate((0, 1), output_dims=(11, 11)) >>> dims = self.shape[0:2] >>> multi_poly1 = self.to_multi_polygon(pixels_are='points') >>> multi_poly2 = self.to_multi_polygon(pixels_are='areas') >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pretty_data = kwplot.make_heatmask(self.data/1.0, cmap='magma')[..., 0:3] >>> def _pixel_grid_lines(self, ax): >>> h, w = self.data.shape[0:2] >>> ybasis = np.arange(0, h) + 0.5 >>> xbasis = np.arange(0, w) + 0.5 >>> xmin = 0 - 0.5 >>> xmax = w - 0.5 >>> ymin = 0 - 0.5 >>> ymax = h - 0.5 >>> ax.hlines(y=ybasis, xmin=xmin, xmax=xmax, color="gainsboro") >>> ax.vlines(x=xbasis, ymin=ymin, ymax=ymax, color="gainsboro") >>> def _setup_grid(self, pnum): >>> ax = kwplot.imshow(pretty_data, show_ticks=True, pnum=pnum)[1] >>> # The gray ticks show the center of the pixels >>> ax.grid(color='dimgray', linewidth=0.5) >>> ax.set_xticks(np.arange(self.data.shape[1])) >>> ax.set_yticks(np.arange(self.data.shape[0])) >>> # Also draw black lines around the edges of the pixels >>> _pixel_grid_lines(self, ax=ax) >>> return ax >>> # Overlay the extracted polygons >>> ax = _setup_grid(self, pnum=(2, 3, 1)) >>> ax.set_title('input binary mask data') >>> ax = _setup_grid(self, pnum=(2, 3, 2)) >>> multi_poly1.draw(linewidth=5, alpha=0.5, radius=0.2, ax=ax, fill=False, vertex=0.2) >>> ax.set_title('opencv "point" polygons') >>> ax = _setup_grid(self, pnum=(2, 3, 3)) >>> multi_poly2.draw(linewidth=5, alpha=0.5, radius=0.2, color='limegreen', ax=ax, fill=False, vertex=0.2) >>> ax.set_title('raterio "area" polygons') >>> ax.figure.suptitle(ub.codeblock( >>> ''' >>> Gray lines are coordinates and pass through pixel centers (integer coords) >>> White lines trace pixel boundaries (fractional coords) >>> ''')) >>> raster1 = multi_poly1.to_mask(dims, pixels_are='points') >>> raster2 = multi_poly2.to_mask(dims, pixels_are='areas') >>> kwplot.imshow(raster1.draw_on(), pnum=(2, 3, 5), title='rasterized') >>> kwplot.imshow(raster2.draw_on(), pnum=(2, 3, 6), title='rasterized')
- get_convex_hull()[source]¶
Returns a list of xy points around the convex hull of this mask
Note
The returned polygon may not surround points that are only one pixel thick.
Example
>>> # xdoctest: +REQUIRES(--mask) >>> self = Mask.random(shape=(8, 8), rng=0) >>> polygons = self.get_convex_hull() >>> print('polygons = ' + ub.urepr(polygons)) >>> other = Mask.from_polygons(polygons, self.shape)
- iou(other)[source]¶
The area of intersection over the area of union
Todo
[ ] Write plural Masks version of this class, which should be able to perform this operation more efficiently.
CommandLine
xdoctest -m kwimage.structs.mask Mask.iou
Example
>>> # xdoctest: +REQUIRES(--mask) >>> self = Mask.demo() >>> other = self.translate(1) >>> iou = self.iou(other) >>> print('iou = {:.4f}'.format(iou)) iou = 0.0830 >>> iou2 = self.intersection(other).area / self.union(other).area >>> print('iou2 = {:.4f}'.format(iou2))
- classmethod coerce(data, dims=None)[source]¶
Attempts to auto-inspect the format of the data and conver to Mask
- Parameters:
data (Any) – the data to coerce
dims (Tuple) – required for certain formats like polygons height / width of the source image
- Returns:
the constructed mask object
- Return type:
Example
>>> # xdoctest: +REQUIRES(--mask) >>> segmentation = {'size': [5, 9], 'counts': ';?1B10O30O4'} >>> polygon = [ >>> [np.array([[3, 0],[2, 1],[2, 4],[4, 4],[4, 3],[7, 0]])], >>> [np.array([[2, 1],[2, 2],[4, 2],[4, 1]])], >>> ] >>> dims = (9, 5) >>> mask = (np.random.rand(32, 32) > .5).astype(np.uint8) >>> Mask.coerce(polygon, dims).to_bytes_rle() >>> Mask.coerce(segmentation).to_bytes_rle() >>> Mask.coerce(mask).to_bytes_rle()
- to_coco(style='orig')[source]¶
Convert the Mask to a COCO json representation based on the current format.
A COCO mask is formatted as a run-length-encoding (RLE), of which there are two variants: (1) a array RLE, which is slightly more readable and extensible, and (2) a bytes RLE, which is slightly more concise. The returned format will depend on the current format of the Mask object. If it is in “bytes_rle” format, it will be returned in that format, otherwise it will be converted to the “array_rle” format and returned as such.
- Parameters:
style (str) – Does nothing for this particular method, exists for API compatibility and if alternate encoding styles are implemented in the future.
- Returns:
- either a bytes-rle or array-rle encoding, depending
on the current mask format. The keys in this dictionary are as follows:
counts (List[int] | str): the array or bytes rle encoding
- size (Tuple[int]): the height and width of the encoded mask
see note.
- shape (Tuple[int]): only present in array-rle mode. This
is also the height/width of the underlying encoded array. This exists for semantic consistency with other kwimage conventions, and is not part of the original coco spec.
- order (str): only present in array-rle mode.
Either C or F, indicating if counts is aranged in row-major or column-major order. For COCO-compatibility this is always returned in F (column-major) order.
- binary (bool): only present in array-rle mode.
For COCO-compatibility this is always returned as False, indicating the mask only contains binary 0 or 1 values.
- Return type:
Note
The output dictionary will contain a key named “size”, this is the only location in kwimage where “size” refers to a tuple in (height/width) order, in order to be backwards compatible with the original coco spec. In all other locations in kwimage a “size” will refer to a (width/height) ordered tuple.
- SeeAlso:
- func:
kwimage.im_runlen.encode_run_length - backend function that does array-style run length encoding.
Example
>>> # xdoctest: +REQUIRES(--mask) >>> from kwimage.structs.mask import * # NOQA >>> self = Mask.demo() >>> coco_data1 = self.toformat('array_rle').to_coco() >>> coco_data2 = self.toformat('bytes_rle').to_coco() >>> print('coco_data1 = {}'.format(ub.urepr(coco_data1, nl=1))) >>> print('coco_data2 = {}'.format(ub.urepr(coco_data2, nl=1))) coco_data1 = { 'binary': True, 'counts': [47, 5, 3, 1, 14, ... 1, 4, 19, 141], 'order': 'F', 'shape': (23, 32), 'size': (23, 32), } coco_data2 = { 'counts': '_153L;4EL...ON3060L0N060L0Nb0Y4', 'size': [23, 32], }
- class kwimage.MaskList(data, meta=None)[source]¶
Bases:
ObjectListStore and manipulate multiple masks, usually within the same image
- to_polygon_list()[source]¶
Converts all mask objects to multi-polygon objects
- Returns:
kwimage.PolygonList
- class kwimage.Matrix(matrix)[source]¶
Bases:
TransformBase class for matrix-based transform.
Example
>>> from kwimage.transform import * # NOQA >>> ms = {} >>> ms['random()'] = Matrix.random() >>> ms['eye()'] = Matrix.eye() >>> ms['random(3)'] = Matrix.random(3) >>> ms['random(4, 4)'] = Matrix.random(4, 4) >>> ms['eye(3)'] = Matrix.eye(3) >>> ms['explicit'] = Matrix(np.array([[1.618]])) >>> for k, m in ms.items(): >>> print('----') >>> print(f'{k} = {m}') >>> print(f'{k}.inv() = {m.inv()}') >>> print(f'{k}.T = {m.T}') >>> print(f'{k}.det() = {m.det()}')
- property shape¶
- classmethod coerce(data=None, **kwargs)[source]¶
Example
>>> Matrix.coerce({'type': 'matrix', 'matrix': [[1, 0, 0], [0, 1, 0]]}) >>> Matrix.coerce(np.eye(3)) >>> Matrix.coerce(None)
- inv()[source]¶
Returns the inverse of this matrix
- Returns:
Matrix
Example
>>> # Test with rationals >>> # xdoctest: +REQUIRES(module:sympy) >>> import kwimage >>> self = kwimage.Matrix.random((3, 3)).rationalize() >>> inv = self.inv() >>> eye = self @ inv >>> eye.isclose_identity(0, 0)
- property T¶
Transpose the underlying matrix
- rationalize()[source]¶
Convert the underlying matrix to a rational type to avoid floating point errors. This does decrease efficiency.
Todo
[ ] mpmath for arbitrary precision? It doesn’t seem to do
inverses correct whereas sympy does.
Example
>>> # xdoctest: +REQUIRES(module:sympy) >>> import kwimage >>> self = kwimage.Matrix.random((3, 3)) >>> mat = self.rationalize() >>> mat2 = kwimage.Matrix.random((3, 3)) >>> mat3 = mat @ mat2 >>> #assert 'sympy' in mat3.matrix.__class__.__module__ >>> mat3 = mat2 @ mat >>> #assert 'sympy' in mat3.matrix.__class__.__module__ >>> assert not mat.isclose_identity() >>> assert (mat @ mat.inv()).isclose_identity(rtol=0, atol=0)
- class kwimage.MultiPolygon(data, meta=None)[source]¶
Bases:
ObjectList,_ShapelyMixinData structure for storing multiple polygons (typically related to the same underlying but potentitally disjoing object)
- Variables:
data (List[Polygon]) –
- classmethod random(n=3, n_holes=0, rng=None, tight=False)[source]¶
Create a random MultiPolygon
- Returns:
MultiPolygon
- fill(image, value=1, pixels_are='points', assert_inplace=False)[source]¶
Inplace fill in an image based on this multi-polyon.
- Parameters:
image (ndarray) – image to draw on (inplace)
value (int | Tuple[int, …]) – value fill in with. Defaults to 1.0
- Returns:
the image that has been modified in place
- Return type:
ndarray
- bounding_box()[source]¶
Return the bounding box of the multi polygon
DEPRECATED: Use singular
box()instead.- Returns:
- a Boxes object with one box that encloses all
polygons
- Return type:
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(rng=0, n=10) >>> boxes = self.to_boxes() >>> sub_boxes = [d.to_boxes() for d in self.data] >>> areas1 = np.array([s.intersection(boxes).area[0] for s in sub_boxes]) >>> areas2 = np.array([s.area[0] for s in sub_boxes]) >>> assert np.allclose(areas1, areas2)
- box()[source]¶
Returns an axis-aligned bounding box for the segmentation
- Returns:
kwimage.Box
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(rng=0, n=10) >>> boxes = self.box() >>> sub_boxes = [d.box() for d in self.data] >>> areas1 = np.array([s.intersection(boxes).area for s in sub_boxes]) >>> areas2 = np.array([s.area for s in sub_boxes]) >>> assert np.allclose(areas1, areas2)
- to_mask(dims=None, pixels_are='points')[source]¶
Returns a mask object indication regions occupied by this multipolygon
- Returns:
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> s = 100 >>> self = MultiPolygon.random(rng=0).scale(s) >>> dims = (s, s) >>> mask = self.to_mask(dims) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> ax = plt.gca() >>> ax.set_xlim(0, s) >>> ax.set_ylim(0, s) >>> self.draw(color='red', alpha=.4) >>> mask.draw(color='blue', alpha=.4)
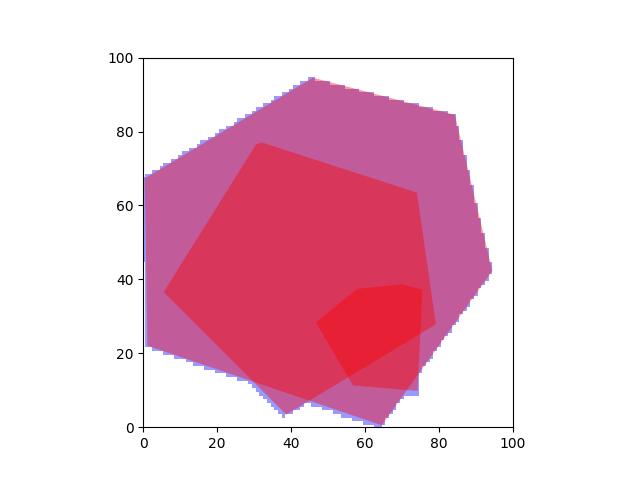
- to_relative_mask(return_offset=False)[source]¶
Returns a translated mask such the mask dimensions are minimal.
In other words, we move the polygon all the way to the top-left and return a mask just big enough to fit the polygon.
- Returns:
kwimage.Mask
- classmethod coerce(data, dims=None)[source]¶
Attempts to construct a MultiPolygon instance from the input data
See Segmentation.coerce
- Returns:
None | MultiPolygon
Example
>>> import kwimage >>> dims = (32, 32) >>> kw_poly = kwimage.Polygon.random().scale(dims) >>> kw_multi_poly = kwimage.MultiPolygon.random().scale(dims) >>> forms = [kw_poly, kw_multi_poly] >>> forms.append(kw_poly.to_shapely()) >>> forms.append(kw_poly.to_mask((32, 32))) >>> forms.append(kw_poly.to_geojson()) >>> forms.append(kw_poly.to_coco(style='orig')) >>> forms.append(kw_poly.to_coco(style='new')) >>> forms.append(kw_multi_poly.to_shapely()) >>> forms.append(kw_multi_poly.to_mask((32, 32))) >>> forms.append(kw_multi_poly.to_geojson()) >>> forms.append(kw_multi_poly.to_coco(style='orig')) >>> forms.append(kw_multi_poly.to_coco(style='new')) >>> for data in forms: >>> result = kwimage.MultiPolygon.coerce(data, dims=dims) >>> assert isinstance(result, kwimage.MultiPolygon)
- to_shapely(fix=False)[source]¶
- Parameters:
fix (bool) – if True, will check for validity and if any simple fixes can be applied, otherwise it returns the data as is.
- Returns:
shapely.geometry.MultiPolygon
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:shapely) >>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(rng=0) >>> geom = self.to_shapely() >>> print('geom = {!r}'.format(geom))
- classmethod from_shapely(geom)[source]¶
Convert a shapely polygon or multipolygon to a kwimage.MultiPolygon
- Parameters:
geom (shapely.geometry.MultiPolygon | shapely.geometry.Polygon)
- Returns:
MultiPolygon
Example
>>> import kwimage >>> sh_poly = kwimage.Polygon.random().to_shapely() >>> sh_multi_poly = kwimage.MultiPolygon.random().to_shapely() >>> kwimage.MultiPolygon.from_shapely(sh_poly) >>> kwimage.MultiPolygon.from_shapely(sh_multi_poly)
- classmethod from_geojson(data_geojson)[source]¶
Convert a geojson polygon or multipolygon to a kwimage.MultiPolygon
- Parameters:
data_geojson (Dict) – geojson data
- Returns:
MultiPolygon
Example
>>> import kwimage >>> orig = kwimage.MultiPolygon.random() >>> data_geojson = orig.to_geojson() >>> self = kwimage.MultiPolygon.from_geojson(data_geojson)
- classmethod from_coco(data, dims=None)[source]¶
Accepts either new-style or old-style coco multi-polygons
- Parameters:
data (List[List[Number] | Dict]) – a new or old style coco multi polygon
dims (None | Tuple[int, …]) – the shape dimensions of the canvas. Unused. Exists for compatibility with masks.
- Returns:
MultiPolygon
- to_coco(style='orig')[source]¶
- Parameters:
style (str) – can be “orig” or “new”
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = MultiPolygon.random(1, rng=0) >>> self.to_coco()
- class kwimage.Points(data=None, meta=None, datakeys=None, metakeys=None, **kwargs)[source]¶
Bases:
Spatial,_PointsWarpMixinStores multiple keypoints for a single object.
This stores both the geometry and the class metadata if available
Example
>>> from kwimage.structs.points import * # NOQA >>> xy = np.random.rand(10, 2) >>> pts = Points(xy=xy) >>> print('pts = {!r}'.format(pts))
- property shape¶
- property xy¶
- classmethod random(num=1, classes=None, rng=None)[source]¶
Makes random points; typically for testing purposes
Example
>>> import kwimage >>> self = kwimage.Points.random(classes=[1, 2, 3]) >>> self.data >>> print('self.data = {!r}'.format(self.data))
- property _impl¶
- tensor(device=NoParam)[source]¶
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10) >>> self.tensor()
- round(inplace=False)[source]¶
Rounds data to the nearest integer
- Parameters:
inplace (bool) – if True, modifies this object
Example
>>> import kwimage >>> self = kwimage.Points.random(3).scale(10) >>> self.round()
- numpy()[source]¶
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10) >>> self.tensor().numpy().tensor().numpy()
- draw_on(image=None, color='white', radius=None, copy=False)[source]¶
- Parameters:
image (ndarray) – image to draw points on.
color (str | Any | List[Any]) – one color for all boxes or a list of colors for each box Can be any type accepted by kwimage.Color.coerce. Extended types: str | ColorLike | List[ColorLike]
radius (None | int) – if an integer, an circle is drawn at each xy point with this radius. if None, attempts to fill a single point with subpixel accuracy, which generally means 4 pixels will be given some weight. Note: color can only be a single value for all points in this case.
copy (bool) – if True, force a copy of the image, otherwise try to draw inplace (may not work depending on dtype).
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/points.py Points.draw_on --show
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> s = 128 >>> image = np.zeros((s, s)) >>> self = Points.random(10).scale(s) >>> image = self.draw_on(image) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.autompl() >>> kwplot.imshow(image) >>> self.draw(radius=3, alpha=.5) >>> kwplot.show_if_requested()
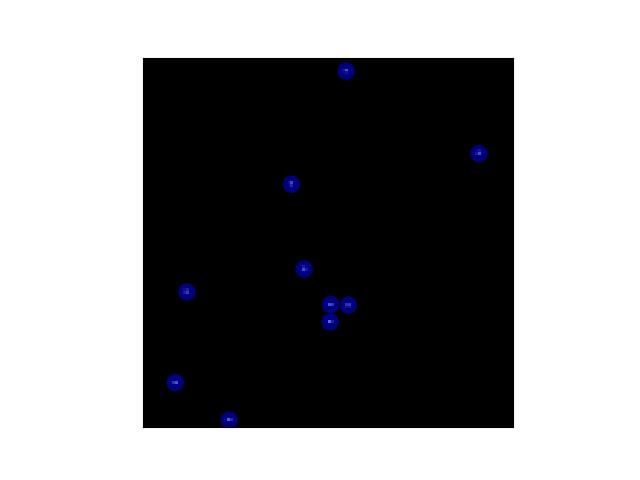

Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> s = 128 >>> image = np.zeros((s, s)) >>> self = Points.random(10).scale(s) >>> image = self.draw_on(image, radius=3, color='distinct') >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.autompl() >>> kwplot.imshow(image) >>> #self.draw(radius=3, alpha=.5, color='classes') >>> kwplot.show_if_requested()
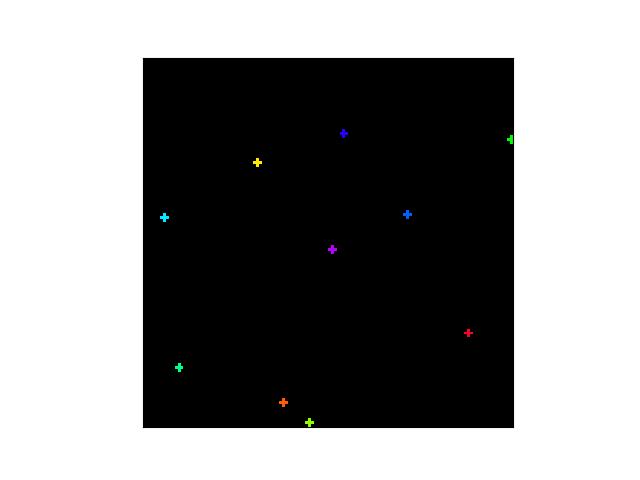

Example
>>> import kwimage >>> s = 32 >>> self = kwimage.Points.random(10).scale(s) >>> color = 'kitware_green' >>> # Test drawing on all channel + dtype combinations >>> im3 = np.zeros((s, s, 3), dtype=np.float32) >>> im_chans = { >>> 'im3': im3, >>> 'im1': kwimage.convert_colorspace(im3, 'rgb', 'gray'), >>> 'im4': kwimage.convert_colorspace(im3, 'rgb', 'rgba'), >>> } >>> inputs = {} >>> for k, im in im_chans.items(): >>> inputs[k + '_01'] = (kwimage.ensure_float01(im.copy()), {'radius': None}) >>> inputs[k + '_255'] = (kwimage.ensure_uint255(im.copy()), {'radius': None}) >>> outputs = {} >>> for k, v in inputs.items(): >>> im, kw = v >>> outputs[k] = self.draw_on(im, color=color, **kw) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=2, doclf=True) >>> plt = kwplot.autoplt() >>> pnum_ = kwplot.PlotNums(nRows=2, nSubplots=len(inputs)) >>> for k in inputs.keys(): >>> kwplot.imshow(outputs[k], fnum=2, pnum=pnum_(), title=k) >>> plt.gcf().suptitle('Test draw points on channel + dtype combos') >>> kwplot.show_if_requested()
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10).scale(32) >>> image = self.draw_on(radius=3, color='distinct') >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image) >>> kwplot.show_if_requested()
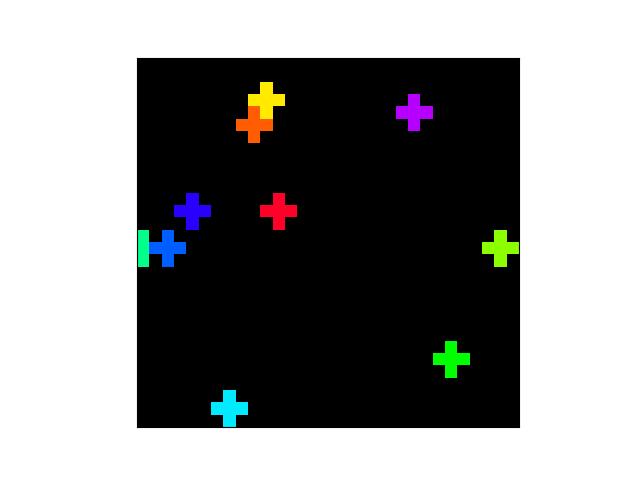

Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # Test cases where single and multiple colors are given >>> # with radius=None and radius=scalar >>> from kwimage.structs.points import * # NOQA >>> import kwimage >>> self = kwimage.Points.random(10).scale(32) >>> image1 = self.draw_on(radius=2, color='blue') >>> image2 = self.draw_on(radius=None, color='blue') >>> image3 = self.draw_on(radius=2, color='distinct') >>> image4 = self.draw_on(radius=None, color='distinct') >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> canvas = kwimage.stack_images_grid( >>> [image1, image2, image3, image4], >>> pad=3, bg_value=(1, 1, 1)) >>> kwplot.autompl() >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()
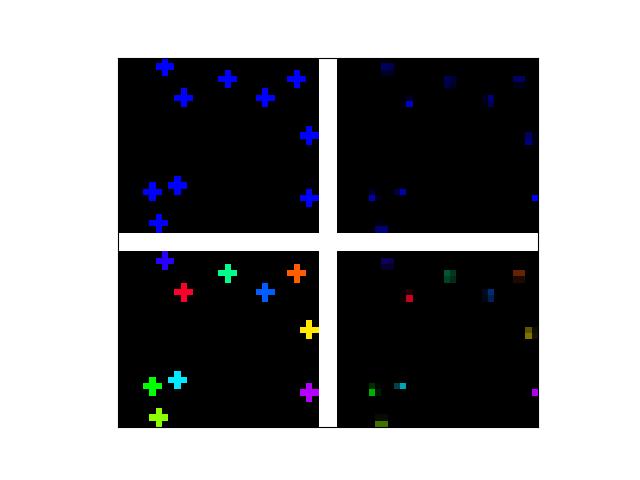
- draw(color='blue', ax=None, alpha=None, radius=1, setlim=False, **kwargs)[source]¶
TODO: can use kwplot.draw_points
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.points import * # NOQA >>> pts = Points.random(10) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(doclf=1) >>> pts.draw(radius=0.01) >>> from kwimage.structs.points import * # NOQA >>> self = Points.random(10, classes=['a', 'b', 'c']) >>> self.draw(radius=0.01, color='classes')
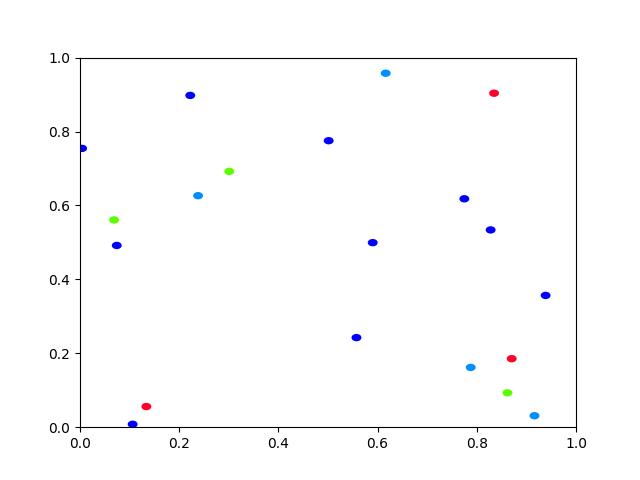
- compress(flags, axis=0, inplace=False)[source]¶
Filters items based on a boolean criterion
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4) >>> flags = [1, 0, 1, 1] >>> other = self.compress(flags) >>> assert len(self) == 4 >>> assert len(other) == 3
>>> # xdoctest: +REQUIRES(module:torch) >>> other = self.tensor().compress(flags) >>> assert len(other) == 3
- take(indices, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4) >>> indices = [1, 3] >>> other = self.take(indices) >>> assert len(self) == 4 >>> assert len(other) == 2
>>> # xdoctest: +REQUIRES(module:torch) >>> other = self.tensor().take(indices) >>> assert len(other) == 2
- to_coco(style='orig')[source]¶
Converts to an mscoco-like representation
Note
items that are usually id-references to other objects may need to be rectified.
- Parameters:
style (str) – either orig, new, new-id, or new-name
- Returns:
mscoco-like representation
- Return type:
Dict
Example
>>> from kwimage.structs.points import * # NOQA >>> self = Points.random(4, classes=['a', 'b']) >>> orig = self._to_coco(style='orig') >>> print('orig = {!r}'.format(orig)) >>> new_name = self._to_coco(style='new-name') >>> print('new_name = {}'.format(ub.urepr(new_name, nl=-1))) >>> # xdoctest: +REQUIRES(module:kwcoco) >>> import kwcoco >>> self.meta['classes'] = kwcoco.CategoryTree.coerce(self.meta['classes']) >>> new_id = self._to_coco(style='new-id') >>> print('new_id = {}'.format(ub.urepr(new_id, nl=-1)))
- classmethod from_coco(coco_kpts, class_idxs=None, classes=None, warn=False)[source]¶
- Parameters:
coco_kpts (list | dict) – either the original list keypoint encoding or the new dict keypoint encoding.
class_idxs (list) – only needed if using old style
classes (list | kwcoco.CategoryTree) – list of all keypoint category names
warn (bool) – if True raise warnings
Example
>>> ## >>> classes = ['mouth', 'left-hand', 'right-hand'] >>> coco_kpts = [ >>> {'xy': (0, 0), 'visible': 2, 'keypoint_category': 'left-hand'}, >>> {'xy': (1, 2), 'visible': 2, 'keypoint_category': 'mouth'}, >>> ] >>> Points.from_coco(coco_kpts, classes=classes) >>> # Test without classes >>> Points.from_coco(coco_kpts) >>> # Test without any category info >>> coco_kpts2 = [ub.dict_diff(d, {'keypoint_category'}) for d in coco_kpts] >>> Points.from_coco(coco_kpts2) >>> # Test without category instead of keypoint_category >>> coco_kpts3 = [ub.map_keys(lambda x: x.replace('keypoint_', ''), d) for d in coco_kpts] >>> Points.from_coco(coco_kpts3) >>> # >>> # Old style >>> coco_kpts = [0, 0, 2, 0, 1, 2] >>> Points.from_coco(coco_kpts) >>> # Fail case >>> coco_kpts4 = [{'xy': [4686.5, 1341.5], 'category': 'dot'}] >>> Points.from_coco(coco_kpts4, classes=[])
Example
>>> # xdoctest: +REQUIRES(module:kwcoco) >>> import kwcoco >>> classes = kwcoco.CategoryTree.from_coco([ >>> {'name': 'mouth', 'id': 2}, {'name': 'left-hand', 'id': 3}, {'name': 'right-hand', 'id': 5} >>> ]) >>> coco_kpts = [ >>> {'xy': (0, 0), 'visible': 2, 'keypoint_category_id': 5}, >>> {'xy': (1, 2), 'visible': 2, 'keypoint_category_id': 2}, >>> ] >>> pts = Points.from_coco(coco_kpts, classes=classes) >>> assert pts.data['class_idxs'].tolist() == [2, 0]
- class kwimage.PointsList(data, meta=None)[source]¶
Bases:
ObjectListStores a list of Points, each item usually corresponds to a different object.
Note
# TODO: when the data is homogenous we can use a more efficient # representation, otherwise we have to use heterogenous storage.
- class kwimage.Polygon(data=None, meta=None, datakeys=None, metakeys=None, **kwargs)[source]¶
Bases:
Spatial,_PolyArrayBackend,_PolyWarpMixin,_ShapelyMixin,NiceReprRepresents a single polygon as set of exterior boundary points and a list of internal polygons representing holes.
By convention exterior boundaries should be counterclockwise and interior holes should be clockwise.
Example
>>> import kwimage >>> poly1 = kwimage.Polygon(exterior=[[ 5., 10.], [ 1., 8.], [ 3., 4.], [ 5., 3.], [ 8., 9.], [ 6., 10.]]) >>> poly2 = kwimage.Polygon.random(rng=34214, n_holes=2).scale(10).round() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(doclf=True) >>> poly1.draw(setlim=1.4, vertex=0.2, vertexcolor='kw_orange', color='kw_blue', edgecolor='kw_green', alpha=0.5) >>> poly2.draw(setlim=2.9, vertex=0.2, vertexcolor='kw_red', color='kw_darkgreen', edgecolor='kw_darkblue', alpha=0.5) >>> kwplot.show_if_requested()
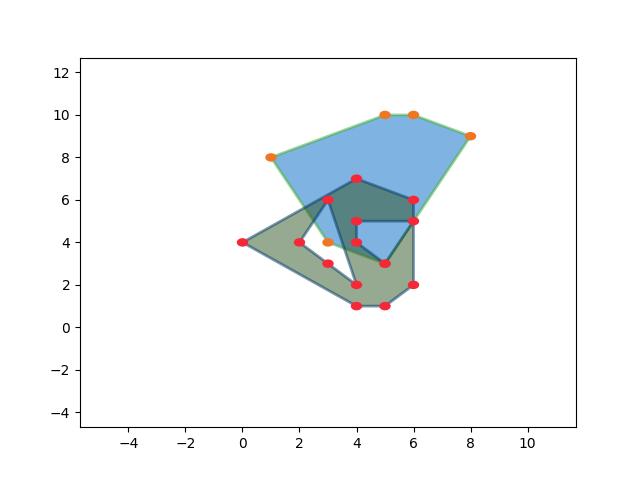
Example
>>> import kwimage >>> data = { >>> 'exterior': np.array([[13, 1], [13, 19], [25, 19], [25, 1]]), >>> 'interiors': [ >>> np.array([[14, 13], [15, 12], [23, 12], [24, 13], [24, 18], >>> [23, 19], [13, 19], [12, 18]]), >>> np.array([[13, 2], [14, 1], [24, 1], [25, 2], [25, 11], >>> [24, 12], [14, 12], [13, 11]])] >>> } >>> self = kwimage.Polygon(**data) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=1.4, vertex=0.2, vertexcolor='kw_orange', color='kw_blue', edgecolor='kw_green')
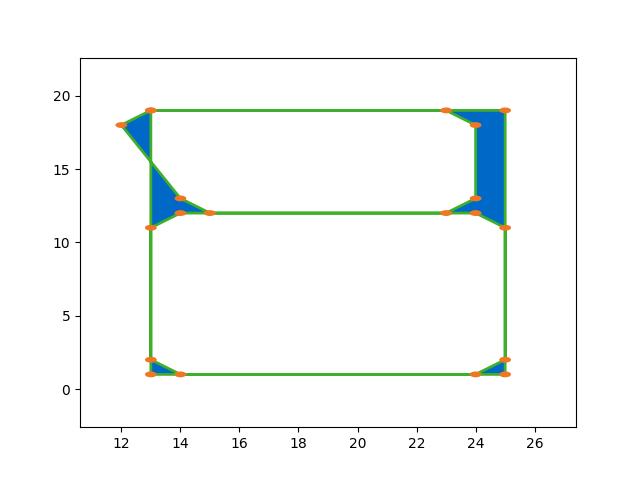
Example
>>> import kwimage >>> data = { >>> 'exterior': np.array([[13, 1], [13, 19], [25, 19], [25, 1]]), >>> 'interiors': [ >>> np.array([[14, 13], [15, 12], [23, 12], [24, 13], [24, 18], >>> [23, 19], [13, 19], [12, 18]]), >>> np.array([[13, 2], [14, 1], [24, 1], [25, 2], [25, 11], >>> [24, 12], [14, 12], [13, 11]])] >>> } >>> self = kwimage.Polygon(**data) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=1.4, vertex=0.2, vertexcolor='kw_orange', color='kw_blue', edgecolor='kw_green')
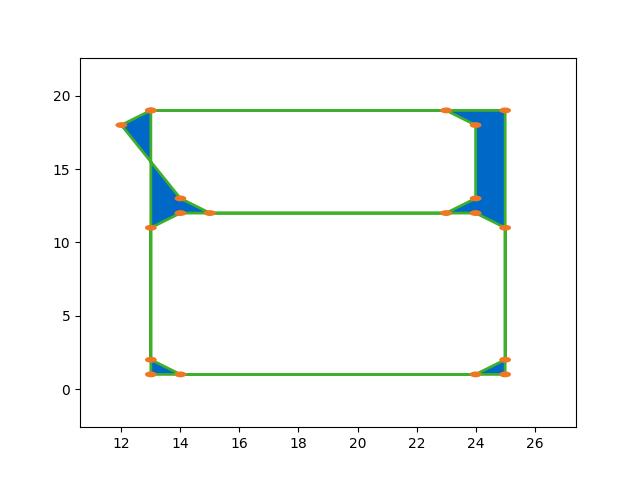
Example
>>> import kwimage >>> self = kwimage.Polygon.random( >>> n=5, n_holes=1, convex=False, rng=0) >>> print('self = {}'.format(self)) self = <Polygon({ 'exterior': <Coords(data= array([[0.30371392, 0.97195856], [0.24372304, 0.60568445], [0.21408694, 0.34884262], [0.5799477 , 0.44020379], [0.83720288, 0.78367234]]))>, 'interiors': [<Coords(data= array([[0.50164209, 0.83520279], [0.25835064, 0.40313428], [0.28778562, 0.74758761], [0.30341266, 0.93748088]]))>], })> >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=True)
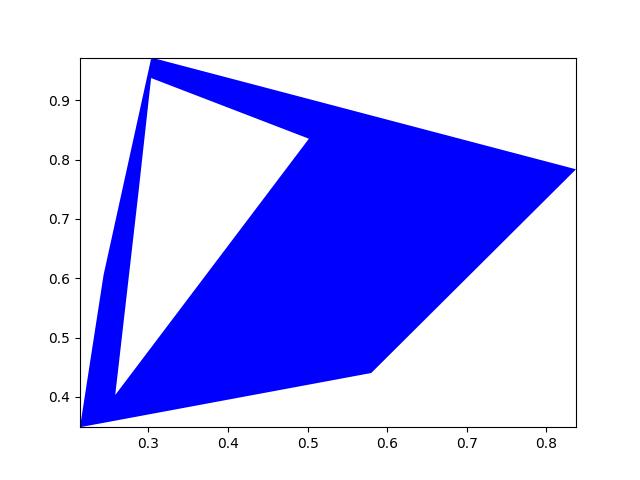
Example
>>> # Test empty polygon >>> import kwimage >>> data = { >>> 'exterior': np.array([]), >>> 'interiors': [],} >>> self = kwimage.Polygon(**data) >>> geos = self.to_geojson() >>> kwimage.Polygon.from_geojson(geos) >>> geom = self.to_shapely() >>> kwimage.Polygon.from_shapely(geom) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw(setlim=True)
- property exterior¶
Returns: kwimage.Coords
- property interiors¶
Returns: List[kwimage.Coords]
- classmethod circle(xy=(0, 0), r=1.0, resolution=64)[source]¶
Create a circular or elliptical polygon.
Might rename to ellipse later?
- Parameters:
xy (Iterable[Number]) – x and y center coordinate
r (float | Number | Tuple[Number, Number]) – circular radius or major and minor elliptical radius
resolution (int) – number of sides
- Returns:
Polygon
Example
>>> import kwimage >>> xy = (0.5, 0.5) >>> r = .3 >>> # Demo with circle >>> circle = kwimage.Polygon.circle(xy, r, resolution=6) >>> # Demo with ellipse >>> xy = (0.5, 0.5) >>> r = (.4, .7) >>> ellipse1 = kwimage.Polygon.circle(xy, r, resolution=12) >>> ellipse2 = kwimage.Polygon.circle(xy, (.7, .4), resolution=12) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> circle.draw(setlim=True, border=1, fill=0, color='kitware_orange') >>> ellipse1.draw(setlim=True, border=1, fill=0, color='kitware_blue') >>> ellipse2.draw(setlim=True, border=1, fill=0, color='kitware_green') >>> plt.gca().set_xlim(-0.5, 1.5) >>> plt.gca().set_ylim(-0.5, 1.5) >>> plt.gca().set_aspect('equal')
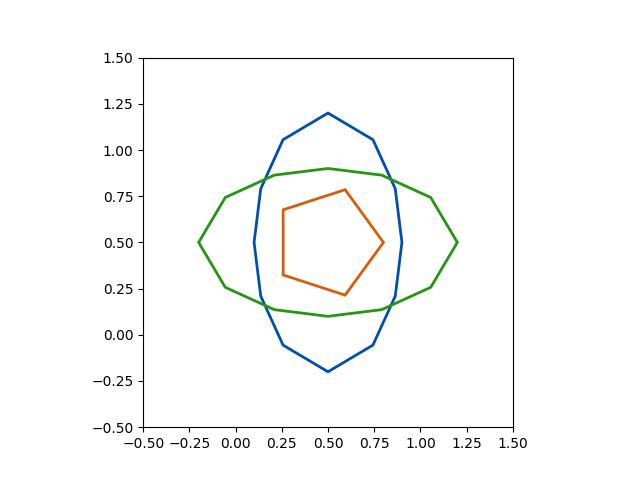
kwimage.Polygon.circle(xy, r, resolution=10).draw()
- classmethod regular(num, xy=(0, 0), r=1)[source]¶
Make a regular polygon with
numsides.Example
>>> import kwimage >>> n_polys = [ >>> kwimage.Polygon.regular(n) >>> for n in range(3, 11) >>> ] >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> fig =kwplot.figure(fnum=1, doclf=True) >>> ax = fig.gca() >>> for i, poly in enumerate(n_polys): >>> poly.translate((i * 2.5, 0), inplace=True) >>> poly.draw(border=True, fill=False) >>> ax.set_aspect('equal') >>> ax.set_xlim(-1, 8 * 2.5) >>> ax.set_ylim(-1, 1)
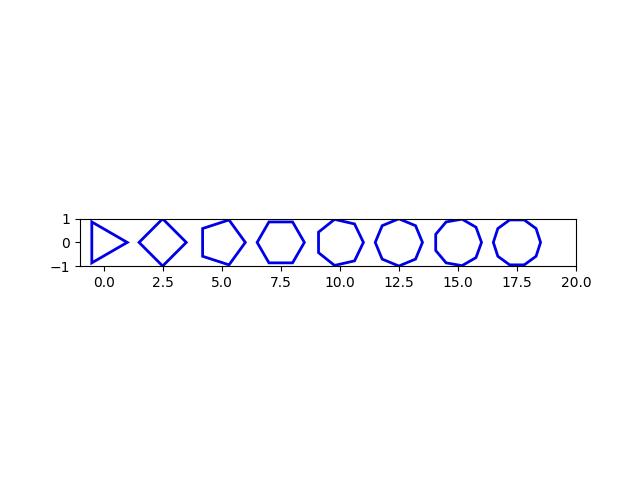
- classmethod star(xy=(0, 0), r=1)[source]¶
Make a star polygon
Example
>>> import kwimage >>> poly = kwimage.Polygon.star() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> fig = kwplot.figure(fnum=1, doclf=True) >>> ax = fig.gca() >>> poly.draw() >>> ax.set_aspect('equal') >>> ax.set_xlim(-1, 1) >>> ax.set_ylim(-1, 1)
- classmethod random(n=6, n_holes=0, convex=True, tight=False, rng=None)[source]¶
- Parameters:
n (int) – number of points in the polygon (must be 3 or more)
n_holes (int) – number of holes
tight (bool) – fits the minimum and maximum points between 0 and 1
convex (bool) – force resulting polygon will be convex (may remove exterior points)
- Returns:
Polygon
CommandLine
xdoctest -m kwimage.structs.polygon Polygon.random
Example
>>> import kwimage >>> self = kwimage.Polygon.random(n=4, rng=None, n_holes=2, convex=1) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> self.draw(color='kw_green', edgecolor='kw_blue', vertexcolor='kw_darkblue', vertex=0.01)
References
https://gis.stackexchange.com/questions/207731/random-multipolygon https://stackoverflow.com/questions/8997099/random-polygon https://stackoverflow.com/questions/27548363/from-voronoi-tessellation-to-shapely-polygons https://stackoverflow.com/questions/8997099/algorithm-to-generate-random-2d-polygon
- property _impl¶
- to_mask(dims=None, pixels_are='points')[source]¶
Convert this polygon to a mask
Todo
[ ] currently not efficient
- Parameters:
dims (Tuple) – height and width of the output mask
pixels_are (str) – either “points” or “areas”
- Returns:
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1).scale(128) >>> mask = self.to_mask((128, 128)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> mask.draw(color='blue') >>> mask.to_multi_polygon().draw(color='red', alpha=.5)
- to_relative_mask(return_offset=False)[source]¶
Returns a translated mask such the mask dimensions are minimal.
In other words, we move the polygon all the way to the top-left and return a mask just big enough to fit the polygon.
- Returns:
kwimage.Mask
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random().scale(8).translate(100, 100) >>> mask = self.to_relative_mask() >>> assert mask.shape <= (8, 8) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> mask.draw(color='blue') >>> mask.to_multi_polygon().draw(color='red', alpha=.5)
- _to_cv_countours()[source]¶
OpenCV polygon representation, which is a list of points. Holes are implicitly represented. When another polygon is drawn over an existing polyon via cv2.fillPoly
- Returns:
- where each ndarray is of shape [N, 1, 2],
where N is the number of points on the boundary, the middle dimension is always 1, and the trailing dimension represents x and y coordinates respectively.
- Return type:
List[ndarray]
- classmethod coerce(data)[source]¶
Routes the input to the proper constructor
Try to autodetermine format of input polygon and coerce it into a kwimage.Polygon.
- Parameters:
data (object) – some type of data that can be interpreted as a polygon.
- Returns:
kwimage.Polygon
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> kwimage.Polygon.coerce(self) >>> kwimage.Polygon.coerce(self.exterior) >>> kwimage.Polygon.coerce(self.exterior.data) >>> kwimage.Polygon.coerce(self.data) >>> kwimage.Polygon.coerce(self.to_geojson()) >>> kwimage.Polygon.coerce('POLYGON ((0.11 0.61, 0.07 0.588, 0.015 0.50, 0.11 0.61))')
- classmethod from_shapely(geom)[source]¶
Convert a shapely polygon to a kwimage.Polygon
- Parameters:
geom (shapely.geometry.polygon.Polygon) – a shapely polygon
- Returns:
kwimage.Polygon
- classmethod from_wkt(data)[source]¶
Convert a WKT string to a kwimage.Polygon
- Parameters:
data (str) – a WKT polygon string
- Returns:
kwimage.Polygon
Example
>>> import kwimage >>> data = 'POLYGON ((0.11 0.61, 0.07 0.588, 0.015 0.50, 0.11 0.61))' >>> self = kwimage.Polygon.from_wkt(data) >>> assert len(self.exterior) == 4
- classmethod from_geojson(data_geojson)[source]¶
Convert a geojson polygon to a kwimage.Polygon
- Parameters:
data_geojson (dict) – geojson data
- Returns:
Polygon
References
https://geojson.org/geojson-spec.html
Example
>>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=2) >>> data_geojson = self.to_geojson() >>> new = Polygon.from_geojson(data_geojson)
- to_shapely(fix=False)[source]¶
- Parameters:
fix (bool) – if True, will check for validity and if any simple fixes can be applied, otherwise it returns the data as is.
- Returns:
shapely.geometry.polygon.Polygon
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:shapely) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1) >>> self = self.scale(100) >>> geom = self.to_shapely() >>> print('geom = {!r}'.format(geom))
- to_geojson()[source]¶
Converts polygon to a geojson structure
- Returns:
Dict[str, object]
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> print(self.to_geojson())
- to_wkt()[source]¶
Convert a kwimage.Polygon to WKT string
- Returns:
str
Example
>>> import kwimage >>> self = kwimage.Polygon.random() >>> print(self.to_wkt())
- classmethod from_coco(data, dims=None)[source]¶
Accepts either new-style or old-style coco polygons
- Parameters:
data (List[Number] | Dict) – A new or old-style coco polygon
dims (None | Tuple[int, …]) – the shape dimensions of the canvas. Unused. Exists for compatibility with masks.
- Returns:
Polygon
- to_coco(style='orig')[source]¶
- Parameters:
style (str) – can be “orig” or “new”
- Returns:
coco-style polygons
- Return type:
List | Dict
- property centroid¶
Returns: Tuple[Number, Number]
- to_box()[source]¶
## DEPRECATED: Use
box()instead. ## Do we deprecate this? Should we stick to the to_ / from_ convention?- Returns:
kwimage.Box
- bounding_box()[source]¶
Returns an axis-aligned bounding box for the segmentation
DEPRECATED: Use singular
box()instead.- Returns:
kwimage.Boxes
- box()[source]¶
Returns an axis-aligned bounding box for the segmentation
- Returns:
kwimage.Box
Example
>>> import kwimage >>> poly = kwimage.Polygon.random() >>> box = poly.box() >>> print('box = {}'.format(ub.urepr(box, nl=1)))
- bounding_box_polygon()[source]¶
Returns an axis-aligned bounding polygon for the segmentation.
Note
This Polygon will be a Box, not a convex hull! Use shapely for convex hulls.
- Returns:
kwimage.Polygon
- clip(x_min, y_min, x_max, y_max, inplace=False)[source]¶
Clip polygon to specified boundaries.
- Returns:
clipped polygon
- Return type:
Example
>>> from kwimage.structs.polygon import * >>> self = Polygon.random().scale(10).translate(-1) >>> self2 = self.clip(1, 1, 3, 3) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self2.draw(setlim=True)
- fill(image, value=1, pixels_are='points', assert_inplace=False)[source]¶
Fill in an image based on this polyon.
- Parameters:
image (ndarray) – image to draw on
value (int | Tuple[int]) – value fill in with. Defaults to 1.
pixels_are (str) – either points or areas
assert_inplace (bool) – if True then the function will error if the modification cannot happen inplace.
- Returns:
the image that has been modified in place
- Return type:
ndarray
Example
>>> # xdoctest: +REQUIRES(module:rasterio) >>> import kwimage >>> mask = kwimage.Mask.random(rng=0) >>> self = mask.to_multi_polygon(pixels_are='areas').data[0] >>> image = np.zeros_like(mask.data) >>> self.fill(image, pixels_are='areas')
Example
>>> # Test case where there are multiple channels >>> import kwimage >>> mask = kwimage.Mask.random(shape=(4, 4), rng=0) >>> self = mask.to_multi_polygon() >>> image = np.zeros(mask.shape[0:2] + (2,), dtype=np.float32) >>> fill_v1 = self.fill(image.copy(), value=1) >>> fill_v2 = self.fill(image.copy(), value=(1, 2)) >>> assert np.all((fill_v1 > 0) == (fill_v2 > 0))
Example
>>> import kwimage >>> # Test dtype with inplace vs not >>> mask = kwimage.Mask.random(shape=(32, 32), rng=0) >>> self = mask.to_multi_polygon() >>> native_dtypes = [] >>> native_dtypes += [np.uint8, np.uint16] >>> native_dtypes += [np.int8, np.int16, np.int32] >>> native_dtypes += [np.float32] >>> for dtype in native_dtypes: >>> image = np.zeros(mask.shape[0:2] + (2,), dtype=dtype) >>> image1 = self.fill(image, value=1, assert_inplace=True) >>> assert image1.sum() > 0 >>> assert image.sum() > 0 >>> print(f'dtype: {dtype} inplace') >>> needfix_dtypes = [np.uint32, np.uint64, np.int64, np.float16, np.float64] >>> for dtype in needfix_dtypes: >>> image = np.zeros(mask.shape[0:2] + (2,), dtype=dtype) >>> image1 = self.fill(image, value=1, assert_inplace=False) >>> assert image1.sum() > 0 >>> assert image.sum() == 0 >>> print(f'dtype: {dtype} not inplace')
- draw_on(image, color='blue', fill=True, border=False, alpha=1.0, edgecolor=None, facecolor=None, copy=False)[source]¶
Rasterizes a polygon on an image. See draw for a vectorized matplotlib version.
- Parameters:
image (ndarray) – image to raster polygon on.
color (str | tuple) – data coercable to a color
fill (bool) – draw the center mass of the polygon. Note: this will be deprecated. Use facecolor instead.
border (bool) – draw the border of the polygon Note: this will be deprecated. Use edgecolor instead.
alpha (float) – polygon transparency (setting alpha < 1 makes this function much slower). Defaults to 1.0
copy (bool) – if False only copies if necessary
edgecolor (str | tuple) – color for the border
facecolor (str | tuple) – color for the fill
- Returns:
np.ndarray
Note
This function will only be inplace if alpha=1.0 and the input has 3 or 4 channels. Otherwise the output canvas is coerced so colors can be drawn on it. In the case where alpha < 1.0,
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1).scale(128) >>> image_in = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image_in) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image_out, fnum=1)
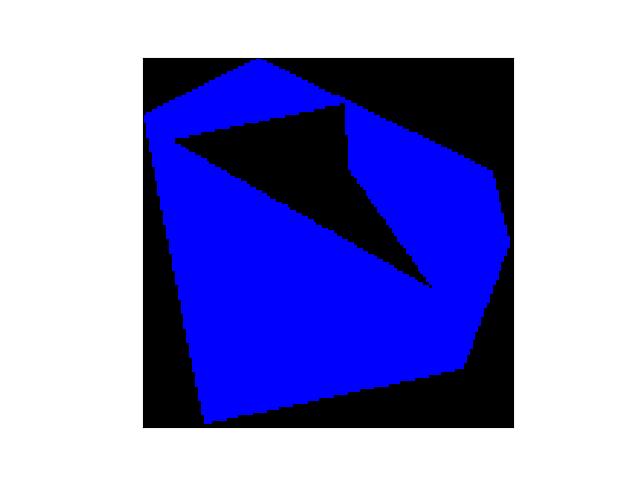

Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # Demo drawing on a RGBA canvas >>> # If you initialize an zero rgba canvas, the alpha values are >>> # filled correctly. >>> from kwimage.structs.polygon import * # NOQA >>> s = 16 >>> self = Polygon.random(n_holes=1, rng=32).scale(s) >>> image_in = np.zeros((s, s, 4), dtype=np.float32) >>> image_out = self.draw_on(image_in, color='black') >>> assert np.all(image_out[..., 0:3] == 0) >>> assert not np.all(image_out[..., 3] == 1) >>> assert not np.all(image_out[..., 3] == 0)
Example
>>> import kwimage >>> color = 'blue' >>> self = kwimage.Polygon.random(n_holes=1).scale(128) >>> image = np.zeros((128, 128), dtype=np.float32) >>> # Test drawing on all channel + dtype combinations >>> im3 = np.random.rand(128, 128, 3) >>> im_chans = { >>> 'im3': im3, >>> 'im1': kwimage.convert_colorspace(im3, 'rgb', 'gray'), >>> #'im0': im3[..., 0], >>> 'im4': kwimage.convert_colorspace(im3, 'rgb', 'rgba'), >>> } >>> inputs = {} >>> for k, im in im_chans.items(): >>> inputs[k + '_f01'] = (kwimage.ensure_float01(im.copy()), {'alpha': None}) >>> inputs[k + '_u255'] = (kwimage.ensure_uint255(im.copy()), {'alpha': None}) >>> inputs[k + '_f01_a'] = (kwimage.ensure_float01(im.copy()), {'alpha': 0.5}) >>> inputs[k + '_u255_a'] = (kwimage.ensure_uint255(im.copy()), {'alpha': 0.5}) >>> # Check cases when image is/isnot written inplace Construct images >>> # with different dtypes / channels and run a draw_on with different >>> # keyword args. For each combination, demo if that results in an >>> # implace operation or not. >>> rows = [] >>> outputs = {} >>> for k, v in inputs.items(): >>> im, kw = v >>> outputs[k] = self.draw_on(im, color=color, **kw) >>> inplace = outputs[k] is im >>> rows.append({'key': k, 'inplace': inplace}) >>> # xdoctest: +REQUIRES(module:pandas) >>> import pandas as pd >>> df = pd.DataFrame(rows).sort_values('inplace') >>> print(df.to_string()) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.figure(fnum=2, doclf=True) >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nCols=2, nRows=len(inputs)) >>> for k in inputs.keys(): >>> kwplot.imshow(inputs[k][0], fnum=2, pnum=pnum_(), title=k) >>> kwplot.imshow(outputs[k], fnum=2, pnum=pnum_(), title=k) >>> kwplot.show_if_requested()
Example
>>> # Test empty polygon draw >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.from_coco([]) >>> image_in = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image_in)
Example
>>> # Test stupid large polygon draw >>> from kwimage.structs.polygon import * # NOQA >>> from kwimage.structs.polygon import _generic >>> import kwimage >>> self = kwimage.Polygon.random().scale(2e11) >>> image = np.zeros((128, 128), dtype=np.float32) >>> image_out = self.draw_on(image)
- draw(color='blue', ax=None, alpha=1.0, radius=1, setlim=False, border=None, linewidth=None, edgecolor=None, facecolor=None, fill=True, vertex=False, vertexcolor=None)[source]¶
Draws polygon in a matplotlib axes. See draw_on for in-memory image modification.
- Parameters:
color (str | Tuple) – coercable color. Default color if specific colors are not given.
alpha (float) – fill transparency
fill (bool) – if True fill the polygon with facecolor, otherwise just draw the border if linewidth > 0
setlim (bool | str) – if True, modify the x and y limits of the matplotlib axes such that the polygon is can be seen. Can also be a string “grow”, which only allows growth of the viewport to accomidate the new polyogn.
border (bool) – if True, draws an edge border on the polygon. DEPRECATED. Use linewidth instead.
linewidth (bool) – width of the border
edgecolor (None | Any) – if None, uses the value of
color. Otherwise the color of the border when linewidth > 0. Extended types Coercible[kwimage.Color].facecolor (None | Any) – if None, uses the value of
color. Otherwise, color of the border when fill=True. Extended types Coercible[kwimage.Color].vertex (float) – if non-zero, draws vertexes on the polygon with this radius.
vertexcolor (Any) – color of vertexes Extended types Coercible[kwimage.Color].
- Returns:
None for am empty polygon
- Return type:
matplotlib.patches.PathPatch | None
Todo
[ ] Rework arguments in favor of matplotlib standards
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1) >>> self = self.scale(100) >>> # xdoctest: +REQUIRES(--show) >>> kwargs = dict(edgecolor='orangered', facecolor='dodgerblue', linewidth=10) >>> self.draw(**kwargs) >>> import kwplot >>> kwplot.autompl() >>> from matplotlib import pyplot as plt >>> kwplot.figure(fnum=2) >>> self.draw(setlim=True, **kwargs)
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> from kwimage.structs.polygon import * # NOQA >>> self = Polygon.random(n_holes=1, rng=33202) >>> import textwrap >>> # Test over a range of parameters >>> basis = { >>> 'linewidth': [0, 4], >>> 'edgecolor': [None, 'gold'], >>> 'facecolor': ['purple'], >>> 'fill': [True, False], >>> 'alpha': [1.0, 0.5], >>> 'vertex': [0, 0.01], >>> 'vertexcolor': ['green'], >>> } >>> grid = list(ub.named_product(basis)) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nSubplots=len(grid)) >>> for kwargs in grid: >>> fig = kwplot.figure(fnum=1, pnum=pnum_()) >>> ax = fig.gca() >>> self.draw(ax=ax, **kwargs) >>> title = ub.urepr(kwargs, compact=True) >>> title = '\n'.join(textwrap.wrap( >>> title.replace(',', ' '), break_long_words=False, >>> width=60)) >>> ax.set_title(title, fontdict={'fontsize': 8}) >>> ax.grid(False) >>> ax.set_xticks([]) >>> ax.set_yticks([]) >>> fig.subplots_adjust(wspace=0.5, hspace=0.3, bottom=0.001, top=0.97) >>> kwplot.show_if_requested()

- _ensure_vertex_order(inplace=False)[source]¶
Fixes vertex ordering so the exterior ring is CCW and the interior rings are CW.
Example
>>> import kwimage >>> self = kwimage.Polygon.random(n=3, n_holes=2, rng=0) >>> print('self = {!r}'.format(self)) >>> new = self._ensure_vertex_order() >>> print('new = {!r}'.format(new))
>>> self = kwimage.Polygon.random(n=3, n_holes=2, rng=0).swap_axes() >>> print('self = {!r}'.format(self)) >>> new = self._ensure_vertex_order() >>> print('new = {!r}'.format(new))
- morph(other, alpha)[source]¶
Perform polygon-to-polygon morphing.
Note
This current algorithm is very basic and does not yet prevent self-intersections in intermediate polygons.
- Parameters:
other (kwimage.Polygon) – the other polygon to morph into
alpha (float | List[float]) – A value between 0 and 1, indicating the fractional position of the new interpolated polygon between
selfandother. If given as a list multiple interpolations are returned.
- Returns:
one ore more interpolated polygons
- Return type:
Todo
[ ] Implement level set method [LevelSet] which rasterizes each
polygon, interpolates the raster, and computes the interpolated polygon as contours in that interpolated raster.
[ ] Implement methods from [Albrecht2006] and [PolygonMorph]
References
[HaudrenShapes]https://github.com/haudren/shapes/blob/master/shapes/shapes.py
[KamvysselisMorph]Example
>>> import kwimage >>> self = kwimage.Polygon.random(3, convex=0) >>> other = kwimage.Polygon.random(4, convex=0).translate((2, 2)) >>> results = self.morph(other, np.linspace(0, 1, 5)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(doclf=1) >>> self.draw(setlim='grow', color='kw_blue', alpha=0.5, vertex=0.02) >>> other.draw(setlim='grow', color='kw_green', alpha=0.5, vertex=0.02) >>> colors = kwimage.Color('kw_blue').morph( >>> 'kw_green', np.linspace(0, 1, 5)) >>> for new, c in zip(results, colors): >>> pt = new.exterior.data[0] >>> new.draw(color=c, alpha=0.5, vertex=0.01) >>> intepolation_lines = np.array([new.exterior.data for new in results]) >>> for interp_line in intepolation_lines.transpose(1, 0, 2)[::8]: >>> plt.plot(*interp_line.T, '--x')
- class kwimage.PolygonList(data, meta=None)[source]¶
Bases:
ObjectListStores and allows manipluation of multiple polygons, usually within the same image.
- to_mask_list(dims=None, pixels_are='points')[source]¶
Converts all items to masks
- Returns:
kwimage.MaskList
- to_segmentation_list()[source]¶
Converts all items to segmentation objects
- Returns:
kwimage.SegmentationList
- to_geojson(as_collection=False)[source]¶
Converts a list of polygons/multipolygons to a geojson structure
- Parameters:
as_collection (bool) – if True, wraps the polygon geojson items in a geojson feature collection, otherwise just return a list of items.
- Returns:
items or geojson data
- Return type:
List[Dict] | Dict
Example
>>> import kwimage >>> data = [kwimage.Polygon.random(), >>> kwimage.Polygon.random(n_holes=1), >>> kwimage.MultiPolygon.random(n_holes=1), >>> kwimage.MultiPolygon.random()] >>> self = kwimage.PolygonList(data) >>> geojson = self.to_geojson(as_collection=True) >>> items = self.to_geojson(as_collection=False) >>> print('geojson = {}'.format(ub.urepr(geojson, nl=-2, precision=1))) >>> print('items = {}'.format(ub.urepr(items, nl=-2, precision=1)))
- class kwimage.Projective(matrix)[source]¶
Bases:
LinearA thin wraper around a 3x3 matrix that represent a projective transform
- Implements methods for:
creating random projective transforms
decomposing the matrix
finding a best-fit transform between corresponding points
TODO: - [ ] fully rational transform
Example
>>> import kwimage >>> import math >>> image = kwimage.grab_test_image() >>> theta = 0.123 * math.tau >>> components = { >>> 'rotate': kwimage.Projective.projective(theta=theta), >>> 'scale': kwimage.Projective.projective(scale=0.5), >>> 'shear': kwimage.Projective.projective(shearx=0.2), >>> 'translation': kwimage.Projective.projective(offset=(100, 200)), >>> 'rotate+translate': kwimage.Projective.projective(theta=0.123 * math.tau, about=(256, 256)), >>> 'perspective': kwimage.Projective.projective(uv=(0.0003, 0.0007)), >>> 'random-composed': kwimage.Projective.random(scale=(0.5, 1.5), translate=(-20, 20), theta=(-theta, theta), shearx=(0, .4), rng=900558176210808600), >>> } >>> warp_stack = [] >>> for key, mat in components.items(): ... warp = kwimage.warp_projective(image, mat) ... warp = kwimage.draw_text_on_image( ... warp, ... ub.urepr(mat.matrix, nl=1, nobr=1, precision=4, si=1, sv=1, with_dtype=0), ... org=(1, 1), ... valign='top', halign='left', ... fontScale=0.8, color='kw_green', ... border={'thickness': 3}, ... ) ... warp = kwimage.draw_header_text(warp, key, color='kw_blue') ... warp_stack.append(warp) >>> warp_canvas = kwimage.stack_images_grid(warp_stack, chunksize=4, pad=10, bg_value='kitware_gray') >>> # xdoctest: +REQUIRES(module:sympy) >>> import sympy >>> # Shows the symbolic construction of the code >>> # https://groups.google.com/forum/#!topic/sympy/k1HnZK_bNNA >>> from sympy.abc import theta >>> params = x0, y0, sx, sy, theta, shearx, tx, ty, u, v = sympy.symbols( >>> 'x0, y0, sx, sy, theta, ex, tx, ty, u, v') >>> # move the center to 0, 0 >>> tr1_ = sympy.Matrix([[1, 0, -x0], >>> [0, 1, -y0], >>> [0, 0, 1]]) >>> P = sympy.Matrix([ # projective part >>> [ 1, 0, 0], >>> [ 0, 1, 0], >>> [ u, v, 1]]) >>> # Define core components of the affine transform >>> S = sympy.Matrix([ # scale >>> [sx, 0, 0], >>> [ 0, sy, 0], >>> [ 0, 0, 1]]) >>> E = sympy.Matrix([ # x-shear >>> [1, shearx, 0], >>> [0, 1, 0], >>> [0, 0, 1]]) >>> R = sympy.Matrix([ # rotation >>> [sympy.cos(theta), -sympy.sin(theta), 0], >>> [sympy.sin(theta), sympy.cos(theta), 0], >>> [ 0, 0, 1]]) >>> T = sympy.Matrix([ # translation >>> [ 1, 0, tx], >>> [ 0, 1, ty], >>> [ 0, 0, 1]]) >>> # move 0, 0 back to the specified origin >>> tr2_ = sympy.Matrix([[1, 0, x0], >>> [0, 1, y0], >>> [0, 0, 1]]) >>> # combine transformations >>> homog_ = sympy.MatMul(tr2_, T, R, E, S, P, tr1_) >>> #with sympy.evaluate(False): >>> # homog_ = sympy.MatMul(tr2_, T, R, E, S, P, tr1_) >>> # sympy.pprint(homog_) >>> homog = homog_.doit() >>> #sympy.pprint(homog) >>> print('homog = {}'.format(ub.urepr(homog.tolist(), nl=1))) >>> # This could be prettier >>> texts = { >>> 'Translation': sympy.pretty(R, use_unicode=0), >>> 'Rotation': sympy.pretty(R, use_unicode=0), >>> 'shEar-X': sympy.pretty(E, use_unicode=0), >>> 'Scale': sympy.pretty(S, use_unicode=0), >>> 'Perspective': sympy.pretty(P, use_unicode=0), >>> } >>> print(ub.urepr(texts, nl=2, sv=1)) >>> equation_stack = [] >>> for text, m in texts.items(): >>> render_canvas = kwimage.draw_text_on_image(None, m, color='kw_green', fontScale=1.0) >>> render_canvas = kwimage.draw_header_text(render_canvas, text, color='kw_blue') >>> render_canvas = kwimage.imresize(render_canvas, scale=1.3) >>> equation_stack.append(render_canvas) >>> equation_canvas = kwimage.stack_images(equation_stack, pad=10, axis=1, bg_value='kitware_gray') >>> render_canvas = kwimage.draw_text_on_image(None, sympy.pretty(homog, use_unicode=0), color='kw_green', fontScale=1.0) >>> render_canvas = kwimage.draw_header_text(render_canvas, 'Full Equation With Pre-Shift', color='kw_blue') >>> # xdoctest: -REQUIRES(module:sympy) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> canvas = kwimage.stack_images([warp_canvas, equation_canvas, render_canvas], pad=20, axis=0, bg_value='kitware_gray', resize='larger') >>> canvas = kwimage.draw_header_text(canvas, 'Projective matrixes can represent', color='kw_blue') >>> kwplot.imshow(canvas) >>> fig = plt.gcf() >>> fig.set_size_inches(13, 13)

- classmethod fit(pts1, pts2)[source]¶
Fit an projective transformation between a set of corresponding points.
See [HomogEst] [SzeleskiBook] and [RansacDummies] for references on the subject.
- Parameters:
pts1 (ndarray) – An Nx2 array of points in “space 1”.
pts2 (ndarray) – A corresponding Nx2 array of points in “space 2”
- Returns:
a transform that warps from “space1” to “space2”.
- Return type:
Note
A projective matrix has 8 degrees of freedome, so at least 8 point pairs are needed.
References
Example
>>> # Create a set of points, warp them, then recover the warp >>> import kwimage >>> points = kwimage.Points.random(9).scale(64) >>> A1 = kwimage.Affine.affine(scale=0.9, theta=-3.2, offset=(2, 3), about=(32, 32), skew=2.3) >>> A2 = kwimage.Affine.affine(scale=0.8, theta=0.8, offset=(2, 0), about=(32, 32)) >>> A12_real = A2 @ A1.inv() >>> points1 = points.warp(A1) >>> points2 = points.warp(A2) >>> # Make the correspondence non-affine >>> points2.data['xy'].data[0, 0] += 3.5 >>> points2.data['xy'].data[3, 1] += 8.5 >>> # Recover the warp >>> pts1, pts2 = points1.xy, points2.xy >>> A_recovered = kwimage.Projective.fit(pts1, pts2) >>> #assert np.all(np.isclose(A_recovered.matrix, A12_real.matrix)) >>> # xdoctest: +REQUIRES(--show) >>> import cv2 >>> import kwplot >>> kwplot.autompl() >>> base1 = np.zeros((96, 96, 3)) >>> base1[32:-32, 5:-5] = 0.5 >>> base2 = np.zeros((96, 96, 3)) >>> img1 = points1.draw_on(base1, radius=3, color='blue') >>> img2 = points2.draw_on(base2, radius=3, color='green') >>> img1_warp = kwimage.warp_projective(img1, A_recovered.matrix, dsize=img1.shape[0:2][::-1]) >>> canvas = kwimage.stack_images([img1, img2, img1_warp], pad=10, axis=1, bg_value=(1., 1., 1.)) >>> kwplot.imshow(canvas)
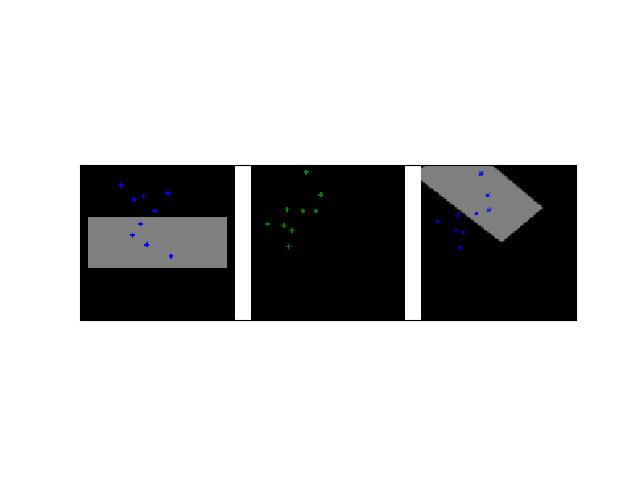

- classmethod projective(scale=None, offset=None, shearx=None, theta=None, uv=None, about=None)[source]¶
Reconstruct from parameters
Sympy
>>> # xdoctest: +SKIP >>> import sympy >>> # Shows the symbolic construction of the code >>> # https://groups.google.com/forum/#!topic/sympy/k1HnZK_bNNA >>> from sympy.abc import theta >>> params = x0, y0, sx, sy, theta, shearx, tx, ty, u, v = sympy.symbols( >>> 'x0, y0, sx, sy, theta, ex, tx, ty, u, v') >>> # move the center to 0, 0 >>> tr1_ = sympy.Matrix([[1, 0, -x0], >>> [0, 1, -y0], >>> [0, 0, 1]]) >>> P = sympy.Matrix([ # projective part >>> [ 1, 0, 0], >>> [ 0, 1, 0], >>> [ u, v, 1]]) >>> # Define core components of the affine transform >>> S = sympy.Matrix([ # scale >>> [sx, 0, 0], >>> [ 0, sy, 0], >>> [ 0, 0, 1]]) >>> E = sympy.Matrix([ # x-shear >>> [1, shearx, 0], >>> [0, 1, 0], >>> [0, 0, 1]]) >>> R = sympy.Matrix([ # rotation >>> [sympy.cos(theta), -sympy.sin(theta), 0], >>> [sympy.sin(theta), sympy.cos(theta), 0], >>> [ 0, 0, 1]]) >>> T = sympy.Matrix([ # translation >>> [ 1, 0, tx], >>> [ 0, 1, ty], >>> [ 0, 0, 1]]) >>> # move 0, 0 back to the specified origin >>> tr2_ = sympy.Matrix([[1, 0, x0], >>> [0, 1, y0], >>> [0, 0, 1]]) >>> # combine transformations >>> with sympy.evaluate(False): >>> homog_ = sympy.MatMul(tr2_, T, R, E, S, P, tr1_) >>> sympy.pprint(homog_) >>> homog = homog_.doit() >>> sympy.pprint(homog) >>> print('homog = {}'.format(ub.urepr(homog.tolist(), nl=1)))
- classmethod coerce(data=None, **kwargs)[source]¶
Attempt to coerce the data into an Projective object
- Parameters:
data – some data we attempt to coerce to an Projective matrix
**kwargs – some data we attempt to coerce to an Projective matrix, mutually exclusive with data.
- Returns:
Projective
Example
>>> import kwimage >>> kwimage.Projective.coerce({'type': 'affine', 'matrix': [[1, 0, 0], [0, 1, 0]]}) >>> kwimage.Projective.coerce({'type': 'affine', 'scale': 2}) >>> kwimage.Projective.coerce({'type': 'projective', 'scale': 2}) >>> kwimage.Projective.coerce({'scale': 2}) >>> kwimage.Projective.coerce({'offset': 3}) >>> kwimage.Projective.coerce(np.eye(3)) >>> kwimage.Projective.coerce(None) >>> import skimage >>> kwimage.Projective.coerce(skimage.transform.AffineTransform(scale=30)) >>> kwimage.Projective.coerce(skimage.transform.ProjectiveTransform(matrix=None))
- is_affine()[source]¶
If the bottom row is [[0, 0, 1]], then this can be safely turned into an affine matrix.
- Returns:
bool
Example
>>> import kwimage >>> kwimage.Projective.coerce(scale=2, uv=[1, 1]).is_affine() False >>> kwimage.Projective.coerce(scale=2, uv=[0, 0]).is_affine() True
- to_skimage()[source]¶
- Returns:
skimage.transform.AffineTransform
Example
>>> import kwimage >>> self = kwimage.Projective.random() >>> tf = self.to_skimage() >>> # Transform points with kwimage and scikit-image >>> kw_poly = kwimage.Polygon.random() >>> kw_warp_xy = kw_poly.warp(self.matrix).exterior.data >>> sk_warp_xy = tf(kw_poly.exterior.data) >>> assert np.allclose(sk_warp_xy, sk_warp_xy)
- classmethod random(shape=None, rng=None, **kw)[source]¶
- Example/
>>> import kwimage >>> self = kwimage.Projective.random() >>> print(f'self={self}') >>> params = self.decompose() >>> aff_part = kwimage.Affine.affine(**ub.dict_diff(params, ['uv'])) >>> proj_part = kwimage.Projective.coerce(uv=params['uv']) >>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(--show) >>> import cv2 >>> import kwplot >>> dsize = (256, 256) >>> kwplot.autompl() >>> img1 = kwimage.grab_test_image(dsize=dsize) >>> img1_affonly = kwimage.warp_projective(img1, aff_part.matrix, dsize=img1.shape[0:2][::-1]) >>> img1_projonly = kwimage.warp_projective(img1, proj_part.matrix, dsize=img1.shape[0:2][::-1]) >>> ### >>> img2 = kwimage.ensure_uint255(kwimage.atleast_3channels(kwimage.checkerboard(dsize=dsize))) >>> img1_fullwarp = kwimage.warp_projective(img1, self.matrix, dsize=img1.shape[0:2][::-1]) >>> img2_affonly = kwimage.warp_projective(img2, aff_part.matrix, dsize=img2.shape[0:2][::-1]) >>> img2_projonly = kwimage.warp_projective(img2, proj_part.matrix, dsize=img2.shape[0:2][::-1]) >>> img2_fullwarp = kwimage.warp_projective(img2, self.matrix, dsize=img2.shape[0:2][::-1]) >>> canvas1 = kwimage.stack_images([img1, img1_projonly, img1_affonly, img1_fullwarp], pad=10, axis=1, bg_value=(0.5, 0.9, 0.1)) >>> canvas2 = kwimage.stack_images([img2, img2_projonly, img2_affonly, img2_fullwarp], pad=10, axis=1, bg_value=(0.5, 0.9, 0.1)) >>> canvas = kwimage.stack_images([canvas1, canvas2], axis=0) >>> kwplot.imshow(canvas)
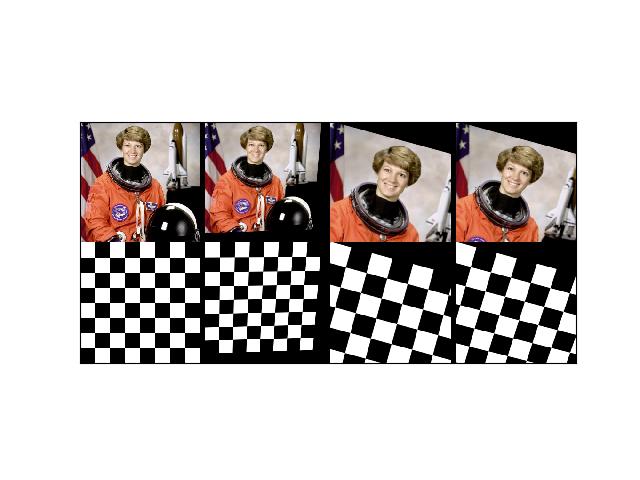

- decompose()[source]¶
Based on the analysis done in [ME1319680].
- Return type:
Dict
References
Example
>>> # Create a set of points, warp them, then recover the warp >>> import kwimage >>> points = kwimage.Points.random(9).scale(64) >>> A1 = kwimage.Affine.affine(scale=0.9, theta=-3.2, offset=(2, 3), about=(32, 32), skew=2.3) >>> A2 = kwimage.Affine.affine(scale=0.8, theta=0.8, offset=(2, 0), about=(32, 32)) >>> A12_real = A2 @ A1.inv() >>> points1 = points.warp(A1) >>> points2 = points.warp(A2) >>> # Make the correspondence non-affine >>> points2.data['xy'].data[0, 0] += 3.5 >>> points2.data['xy'].data[3, 1] += 8.5 >>> # Recover the warp >>> pts1, pts2 = points1.xy, points2.xy >>> self = kwimage.Projective.random() >>> self.decompose()
Example
>>> # xdoctest: +REQUIRES(module:sympy) >>> from kwimage.transform import * # NOQA >>> import kwimage >>> from kwimage.transform import _RationalNDArray >>> self = kwimage.Projective.random().rationalize() >>> rat_decomp = self.decompose() >>> print('rat_decomp = {}'.format(ub.urepr(rat_decomp, nl=1))) >>> #### >>> import sympy >>> cells = sympy.symbols('h1, h2, h3, h4, h5, h6, h7, h8, h9') >>> matrix = _RationalNDArray(cells).reshape(3, 3) >>> # Symbolic decomposition. Neat. >>> self = kwimage.Projective(matrix) >>> self.decompose()
- class kwimage.Segmentation(data, format=None)[source]¶
Bases:
_WrapperObjectEither holds a MultiPolygon, Polygon, or Mask
- Parameters:
data (object) – the underlying object
format (str) – either ‘mask’, ‘polygon’, or ‘multipolygon’
- classmethod random(rng=None)[source]¶
Example
>>> self = Segmentation.random() >>> print('self = {!r}'.format(self)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> self.draw() >>> kwplot.show_if_requested()
- property meta¶
- class kwimage.SegmentationList(data, meta=None)[source]¶
Bases:
ObjectListStore and manipulate multiple segmentations (masks or polygons), usually within the same image
- kwimage.add_homog(pts)[source]¶
Add a homogenous coordinate to a point array
This is a convinience function, it is not particularly efficient.
- SeeAlso:
cv2.convertPointsToHomogeneous
Example
>>> pts = np.random.rand(10, 2) >>> add_homog(pts)
Benchmark
>>> import timerit >>> ti = timerit.Timerit(1000, bestof=10, verbose=2) >>> pts = np.random.rand(1000, 2) >>> for timer in ti.reset('kwimage'): >>> with timer: >>> kwimage.add_homog(pts) >>> for timer in ti.reset('cv2'): >>> with timer: >>> cv2.convertPointsToHomogeneous(pts) >>> # cv2 is 4x faster, but has more restrictive inputs
- kwimage.atleast_3channels(arr, copy=True)[source]¶
Ensures that there are 3 channels in the image
- Parameters:
arr (ndarray) – an image with 2 or 3 dims.
copy (bool) – Always copies if True, if False, then copies only when the size of the array must change. Defaults to True.
- Returns:
with shape (N, M, C), where C in {3, 4}
- Return type:
ndarray
Doctest
>>> assert atleast_3channels(np.zeros((10, 10))).shape[-1] == 3 >>> assert atleast_3channels(np.zeros((10, 10, 1))).shape[-1] == 3 >>> assert atleast_3channels(np.zeros((10, 10, 3))).shape[-1] == 3 >>> assert atleast_3channels(np.zeros((10, 10, 4))).shape[-1] == 4
- kwimage.available_nms_impls()[source]¶
List available values for the impl kwarg of non_max_supression
CommandLine
xdoctest -m kwimage.algo.algo_nms available_nms_impls
Example
>>> impls = available_nms_impls() >>> assert 'numpy' in impls >>> print('impls = {!r}'.format(impls))
- kwimage.checkerboard(num_squares='auto', square_shape='auto', dsize=(512, 512), dtype=<class 'float'>, on_value=1, off_value=0)[source]¶
Creates a checkerboard image
- Parameters:
num_squares (int | str) – Number of squares in a row. If ‘auto’ defaults to 8
square_shape (int | Tuple[int, int] | str) – If ‘auto’, chosen based on num_squares. Otherwise this is the height, width of each square in pixels.
dsize (Tuple[int, int]) – width and height
dtype (type) – return data type
on_value (Number | int) – The value of one checker. Defaults to 1.
off_value (Number | int) – The value off the other checker. Defaults to 0.
References
https://stackoverflow.com/questions/2169478/how-to-make-a-checkerboard-in-numpy
Example
>>> import kwimage >>> import numpy as np >>> img = kwimage.checkerboard() >>> print(kwimage.checkerboard(dsize=(16, 16)).shape) >>> print(kwimage.checkerboard(num_squares=4, dsize=(16, 16)).shape) >>> print(kwimage.checkerboard(square_shape=3, dsize=(23, 17)).shape) >>> print(kwimage.checkerboard(square_shape=3, dsize=(1451, 1163)).shape) >>> print(kwimage.checkerboard(square_shape=3, dsize=(1202, 956)).shape) >>> print(kwimage.checkerboard(dsize=(4, 4), on_value=(255, 0, 0), off_value=(0, 0, 1), dtype=np.uint8))
Example
>>> import kwimage >>> img = kwimage.checkerboard( >>> dsize=(64, 64), on_value='kw_green', off_value='kw_blue') >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autoplt() >>> kwplot.imshow(img) >>> kwplot.show_if_requested()

- kwimage.connected_components(image, connectivity=8, ltype=<class 'numpy.int32'>, with_stats=True, algo='default')[source]¶
Find connected components in a binary image.
Wrapper around
cv2.connectedComponentsWithStats().- Parameters:
image (ndarray) – a binary uint8 image. Zeros denote the background, and non-zeros numbers are foreground regions that will be partitioned into connected components.
connectivity (int) – either 4 or 8
ltype (numpy.dtype | str | int) – The dtype for the output label array. Can be either ‘int32’ or ‘uint16’, and this can be specified as a cv2 code or a numpy dtype.
algo (str) – The underlying algorithm to use. See [Cv2CCAlgos] for details. Options are spaghetti, sauf, bbdt. (default is spaghetti)
- Returns:
The label array and an information dictionary
- Return type:
Tuple[ndarray, dict]
Todo
Document the details of which type of coordinates we are using. I.e. are pixels points or areas? (I think this uses the points convention?)
Note
opencv 4.5.5 will segfault if connectivity=4 See: [CvIssue21366].
Note
Based on information in [SO35854197].
References
CommandLine
xdoctest -m kwimage.im_cv2 connected_components:0 --show
Example
>>> import kwimage >>> from kwimage.im_cv2 import * # NOQA >>> mask = kwimage.Mask.demo() >>> image = mask.data >>> labels, info = connected_components(image) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> canvas0 = kwimage.atleast_3channels(mask.data * 255) >>> canvas2 = canvas0.copy() >>> canvas3 = canvas0.copy() >>> boxes = info['label_boxes'] >>> centroids = info['label_centroids'] >>> label_colors = kwimage.Color.distinct(info['num_labels']) >>> index_to_color = np.array([kwimage.Color('black').as01()] + label_colors) >>> canvas2 = centroids.draw_on(canvas2, color=label_colors, radius=None) >>> boxes.draw_on(canvas3, color=label_colors, thickness=1) >>> legend = kwplot.make_legend_img(ub.dzip(range(len(index_to_color)), index_to_color)) >>> colored_label_img = index_to_color[labels] >>> canvas1 = kwimage.stack_images([colored_label_img, legend], axis=1, resize='smaller') >>> kwplot.imshow(canvas0, pnum=(1, 4, 1), title='input image') >>> kwplot.imshow(canvas1, pnum=(1, 4, 2), title='label image (colored w legend)') >>> kwplot.imshow(canvas2, pnum=(1, 4, 3), title='component centroids') >>> kwplot.imshow(canvas3, pnum=(1, 4, 4), title='component bounding boxes')

- kwimage.convert_colorspace(img, src_space, dst_space, copy=False, implicit=False, dst=None)[source]¶
Converts colorspace of img.
Convenience function around
cv2.cvtColor()- Parameters:
img (ndarray) – image data with float32 or uint8 precision
src_space (str) – input image colorspace. (e.g. BGR, GRAY)
dst_space (str) – desired output colorspace. (e.g. RGB, HSV, LAB)
implicit (bool) –
- if False, the user must correctly specify if the input/output
colorspaces contain alpha channels.
- If True and the input image has an alpha channel, we modify
src_space and dst_space to ensure they both end with “A”.
dst (ndarray[Any, UInt8]) – inplace-output array.
- Returns:
img - image data
- Return type:
ndarray
Note
Note the LAB and HSV colorspaces in float do not go into the 0-1 range.
- For HSV the floating point range is:
0:360, 0:1, 0:1
- For LAB the floating point range is:
0:100, -86.1875:98.234375, -107.859375:94.46875 (Note, that some extreme combinations of a and b are not valid)
Example
>>> import numpy as np >>> convert_colorspace(np.array([[[0, 0, 1]]], dtype=np.float32), 'RGB', 'LAB') >>> convert_colorspace(np.array([[[0, 1, 0]]], dtype=np.float32), 'RGB', 'LAB') >>> convert_colorspace(np.array([[[1, 0, 0]]], dtype=np.float32), 'RGB', 'LAB') >>> convert_colorspace(np.array([[[1, 1, 1]]], dtype=np.float32), 'RGB', 'LAB') >>> convert_colorspace(np.array([[[0, 0, 1]]], dtype=np.float32), 'RGB', 'HSV')
- kwimage.daq_spatial_nms(ltrb, scores, diameter, thresh, max_depth=6, stop_size=2048, recsize=2048, impl='auto', device_id=None)[source]¶
Divide and conquor speedup non-max-supression algorithm for when bboxes have a known max size
- Parameters:
ltrb (ndarray) – boxes in (tlx, tly, brx, bry) format
scores (ndarray) – scores of each box
diameter (int | Tuple[int, int]) – Distance from split point to consider rectification. If specified as an integer, then number is used for both height and width. If specified as a tuple, then dims are assumed to be in [height, width] format.
thresh (float) – iou threshold. Boxes are removed if they overlap greater than this threshold. 0 is the most strict, resulting in the fewest boxes, and 1 is the most permissive resulting in the most.
max_depth (int) – maximum number of times we can divide and conquor
stop_size (int) – number of boxes that triggers full NMS computation
recsize (int) – number of boxes that triggers full NMS recombination
impl (str) – algorithm to use
Note
# TODO: Look Into
# Didn’t read yet but it seems similar
http://www.cyberneum.de/fileadmin/user_upload/files/publications/CVPR2010-Lampert_[0].pdf
https://www.researchgate.net/publication/220929789_Efficient_Non-Maximum_Suppression
# This seems very similar
https://projet.liris.cnrs.fr/m2disco/pub/Congres/2006-ICPR/DATA/C03_0406.PDF
Example
>>> import kwimage >>> # Make a bunch of boxes with the same width and height >>> #boxes = kwimage.Boxes.random(230397, scale=1000, format='cxywh') >>> boxes = kwimage.Boxes.random(237, scale=1000, format='cxywh') >>> boxes.data.T[2] = 10 >>> boxes.data.T[3] = 10 >>> # >>> ltrb = boxes.to_ltrb().data.astype(np.float32) >>> scores = np.arange(0, len(ltrb)).astype(np.float32) >>> # >>> n_megabytes = (ltrb.size * ltrb.dtype.itemsize) / (2 ** 20) >>> print('n_megabytes = {!r}'.format(n_megabytes)) >>> # >>> thresh = iou_thresh = 0.01 >>> impl = 'auto' >>> max_depth = 20 >>> diameter = 10 >>> stop_size = 2000 >>> recsize = 500 >>> # >>> import ubelt as ub >>> # >>> with ub.Timer(label='daq'): >>> keep1 = daq_spatial_nms(ltrb, scores, >>> diameter=diameter, thresh=thresh, max_depth=max_depth, >>> stop_size=stop_size, recsize=recsize, impl=impl) >>> # >>> with ub.Timer(label='full'): >>> keep2 = non_max_supression(ltrb, scores, >>> thresh=thresh, impl=impl) >>> # >>> # Due to the greedy nature of the algorithm, there will be slight >>> # differences in results, but they will be mostly similar. >>> similarity = len(set(keep1) & set(keep2)) / len(set(keep1) | set(keep2)) >>> print('similarity = {!r}'.format(similarity))
- kwimage.decode_run_length(counts, shape, binary=False, dtype=<class 'numpy.uint8'>, order='C')[source]¶
Decode run length encoding back into an image.
- Parameters:
counts (ndarray) – the run-length encoding
shape (Tuple[int, int]) – the height / width of the mask
binary (bool) – if the RLE is binary or non-binary. Set to True for compatibility with COCO.
dtype (type) – data type for decoded image. Defaults to np.uint8.
order (str) – Order of the encoding. Either ‘C’ for row major or ‘F’ for column-major. Defaults to ‘C’.
- Returns:
the reconstructed image
- Return type:
ndarray
Example
>>> from kwimage.im_runlen import * # NOQA >>> img = np.array([[1, 0, 1, 1, 1, 0, 0, 1, 0]]) >>> encoded = encode_run_length(img, binary=True) >>> recon = decode_run_length(**encoded) >>> assert np.all(recon == img)
>>> import ubelt as ub >>> lines = ub.codeblock( >>> ''' >>> .......... >>> ......111. >>> ..2...111. >>> .222..111. >>> 22222..... >>> .222...... >>> ..2....... >>> ''').replace('.', '0').splitlines() >>> img = np.array([list(map(int, line)) for line in lines]) >>> encoded = encode_run_length(img) >>> recon = decode_run_length(**encoded) >>> assert np.all(recon == img)
- kwimage.draw_boxes_on_image(img, boxes, color='blue', thickness=1, box_format=None, colorspace='rgb')[source]¶
Draws boxes on an image.
- Parameters:
img (ndarray) – image to copy and draw on
boxes (kwimage.Boxes | ndarray) – boxes to draw
colorspace (str) – string code of the input image colorspace
Example
>>> import kwimage >>> import numpy as np >>> img = np.zeros((10, 10, 3), dtype=np.uint8) >>> color = 'dodgerblue' >>> thickness = 1 >>> boxes = kwimage.Boxes([[1, 1, 8, 8]], 'ltrb') >>> img2 = draw_boxes_on_image(img, boxes, color, thickness) >>> assert tuple(img2[1, 1]) == (30, 144, 255) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() # xdoctest: +SKIP >>> kwplot.figure(doclf=True, fnum=1) >>> kwplot.imshow(img2)

- kwimage.draw_clf_on_image(im, classes, tcx=None, probs=None, pcx=None, border=1)[source]¶
Draws classification label on an image.
Works best with image chips sized between 200x200 and 500x500
- Parameters:
im (ndarray) – the image
classes (Sequence[str] | kwcoco.CategoryTree) – list of class names
tcx (int) – true class index if known
probs (ndarray) – predicted class probs for each class
pcx (int) – predicted class index. (if None but probs is specified uses argmax of probs)
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> import kwarray >>> import kwimage >>> rng = kwarray.ensure_rng(0) >>> im = (rng.rand(300, 300) * 255).astype(np.uint8) >>> classes = ['cls_a', 'cls_b', 'cls_c'] >>> tcx = 1 >>> probs = rng.rand(len(classes)) >>> probs[tcx] = 0 >>> probs = torch.FloatTensor(probs).softmax(dim=0).numpy() >>> im1_ = kwimage.draw_clf_on_image(im, classes, tcx, probs) >>> probs[tcx] = .9 >>> probs = torch.FloatTensor(probs).softmax(dim=0).numpy() >>> im2_ = kwimage.draw_clf_on_image(im, classes, tcx, probs) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(im1_, colorspace='rgb', pnum=(1, 2, 1), fnum=1, doclf=True) >>> kwplot.imshow(im2_, colorspace='rgb', pnum=(1, 2, 2), fnum=1) >>> kwplot.show_if_requested()
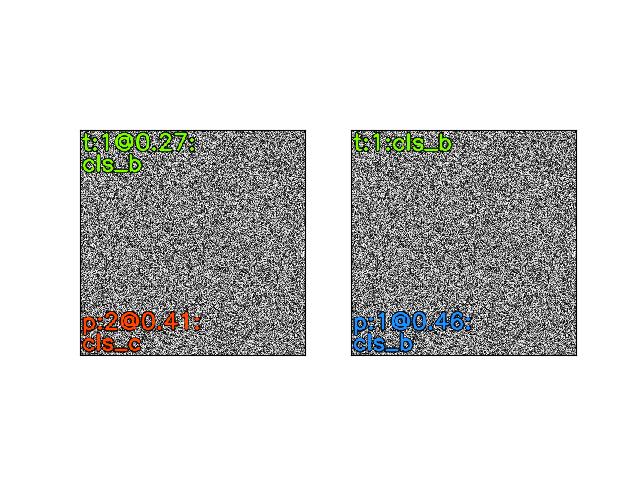

- kwimage.draw_header_text(image=None, text=None, fit=False, color='strawberry', halign='center', stack='auto', bg_color='black', **kwargs)[source]¶
Places a black bar on top of an image and writes text in it
- Parameters:
image (ndarray | dict | None) – numpy image or dictionary containing a key width
text (str) – text to draw
fit (bool | str) – If False, will draw as much text within the given width as possible. If True, will draw all text and then resize to fit in the given width If “shrink”, will only resize the text if it is too big to fit, in other words this is like fit=True, but it wont enlarge the text.
color (str | Tuple) – a color coercable to
kwimage.Color.halign (str) – Horizontal alignment. Can be left, center, or right.
stack (bool | str) – if True returns the stacked image, otherwise just returns the header. If ‘auto’, will only stack if an image is given as an ndarray.
**kwargs – used only for parameter aliases. Currently accepts:
bg_value as an alias for bg_color.
fontScale
fontFace
thickness
- Returns:
ndarray
Example
>>> from kwimage.im_draw import * # NOQA >>> import kwimage >>> image = kwimage.grab_test_image() >>> tiny_image = kwimage.imresize(image, dsize=(64, 64)) >>> canvases = [] >>> canvases += [draw_header_text(image=image, text='unfit long header ' * 5, fit=False)] >>> canvases += [draw_header_text(image=image, text='shrunk long header ' * 5, fit='shrink')] >>> canvases += [draw_header_text(image=image, text='left header', fit=False, halign='left')] >>> canvases += [draw_header_text(image=image, text='center header', fit=False, halign='center')] >>> canvases += [draw_header_text(image=image, text='right header', fit=False, halign='right')] >>> canvases += [draw_header_text(image=image, text='shrunk header', fit='shrink', halign='left')] >>> canvases += [draw_header_text(image=tiny_image, text='shrunk header-center', fit='shrink', halign='center')] >>> canvases += [draw_header_text(image=image, text='fit header', fit=True, halign='left')] >>> canvases += [draw_header_text(image={'width': 200}, text='header only', fit=True, halign='left')] >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nCols=3, nSubplots=len(canvases)) >>> for c in canvases: >>> kwplot.imshow(c, pnum=pnum_()) >>> kwplot.show_if_requested()

Example
>>> # Test fontScale >>> from kwimage.im_draw import * # NOQA >>> import kwimage >>> test_image = kwimage.grab_test_image(dsize=(900, 120)) >>> canvases = [] >>> for image in [test_image, None]: >>> canvases += [draw_header_text(image=image, text='Scale=1.0 Fit=False', fit=False, fontScale=1.0)] >>> canvases += [draw_header_text(image=image, text='Scale=2.0 Fit=False', fit=False, fontScale=2.0)] >>> canvases += [draw_header_text(image=image, text='Scale=6.0 Fit=False', fit=False, fontScale=6.0)] >>> canvases += [draw_header_text(image=image, text='Scale=1.0 Fit=True', fit=True, fontScale=1.0)] >>> canvases += [draw_header_text(image=image, text='Scale=2.0 Fit=True', fit=True, fontScale=2.0)] >>> canvases += [draw_header_text(image=image, text='Scale=6.0 Fit=True', fit=True, fontScale=6.0)] >>> canvases += [draw_header_text(image=image, text='Scale=1.0 Fit=shrink', fit='shrink', fontScale=1.0)] >>> canvases += [draw_header_text(image=image, text='Scale=2.0 Fit=shrink', fit='shrink', fontScale=2.0)] >>> canvases += [draw_header_text(image=image, text='Scale=6.0 Fit=shrink', fit='shrink', fontScale=6.0)] >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nCols=3, nSubplots=len(canvases)) >>> for c in canvases: >>> kwplot.imshow(c, pnum=pnum_(), title=str(c.shape)) >>> kwplot.show_if_requested()

- kwimage.draw_line_segments_on_image(img, pts1, pts2, color='blue', colorspace='rgb', thickness=1, **kwargs)[source]¶
Draw line segments between pts1 and pts2 on an image.
- Parameters:
pts1 (ndarray) – xy coordinates of starting points
pts2 (ndarray) – corresponding xy coordinates of ending points
color (str | List) – color code or a list of colors for each line segment
colorspace (str) – colorspace of image. Defaults to ‘rgb’
thickness (int) – Defaults to 1
lineType (int) – option for cv2.line
- Returns:
the modified image (inplace if possible)
- Return type:
ndarray
Example
>>> from kwimage.im_draw import * # NOQA >>> pts1 = np.array([[2, 0], [2, 20], [2.5, 30]]) >>> pts2 = np.array([[10, 5], [30, 28], [100, 50]]) >>> img = np.ones((100, 100, 3), dtype=np.uint8) * 255 >>> color = 'blue' >>> colorspace = 'rgb' >>> img2 = draw_line_segments_on_image(img, pts1, pts2, thickness=2) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() # xdoctest: +SKIP >>> kwplot.figure(doclf=True, fnum=1) >>> kwplot.imshow(img2)
Example
>>> import kwimage >>> # xdoctest: +REQUIRES(module:matplotlib) >>> pts1 = kwimage.Points.random(10).scale(512).xy >>> pts2 = kwimage.Points.random(10).scale(512).xy >>> img = np.ones((512, 512, 3), dtype=np.uint8) * 255 >>> color = kwimage.Color.distinct(10) >>> img2 = kwimage.draw_line_segments_on_image(img, pts1, pts2, color=color) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() # xdoctest: +SKIP >>> kwplot.figure(doclf=True, fnum=1) >>> kwplot.imshow(img2)
- kwimage.draw_text_on_image(img, text, org=None, return_info=False, **kwargs)[source]¶
Draws multiline text on an image using opencv
- Parameters:
img (ndarray | None | dict) – Generally a numpy image to draw on (inplace). Otherwise a canvas will be constructed such that the text will fit. The user may specify a dictionary with keys width and height to have more control over the constructed canvas.
text (str) – text to draw
org (Tuple[int, int]) – The x, y location of the text string “anchor” in the image as specified by halign and valign. For instance, If valign=’bottom’, halign=’left’, this where the bottom left corner of the text will be placed.
return_info (bool) – if True, also returns information about the positions the text was drawn on.
**kwargs – color (tuple): default blue
thickness (int): defaults to 2
fontFace (int): defaults to cv2.FONT_HERSHEY_SIMPLEX
fontScale (float): defaults to 1.0
valign (str): either top, center, or bottom. Defaults to “bottom” NOTE: this default may change to “top” in the future.
halign (str): either left, center, or right. Defaults to “left”.
border (dict | int): If specified as an integer, draws a black border with that given thickness. If specified as a dictionary, draws a border with color specified parameters. “color”: border color, defaults to “black”. “thickness”: border thickness, defaults to 1.
- Returns:
The image that was drawn on and optionally an information dictionary if return_info was True.
- Return type:
ndarray | Tuple[ndarray, dict]
Note
The image is modified inplace. If the image is non-contiguous then this returns a UMat instead of a ndarray, so be careful with that.
- Related:
The logic in this function is related to the following stack overflow posts [SO27647424] [SO51285616]
References
Example
>>> import kwimage >>> img = kwimage.grab_test_image(space='rgb') >>> img2 = kwimage.draw_text_on_image(img.copy(), 'FOOBAR', org=(0, 0), valign='top') >>> assert img2.shape == img.shape >>> assert np.any(img2 != img) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img2) >>> kwplot.show_if_requested()

Example
>>> import kwimage >>> # Test valign >>> img = kwimage.grab_test_image(space='rgb', dsize=(640, 640)) >>> img2 = kwimage.draw_text_on_image(img, 'VALIGN-top\nbazbiz\nspam', org=(0, 100), valign='top', border=2) >>> img2 = kwimage.draw_text_on_image(img, 'VALIGN-center\nbazbiz\nspam', org=(200, 100), valign='center', border=2) >>> img2 = kwimage.draw_text_on_image(img, 'VALIGN-bottom\nbazbiz\nspam', org=(450, 100), valign='bottom', border=2) >>> # Test halign >>> img2 = kwimage.draw_text_on_image(img, 'HALIGN-right\nbazbiz\nspam', org=(250, 300), halign='right', border=2) >>> img2 = kwimage.draw_text_on_image(img, 'HALIGN-center\nbazbiz\nspam', org=(250, 400), halign='center', border=2) >>> img2 = kwimage.draw_text_on_image(img, 'HALIGN-left\nbazbiz\nspam', org=(250, 500), halign='left', border=2) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img2) >>> kwplot.show_if_requested()

Example
>>> # Ensure the function works with float01 or uint255 images >>> import kwimage >>> img = kwimage.grab_test_image(space='rgb') >>> img = kwimage.ensure_float01(img) >>> img2 = img.copy() >>> img2 = kwimage.draw_text_on_image(img2, 'FOOBAR\nbazbiz\nspam', org=(0, 0), valign='top', border=2, fontScale=1.0) >>> img2 = kwimage.draw_text_on_image(img2, 'FOOBAR\nbazbiz\nspam', org=(0, 200), valign='top', border=2, fontScale=2.0) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img2) >>> kwplot.show_if_requested()

Example
>>> # Test dictionary border >>> import kwimage >>> img = kwimage.draw_text_on_image(None, 'Battery\nFraction', org=(100, 100), valign='top', halign='center', border={'color': 'green', 'thickness': 9}) >>> #img = kwimage.draw_text_on_image(None, 'hello\neveryone', org=(0, 0), valign='top') >>> #img = kwimage.draw_text_on_image(None, 'hello', org=(0, 60), valign='top', halign='center', border=0) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img) >>> kwplot.show_if_requested()

Example
>>> # Test dictionary image >>> import kwimage >>> img = kwimage.draw_text_on_image({'width': 300}, 'Arbitrary\nText', org=(150, 0), valign='top', halign='center', border={'color': 'green', 'thickness': 0}) >>> print('img.shape = {!r}'.format(img.shape)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img) >>> kwplot.show_if_requested()

Example
>>> # Test fontScale >>> import kwimage >>> canvases = [] >>> canvases.append(kwimage.draw_text_on_image(None, 'FontScale=1.0', fontScale=1.0)) >>> canvases.append(kwimage.draw_text_on_image(None, 'FontScale=2.0', fontScale=2.0)) >>> canvases.append(kwimage.draw_text_on_image(None, 'FontScale=3.0', fontScale=3.0)) >>> # xdoctest: +REQUIRES(--show) >>> canvas = kwimage.stack_images_grid(canvases, pad=10, bg_value=(255, 255, 255)) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()

Example
>>> import ubelt as ub >>> import kwimage >>> grid = list(ub.named_product({ >>> 'halign': ['left', 'center', 'right', None], >>> 'valign': ['top', 'center', 'bottom', None], >>> 'border': [0, 3] >>> })) >>> canvases = [] >>> text = 'small-line\na-much-much-much-bigger-line\nanother-small\n.' >>> for kw in grid: >>> header = kwimage.draw_text_on_image({}, ub.urepr(kw, compact=1), color='blue') >>> canvas = kwimage.draw_text_on_image({'color': 'white'}, text, org=None, **kw) >>> canvases.append(kwimage.stack_images([header, canvas], axis=0, bg_value=(255, 255, 255), pad=5)) >>> # xdoctest: +REQUIRES(--show) >>> canvas = kwimage.stack_images_grid(canvases, pad=10, bg_value=(255, 255, 255)) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()

- kwimage.draw_vector_field(image, dx, dy, stride=0.02, thresh=0.0, scale=1.0, alpha=1.0, color='strawberry', thickness=1, tipLength=0.1, line_type='aa')[source]¶
Create an image representing a 2D vector field.
- Parameters:
image (ndarray) – image to draw on
dx (ndarray) – grid of vector x components
dy (ndarray) – grid of vector y components
stride (int | float) – sparsity of vectors, int specifies stride step in pixels, a float specifies it as a percentage.
thresh (float) – only plot vectors with magnitude greater than thres
scale (float) – multiply magnitude for easier visualization
alpha (float) – alpha value for vectors. Non-vector regions receive 0 alpha (if False, no alpha channel is used)
color (str | tuple | kwimage.Color) – RGB color of the vectors
thickness (int) – thickness of arrows
tipLength (float) – fraction of line length
line_type (int | str) – either cv2.LINE_4, cv2.LINE_8, or cv2.LINE_AA or ‘aa’
- Returns:
The image with vectors overlaid. If image=None, then an rgb/a image is created and returned.
- Return type:
ndarray[Any, Float32]
Example
>>> from kwimage.im_draw import * # NOQA >>> import kwimage >>> width, height = 512, 512 >>> image = kwimage.grab_test_image(dsize=(width, height)) >>> x, y = np.meshgrid(np.arange(height), np.arange(width)) >>> dx, dy = x - width / 2, y - height / 2 >>> radians = np.arctan2(dx, dy) >>> mag = np.sqrt(dx ** 2 + dy ** 2) + 1e-3 >>> dx, dy = dx / mag, dy / mag >>> img = kwimage.draw_vector_field(image, dx, dy, scale=10, alpha=False) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img, title='draw_vector_field') >>> kwplot.show_if_requested()

- kwimage.encode_run_length(img, binary=False, order='C')[source]¶
Construct the run length encoding (RLE) of an image.
- Parameters:
img (ndarray) – 2D image
binary (bool) – If true, assume that the input image only contains 0’s and 1’s. Set to True for compatibility with COCO (which does not support multi-value RLE encodings).
order (str) – Order of the encoding. Either ‘C’ for row major or ‘F’ for column-major. Defaults to ‘C’.
- Returns:
encoding: dictionary items are:
counts (ndarray): the run length encoding
- shape (Tuple): the original image shape.
This should be in standard shape row-major (e.g. h/w) order
- binary (bool):
if True, the counts are assumed to encode only 0’s and 1’s, otherwise the counts encoding specifies any numeric values.
- order (str):
Encoding order, either ‘C’ for row major or ‘F’ for column-major. Defaults to ‘C’.
- Return type:
- SeeAlso:
kwimage.Mask-cython-backed data structure to handle coco-style RLEs
Example
>>> import ubelt as ub >>> lines = ub.codeblock( >>> ''' >>> .......... >>> ......111. >>> ..2...111. >>> .222..111. >>> 22222..... >>> .222...... >>> ..2....... >>> ''').replace('.', '0').splitlines() >>> img = np.array([list(map(int, line)) for line in lines]) >>> encoding = encode_run_length(img) >>> target = np.array([0,16,1,3,0,3,2,1,0,3,1,3,0,2,2,3,0,2,1,3,0,1,2,5,0,6,2,3,0,8,2,1,0,7]) >>> assert np.all(target == encoding['counts'])
Example
>>> binary = True >>> img = np.array([[1, 0, 1, 1, 1, 0, 0, 1, 0]]) >>> encoding = encode_run_length(img, binary=True) >>> assert encoding['counts'].tolist() == [0, 1, 1, 3, 2, 1, 1]
Example
>>> # Test empty case >>> from kwimage.im_runlen import * # NOQA >>> binary = True >>> img = np.zeros((0, 0), dtype=np.uint8) >>> encoding = encode_run_length(img, binary=True) >>> assert encoding['counts'].tolist() == [] >>> recon = decode_run_length(**encoding) >>> assert np.all(recon == img)
Example
>>> # Test small full cases >>> for d in [0, 1, 2, 3]: >>> img = np.zeros((d, d), dtype=np.uint8) >>> encoding = encode_run_length(img, binary=True) >>> recon = decode_run_length(**encoding) >>> assert np.all(recon == img) >>> img = np.ones((d, d), dtype=np.uint8) >>> encoding = encode_run_length(img, binary=True) >>> recon = decode_run_length(**encoding) >>> assert np.all(recon == img)
- kwimage.ensure_alpha_channel(img, alpha=1.0, dtype=<class 'numpy.float32'>, copy=False)[source]¶
Returns the input image with 4 channels.
- Parameters:
img (ndarray) – an image with shape [H, W], [H, W, 1], [H, W, 3], or [H, W, 4].
alpha (float | ndarray) – default scalar value for missing alpha channel, or an ndarray with the same height / width to use explicitly.
dtype (type) – The final output dtype. Should be numpy.float32 or numpy.float64.
copy (bool) – always copy if True, else copy if needed.
- Returns:
an image with specified dtype with shape [H, W, 4].
- Return type:
ndarray
- Raises:
ValueError - if the input image does not have 1, 3, or 4 input channels – or if the image cannot be converted into a float01 representation
Example
>>> # Demo with a scalar default alpha value >>> import kwimage >>> data0 = np.zeros((5, 5)) >>> data1 = np.zeros((5, 5, 1)) >>> data2 = np.zeros((5, 5, 3)) >>> data3 = np.zeros((5, 5, 4)) >>> ensured0 = kwimage.ensure_alpha_channel(data0, alpha=0.5) >>> ensured1 = kwimage.ensure_alpha_channel(data1, alpha=0.5) >>> ensured2 = kwimage.ensure_alpha_channel(data2, alpha=0.5) >>> ensured3 = kwimage.ensure_alpha_channel(data3, alpha=0.5) >>> assert np.all(ensured0[..., 3] == 0.5), 'should have been populated' >>> assert np.all(ensured1[..., 3] == 0.5), 'should have been populated' >>> assert np.all(ensured2[..., 3] == 0.5), 'should have been populated' >>> assert np.all(ensured3[..., 3] == 0.0), 'last image already had alpha'
Example
>>> import kwimage >>> # Demo with a explicit alpha channel >>> alpha = np.random.rand(5, 5) >>> data0 = np.zeros((5, 5)) >>> data1 = np.zeros((5, 5, 1)) >>> data2 = np.zeros((5, 5, 3)) >>> data3 = np.zeros((5, 5, 4)) >>> ensured0 = kwimage.ensure_alpha_channel(data0, alpha=alpha) >>> ensured1 = kwimage.ensure_alpha_channel(data1, alpha=alpha) >>> ensured2 = kwimage.ensure_alpha_channel(data2, alpha=alpha) >>> ensured3 = kwimage.ensure_alpha_channel(data3, alpha=alpha) >>> assert np.all(ensured0[..., 3] == alpha), 'should have been populated' >>> assert np.all(ensured1[..., 3] == alpha), 'should have been populated' >>> assert np.all(ensured2[..., 3] == alpha), 'should have been populated' >>> assert np.all(ensured3[..., 3] == 0.0), 'last image already had alpha'
- kwimage.ensure_float01(img, dtype=<class 'numpy.float32'>, copy=True)[source]¶
Ensure that an image is encoded using a float32 properly
- Parameters:
img (ndarray) – an image in uint255 or float01 format. Other formats will raise errors.
dtype (type) – a numpy floating type defaults to np.float32
copy (bool) – Always copy if True, else copy if needed. Defaults to True.
- Returns:
an array of floats in the range 0-1
- Return type:
ndarray
- Raises:
ValueError – if the image type is integer and not in [0-255]
Example
>>> ensure_float01(np.array([[0, .5, 1.0]])) array([[0. , 0.5, 1. ]], dtype=float32) >>> ensure_float01(np.array([[0, 1, 200]])) array([[0..., 0.0039..., 0.784...]], dtype=float32)
- kwimage.ensure_uint255(img, copy=True)[source]¶
Ensure that an image is encoded using a uint8 properly. Either
- Parameters:
img (ndarray) – an image in uint255 or float01 format. Other formats will raise errors.
copy (bool) – always copy if True, else copy if needed. Defaults to True.
- Returns:
an array of bytes in the range 0-255
- Return type:
ndarray
- Raises:
ValueError – if the image type is float and not in [0-1]
ValueError – if the image type is integer and not in [0-255]
Example
>>> ensure_uint255(np.array([[0, .5, 1.0]])) array([[ 0, 127, 255]], dtype=uint8) >>> ensure_uint255(np.array([[0, 1, 200]])) array([[ 0, 1, 200]], dtype=uint8)
- kwimage.exactly_1channel(image, ndim=2)[source]¶
Returns a 1-channel image as either a 2D or 3D array.
- Parameters:
image (ndarray) – an image with shape (H, W, 1) or (H, W).
ndim (int) – number of dimensions in the output array. Can be either 2 or 3.
- Returns:
if ndim is 2, returns a (H, W) image. if ndim is 3, returns a (H, W, 1) image.
- Return type:
ndarray
- Raises:
ValueError – if assumptions are not met.
Example
>>> import kwimage >>> assert kwimage.exactly_1channel(np.empty((3, 3)), ndim=2).shape == (3, 3) >>> assert kwimage.exactly_1channel(np.empty((3, 3)), ndim=3).shape == (3, 3, 1) >>> assert kwimage.exactly_1channel(np.empty((3, 3, 1)), ndim=2).shape == (3, 3) >>> assert kwimage.exactly_1channel(np.empty((3, 3, 1)), ndim=3).shape == (3, 3, 1) >>> import pytest >>> with pytest.raises(ValueError): >>> kwimage.exactly_1channel(np.empty((3, 3, 2)), ndim=2) >>> with pytest.raises(ValueError): >>> kwimage.exactly_1channel(np.empty((3)), ndim=3)
- kwimage.fill_nans_with_checkers(canvas, square_shape=8, on_value='auto', off_value='auto')[source]¶
Fills nan or masked values with a 2d checkerboard pattern.
- Parameters:
canvas (np.ndarray) – data replace nans in
square_shape (int | Tuple[int, int] | str) – Size of the checker squares. Defaults to 8.
on_value (Number | str) – The value of one checker. Defaults to a dark-gray color, 0.3 for floats and 77 for ints.
off_value (Number | str) – The value off the other checker. Defaults to black, which is 0.
- Returns:
the inplace modified canvas
- Return type:
np.ndarray
- SeeAlso:
nodata_checkerboard()- similar, but operates on nans or masked arrays.
Example
>>> from kwimage.im_draw import * # NOQA >>> import kwimage >>> orig_img = kwimage.ensure_float01(kwimage.grab_test_image()) >>> poly1 = kwimage.Polygon.random(rng=1).scale(orig_img.shape[0] // 2) >>> poly2 = kwimage.Polygon.random(rng=3).scale(orig_img.shape[0]) >>> poly3 = kwimage.Polygon.random(rng=4).scale(orig_img.shape[0] // 2) >>> poly3 = poly3.translate((0, 200)) >>> poly4 = poly2.translate((100, 0)) >>> poly5 = poly2.translate((50, 100)) >>> img = orig_img.copy() >>> img = poly1.fill(img, np.nan) >>> img = poly3.fill(img, 0) >>> img[:, :, 0] = poly2.fill(np.ascontiguousarray(img[:, :, 0]), np.nan) >>> img[:, :, 2] = poly4.fill(np.ascontiguousarray(img[:, :, 2]), np.nan) >>> img[:, :, 1] = poly5.fill(np.ascontiguousarray(img[:, :, 1]), np.nan) >>> input_img = img.copy() >>> canvas = fill_nans_with_checkers(input_img, on_value=0.3) >>> assert input_img is canvas >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img, pnum=(1, 2, 1), title='matplotlib treats nans as zeros') >>> kwplot.imshow(canvas, pnum=(1, 2, 2), title='checkers highlight real nans')
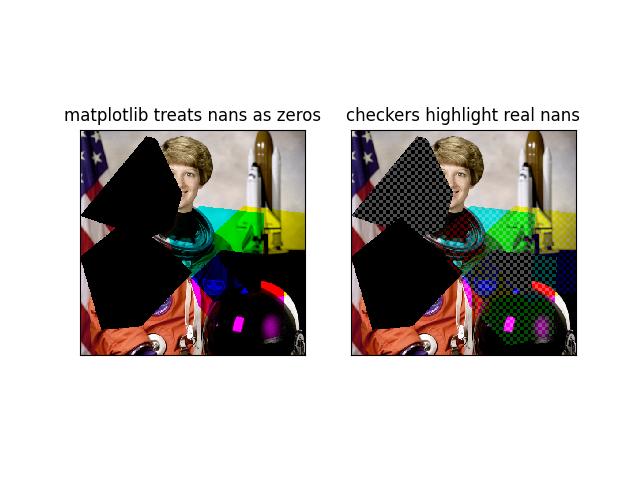

Example
>>> # Test grayscale >>> from kwimage.im_draw import * # NOQA >>> import kwimage >>> orig_img = kwimage.ensure_float01(kwimage.grab_test_image()) >>> poly1 = kwimage.Polygon.random().scale(orig_img.shape[0] // 2) >>> poly2 = kwimage.Polygon.random().scale(orig_img.shape[0]) >>> img = orig_img.copy() >>> img = poly1.fill(img, np.nan) >>> img[:, :, 0] = poly2.fill(np.ascontiguousarray(img[:, :, 0]), np.nan) >>> img = kwimage.convert_colorspace(img, 'rgb', 'gray') >>> canvas = img.copy() >>> canvas = fill_nans_with_checkers(canvas) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img, pnum=(1, 2, 1)) >>> kwplot.imshow(canvas, pnum=(1, 2, 2))
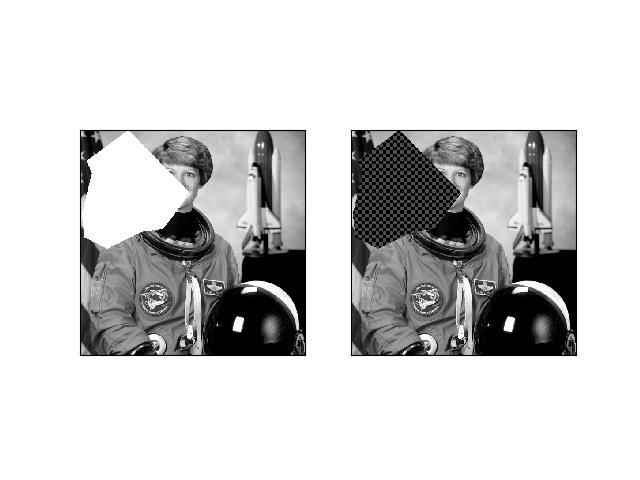

- kwimage.find_robust_normalizers(data, params='auto')[source]¶
Finds robust normalization statistics for a single observation
DEPRECATED IN FAVOR of kwarray.find_robust_normalizers
- Parameters:
data (ndarray) – a 1D numpy array where invalid data has already been removed
params (str | dict) – normalization params
- Returns:
normalization parameters
- Return type:
Todo
[ ] No Magic Numbers! Use first principles to deterimine defaults.
[ ] Probably a lot of literature on the subject.
[ ] Is this a kwarray function in general?
Example
>>> from kwimage.im_core import * # NOQA >>> data = np.random.rand(100) >>> norm_params1 = find_robust_normalizers(data, params='auto') >>> norm_params2 = find_robust_normalizers(data, params={'low': 0, 'high': 1.0}) >>> norm_params3 = find_robust_normalizers(np.empty(0), params='auto') >>> print('norm_params1 = {}'.format(ub.urepr(norm_params1, nl=1))) >>> print('norm_params2 = {}'.format(ub.urepr(norm_params2, nl=1))) >>> print('norm_params3 = {}'.format(ub.urepr(norm_params3, nl=1)))
- kwimage.fourier_mask(img_hwc, mask, axis=None, clip=None, backend='cv2')[source]¶
Applies a mask to the fourier spectrum of an image
- Parameters:
img_hwc (ndarray) – assumed to be float 01
mask (ndarray) –
- mask used to modulate the image in the fourier domain.
Usually these are boolean values (hence the name mask), but any numerical value is technically allowed.
- backend (str):
which implementation of DFT to use. Can be ‘cv2’ or ‘numpy’. Defaults to cv2.
CommandLine
XDEV_PROFILE=1 xdoctest -m kwimage.im_filter fourier_mask --show
import kwimage img_hwc = kwimage.grab_test_image(space=’gray’) mask = np.random.rand(*img_hwc.shape[0:2]) out_hwc = fourier_mask(img_hwc, mask) for timer in ti.reset(‘fft mask with numpy’):
- with timer:
fourier_mask(out_hwc, mask, backend=’numpy’)
- for timer in ti.reset(‘fft mask with cv2’):
- with timer:
fourier_mask(out_hwc, mask, backend=’cv2’)
Example
>>> from kwimage.im_filter import * # NOQA >>> import kwimage >>> img_hwc = kwimage.grab_test_image(space='gray') >>> mask = np.random.rand(*img_hwc.shape[0:2]) >>> mask = (kwimage.gaussian_blur(mask) > 0.5) >>> out_hwc_cv2 = fourier_mask(img_hwc, mask, backend='numpy') >>> out_hwc_np = fourier_mask(img_hwc, mask, backend='cv2') >>> # xdoctest: REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img_hwc, pnum=(1, 4, 1), fnum=1, title='input') >>> kwplot.imshow(mask, pnum=(1, 4, 2), fnum=1, title='mask') >>> kwplot.imshow(out_hwc_cv2, pnum=(1, 4, 3), fnum=1, title='numpy') >>> kwplot.imshow(out_hwc_np, pnum=(1, 4, 4), fnum=1, title='cv2') >>> kwplot.show_if_requested()

Example
>>> from kwimage.im_filter import * # NOQA >>> import kwimage >>> img_hwc = kwimage.grab_test_image(space='gray') >>> mask = kwimage.gaussian_patch(img_hwc.shape[0:2]) >>> mask = (mask / mask.max()) ** 32 >>> out_hwc_cv2 = fourier_mask(img_hwc, mask, backend='numpy') >>> out_hwc_np = fourier_mask(img_hwc, mask, backend='cv2') >>> # xdoctest: REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img_hwc, pnum=(1, 4, 1), fnum=1, title='input') >>> kwplot.imshow(mask, pnum=(1, 4, 2), fnum=1, title='mask') >>> kwplot.imshow(out_hwc_cv2, pnum=(1, 4, 3), fnum=1, title='numpy') >>> kwplot.imshow(out_hwc_np, pnum=(1, 4, 4), fnum=1, title='cv2') >>> kwplot.show_if_requested()
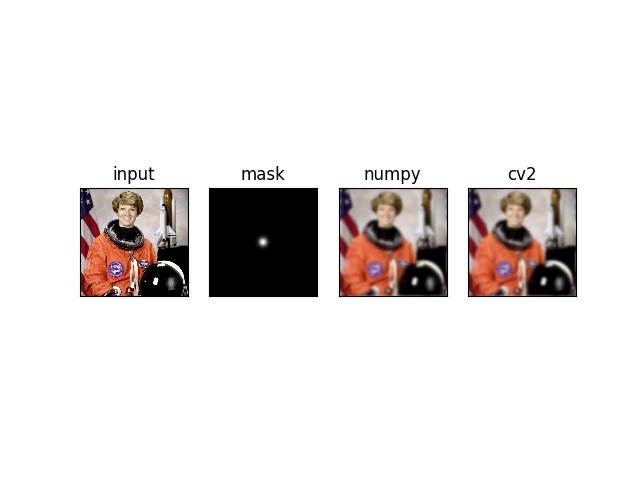

Example
>>> from kwimage.im_filter import * # NOQA >>> import kwimage >>> img_hwc = kwimage.grab_test_image(space='gray') >>> mask = kwimage.gaussian_patch(img_hwc.shape[0:2]) >>> mask = 1 - (mask / mask.max()) ** 32 >>> out_hwc_cv2 = fourier_mask(img_hwc, mask, backend='numpy') >>> out_hwc_np = fourier_mask(img_hwc, mask, backend='cv2') >>> # xdoctest: REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img_hwc, pnum=(1, 4, 1), fnum=1, title='input') >>> kwplot.imshow(mask, pnum=(1, 4, 2), fnum=1, title='mask') >>> kwplot.imshow(out_hwc_cv2, pnum=(1, 4, 3), fnum=1, title='numpy') >>> kwplot.imshow(out_hwc_np, pnum=(1, 4, 4), fnum=1, title='cv2') >>> kwplot.show_if_requested()
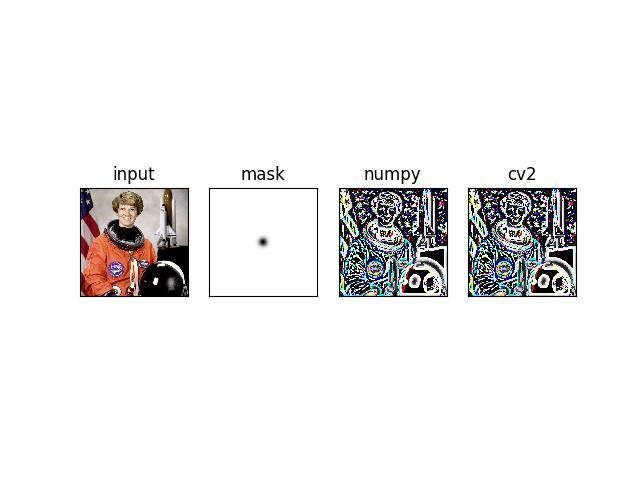

- kwimage.gaussian_blur(image, kernel=None, sigma=None, border_mode=None, dst=None)[source]¶
Apply a gausian blur to an image.
This is a simple wrapper around
cv2.GaussianBlur()with concise parametarization and sane defaults.- Parameters:
image (ndarray) – the input image
kernel (int | Tuple[int, int]) – The kernel size in x and y directions.
sigma (float | Tuple[float, float]) – The gaussian spread in x and y directions.
border_mode (str | int | None) – Border text code or cv2 integer. Border codes are ‘constant’ (default), ‘replicate’, ‘reflect’, ‘reflect101’, and ‘transparent’.
dst (ndarray | None) – optional inplace-output array.
- Returns:
the blurred image
- Return type:
ndarray
Example
>>> import kwimage >>> image = kwimage.ensure_float01(kwimage.grab_test_image('astro')) >>> blurred1 = kwimage.gaussian_blur(image) >>> blurred2 = kwimage.gaussian_blur(image, kernel=9) >>> blurred3 = kwimage.gaussian_blur(image, sigma=2) >>> blurred4 = kwimage.gaussian_blur(image, sigma=(2, 5), kernel=5) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=4, nCols=1) >>> blurs = [blurred1, blurred2, blurred3, blurred4] >>> for blurred in blurs: >>> diff = np.abs(image - blurred) >>> stack = kwimage.stack_images([image, blurred, diff], pad=10, axis=1) >>> kwplot.imshow(stack, pnum=pnum_()) >>> kwplot.show_if_requested()

- kwimage.gaussian_patch(shape=(7, 7), sigma=None)[source]¶
Creates a 2D gaussian patch with a specific size and sigma
- Parameters:
shape (Tuple[int, int]) – patch height and width
sigma (float | Tuple[float, float] | None) – Gaussian standard deviation. If unspecified, it is derived using the formula
0.3 * ((s - 1) * 0.5 - 1) + 0.8as described in [Cv2GaussKern].
- Returns:
ndarray
References
Todo
[ ] Look into this C-implementation https://kwgitlab.kitware.com/computer-vision/heatmap/blob/master/heatmap/heatmap.c
CommandLine
xdoctest -m kwimage.im_cv2 gaussian_patch --show
Example
>>> import numpy as np >>> shape = (88, 24) >>> sigma = None # 1.0 >>> gausspatch = gaussian_patch(shape, sigma) >>> sum_ = gausspatch.sum() >>> assert np.all(np.isclose(sum_, 1.0)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> norm = (gausspatch - gausspatch.min()) / (gausspatch.max() - gausspatch.min()) >>> kwplot.imshow(norm) >>> kwplot.show_if_requested()

Example
>>> import numpy as np >>> shape = (24, 24) >>> sigma = 3.0 >>> gausspatch = gaussian_patch(shape, sigma) >>> sum_ = gausspatch.sum() >>> assert np.all(np.isclose(sum_, 1.0)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> norm = (gausspatch - gausspatch.min()) / (gausspatch.max() - gausspatch.min()) >>> kwplot.imshow(norm) >>> kwplot.show_if_requested()

- kwimage.grab_test_image(key='astro', space='rgb', dsize=None, interpolation='lanczos')[source]¶
Ensures that the test image exists (this might use the network), reads it and returns the the image pixels.
- Parameters:
key (str) – which test image to grab. Valid choices are: astro - an astronaught carl - Carl Sagan paraview - ParaView logo stars - picture of stars in the sky airport - SkySat image of Beijing Capital International Airport on 18 February 2018 See
kwimage.grab_test_image.keysfor a full list.space (str) – which colorspace to return in. Defaults to ‘rgb’
dsize (Tuple[int, int]) – if specified resizes image to this size
- Returns:
the requested image
- Return type:
ndarray
CommandLine
xdoctest -m kwimage.im_demodata grab_test_image
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> key_to_image = {} >>> for key in kwimage.grab_test_image.keys(): >>> print('attempt to grab key = {!r}'.format(key)) >>> # specifying dsize will returned a resized variant >>> imdata = kwimage.grab_test_image(key, dsize=(256, None)) >>> key_to_image[key] = imdata >>> print('grabbed key = {!r}'.format(key)) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autoplt() >>> to_stack = [kwimage.draw_header_text( >>> imdata, text=key, color='kw_blue') >>> for key, imdata in key_to_image.items()] >>> stacked = kwimage.stack_images_grid(to_stack, bg_value='kw_darkgray') >>> stacked = kwimage.draw_header_text(stacked, 'kwimage.grab_test_image', fit=True, color='kitware_green') >>> kwplot.imshow(stacked)
- kwimage.grab_test_image_fpath(key='astro', dsize=None, overviews=None)[source]¶
Ensures that the test image exists (this might use the network) and returns the cached filepath to the requested image.
- Parameters:
key (str) – which test image to grab. Valid choices are: astro - an astronaught carl - Carl Sagan paraview - ParaView logo stars - picture of stars in the sky OR can be an existing path to an image
dsize (None | Tuple[int, int]) – if specified, we will return a variant of the data with the specific dsize
overviews (None | int) – if specified, will return a variant of the data with overviews
- Returns:
path to the requested image
- Return type:
CommandLine
python -c "import kwimage; print(kwimage.grab_test_image_fpath('airport'))"
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> for key in kwimage.grab_test_image.keys(): ... print('attempt to grab key = {!r}'.format(key)) ... kwimage.grab_test_image_fpath(key) ... print('grabbed grab key = {!r}'.format(key))
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> key = ub.peek(kwimage.grab_test_image.keys()) >>> # specifying a dsize will construct a new image >>> fpath1 = kwimage.grab_test_image_fpath(key) >>> fpath2 = kwimage.grab_test_image_fpath(key, dsize=(32, 16)) >>> print('fpath1 = {}'.format(ub.urepr(fpath1, nl=1))) >>> print('fpath2 = {}'.format(ub.urepr(fpath2, nl=1))) >>> assert fpath1 != fpath2 >>> imdata2 = kwimage.imread(fpath2) >>> assert imdata2.shape[0:2] == (16, 32)
- kwimage.imcrop(img, dsize, about=None, origin=None, border_value=None, interpolation='nearest')[source]¶
Crop an image about a specified point, padding if necessary.
This is like
PIL.Image.Image.crop()with more convenient arguments, orcv2.getRectSubPix()without the baked-in bilinear interpolation.- Parameters:
img (ndarray) – image to crop
dsize (Tuple[None | int, None | int]) – the desired width and height of the new image. If a dimension is None, then it is automatically computed to preserve aspect ratio. This can be larger than the original dims; if so, the cropped image is padded with border_value.
about (Tuple[str | int, str | int]) – the location to crop about. Mutually exclusive with origin. Defaults to top left. If ints (w,h) are provided, that will be the center of the cropped image. There are also string codes available: ‘lt’: make the top left point of the image the top left point of the cropped image. This is equivalent to
img[:dsize[1], :dsize[0]], plus padding. ‘rb’: make the bottom right point of the image the bottom right point of the cropped image. This is equivalent toimg[-dsize[1]:, -dsize[0]:], plus padding. ‘cc’: make the center of the image the center of the cropped image. Any combination of these codes can be used, ex. ‘lb’, ‘ct’, (‘r’, 200), …origin (Tuple[int, int] | None) – the origin of the crop in (x,y) order (same order as dsize/about). Mutually exclusive with about. Defaults to top left.
border_value (Number | Tuple | str) – any border border_value accepted by cv2.copyMakeBorder, ex. [255, 0, 0] (blue). Default is 0.
interpolation (str) – Can be ‘nearest’, in which case integral cropping is used. Can also be ‘linear’, in which case cv2.getRectSubPix is used. Defaults to ‘nearest’.
- Returns:
the cropped image
- Return type:
ndarray
- SeeAlso:
kwarray.padded_slice()- a similar function for working with“negative slices”.
Example
>>> import kwimage >>> import numpy as np >>> # >>> img = kwimage.grab_test_image('astro', dsize=(32, 32))[..., 0:3] >>> # >>> # regular crop >>> new_img1 = kwimage.imcrop(img, dsize=(5,6)) >>> assert new_img1.shape[0:2] == (6, 5) >>> # >>> # padding for coords outside the image bounds >>> new_img2 = kwimage.imcrop(img, dsize=(5,6), >>> origin=(-1,0), border_value=[1, 0, 0]) >>> assert np.all(new_img2[:, 0, 0:3] == [1, 0, 0]) >>> # >>> # codes for corner- and edge-centered cropping >>> new_img3 = kwimage.imcrop(img, dsize=(5,6), >>> about='cb') >>> # >>> # special code for bilinear interpolation >>> # with floating-point coordinates >>> new_img4 = kwimage.imcrop(img, dsize=(5,6), >>> about=(5.5, 8.5), interpolation='linear') >>> # >>> # use with bounding boxes >>> bbox = kwimage.Boxes.random(scale=5, rng=132).to_xywh().quantize() >>> origin, dsize = np.split(bbox.data[0], 2) >>> new_img5 = kwimage.imcrop(img, dsize=dsize, >>> origin=origin) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nSubplots=6) >>> kwplot.imshow(img, pnum=pnum_()) >>> kwplot.imshow(new_img1, pnum=pnum_()) >>> kwplot.imshow(new_img2, pnum=pnum_()) >>> kwplot.imshow(new_img3, pnum=pnum_()) >>> kwplot.imshow(new_img4, pnum=pnum_()) >>> kwplot.imshow(new_img5, pnum=pnum_()) >>> kwplot.show_if_requested()

- kwimage.imread(fpath, space='auto', backend='auto', **kw)[source]¶
Reads image data in a specified format using some backend implementation.
- Parameters:
fpath (str) – path to the file to be read
space (str) – The desired colorspace of the image. Can by any colorspace accepted by convert_colorspace, or it can be ‘auto’, in which case the colorspace of the image is unmodified (except in the case where a color image is read by opencv, in which case we convert BGR to RGB by default). If None, then no modification is made to whatever backend is used to read the image. Defaults to ‘auto’.
New in version 0.7.10: when the backend does not resolve to “cv2” the “auto” space resolves to None, thus the image is read as-is.
backend (str) – which backend reader to use. By default the file extension is used to determine this, but it can be manually overridden. Valid backends are ‘gdal’, ‘skimage’, ‘itk’, ‘pil’, and ‘cv2’. Defaults to ‘auto’.
**kw – backend-specific arguments
- The gdal backend accepts:
overview, ignore_color_table, nodata_method, band_indices
- Returns:
the image data in the specified color space.
- Return type:
ndarray
Note
if space is something non-standard like HSV or LAB, then the file must be a normal 8-bit color image, otherwise an error will occur.
Note
Some backends will respect EXIF orientation (skimage) and others will not (gdal, cv2).
The scikit-image backend is itself another multi-backend plugin-based image reader/writer.
- Raises:
IOError - If the image cannot be read –
ImportError - If trying to read a nitf without gdal –
NotImplementedError - if trying to read a corner-case image –
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> import ubelt as ub >>> # Test a non-standard image, which encodes a depth map >>> fpath = ub.grabdata( >>> 'http://www.topcoder.com/contest/problem/UrbanMapper3D/JAX_Tile_043_DTM.tif', >>> hasher='sha256', hash_prefix='64522acba6f0fb7060cd4c202ed32c5163c34e63d386afdada4190cce51ff4d4') >>> img1 = kwimage.imread(fpath) >>> # Check that write + read preserves data >>> dpath = ub.Path.appdir('kwimage/test/imread').ensuredir() >>> tmp_fpath = dpath / 'tmp0.tif' >>> kwimage.imwrite(tmp_fpath, img1) >>> img2 = kwimage.imread(tmp_fpath) >>> assert np.all(img2 == img1) >>> tmp_fpath.delete() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img1, pnum=(1, 2, 1), fnum=1, norm=True, title='tif orig') >>> kwplot.imshow(img2, pnum=(1, 2, 2), fnum=1, norm=True, title='tif io round-trip')
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> import ubelt as ub >>> img1 = kwimage.imread(ub.grabdata( >>> 'http://i.imgur.com/iXNf4Me.png', fname='ada.png', hasher='sha256', >>> hash_prefix='898cf2588c40baf64d6e09b6a93b4c8dcc0db26140639a365b57619e17dd1c77')) >>> dpath = ub.Path.appdir('kwimage/test/imread').ensuredir() >>> tmp_tif_fpath = dpath / 'tmp1.tif' >>> tmp_png_fpath = dpath / 'tmp1.png' >>> kwimage.imwrite(tmp_tif_fpath, img1) >>> kwimage.imwrite(tmp_png_fpath, img1) >>> tif_im = kwimage.imread(tmp_tif_fpath) >>> png_im = kwimage.imread(tmp_png_fpath) >>> assert np.all(tif_im == png_im) >>> tmp_tif_fpath.delete() >>> tmp_png_fpath.delete() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(png_im, pnum=(1, 2, 1), fnum=1, title='tif io') >>> kwplot.imshow(tif_im, pnum=(1, 2, 2), fnum=1, title='png io')
Example
>>> # xdoctest: +REQUIRES(--network) >>> import kwimage >>> import ubelt as ub >>> # FIXME: Dead link... >>> tif_fpath = ub.grabdata( >>> 'https://ghostscript.com/doc/tiff/test/images/rgb-3c-16b.tiff', >>> fname='pepper.tif', hasher='sha256', >>> hash_prefix='31ff3a1f416cb7281acfbcbb4b56ee8bb94e9f91489602ff2806e5a49abc03c0') >>> img1 = kwimage.imread(tif_fpath) >>> dpath = ub.Path.appdir('kwimage/test/imread').ensuredir() >>> tmp_tif_fpath = dpath / 'tmp2.tif' >>> tmp_png_fpath = dpath / 'tmp2.png' >>> kwimage.imwrite(tmp_tif_fpath, img1) >>> kwimage.imwrite(tmp_png_fpath, img1) >>> tif_im = kwimage.imread(tmp_tif_fpath) >>> png_im = kwimage.imread(tmp_png_fpath) >>> assert np.all(tif_im == png_im) >>> tmp_tif_fpath.delete() >>> tmp_png_fpath.delete() >>> assert np.all(tif_im == png_im) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(png_im / 2 ** 16, pnum=(1, 2, 1), fnum=1) >>> kwplot.imshow(tif_im / 2 ** 16, pnum=(1, 2, 2), fnum=1)
Example
>>> # xdoctest: +REQUIRES(module:itk, --network) >>> import kwimage >>> import ubelt as ub >>> # Grab an image that ITK can read >>> fpath = ub.grabdata( >>> url='https://data.kitware.com/api/v1/file/606754e32fa25629b9476f9e/download', >>> fname='brainweb1e5a10f17Rot20Tx20.mha', >>> hash_prefix='08f0812591691ae24a29788ba8cd1942e91', hasher='sha512') >>> # Read the image (this is actually a DxHxW stack of images) >>> img1_stack = kwimage.imread(fpath) >>> # Check that write + read preserves data >>> dpath = ub.Path.appdir('kwimage/test/imread').ensuredir() >>> tmp_fpath = dpath / 'tmp3.mha' >>> kwimage.imwrite(tmp_fpath, img1_stack) >>> recon = kwimage.imread(tmp_fpath) >>> assert not np.may_share_memory(recon, img1_stack) >>> assert np.all(recon == img1_stack) >>> tmp_fpath.delete() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(kwimage.stack_images_grid(recon[0::20]), >>> title='kwimage.imread with a .mha file') >>> kwplot.show_if_requested()
Benchmark
>>> import timerit >>> import kwimage >>> import ubelt as ub >>> # >>> dsize = (1920, 1080) >>> img1 = kwimage.grab_test_image('amazon', dsize=dsize) >>> ti = timerit.Timerit(10, bestof=3, verbose=1, unit='us') >>> formats = {} >>> dpath = ub.Path.appdir('kwimage/bench/im_io').ensuredir() >>> space = 'auto' >>> formats['png'] = kwimage.imwrite(join(dpath, '.png'), img1, space=space, backend='cv2') >>> formats['jpg'] = kwimage.imwrite(join(dpath, '.jpg'), img1, space=space, backend='cv2') >>> formats['tif_raw'] = kwimage.imwrite(join(dpath, '.raw.tif'), img1, space=space, backend='gdal', compress='RAW') >>> formats['tif_deflate'] = kwimage.imwrite(join(dpath, '.deflate.tif'), img1, space=space, backend='gdal', compress='DEFLATE') >>> formats['tif_lzw'] = kwimage.imwrite(join(dpath, '.lzw.tif'), img1, space=space, backend='gdal', compress='LZW') >>> grid = [ >>> ('cv2', 'png'), >>> ('cv2', 'jpg'), >>> ('gdal', 'jpg'), >>> ('turbojpeg', 'jpg'), >>> ('gdal', 'tif_raw'), >>> ('gdal', 'tif_lzw'), >>> ('gdal', 'tif_deflate'), >>> ('skimage', 'tif_raw'), >>> ] >>> backend, filefmt = 'cv2', 'png' >>> for backend, filefmt in grid: >>> for timer in ti.reset(f'imread-{filefmt}-{backend}'): >>> with timer: >>> kwimage.imread(formats[filefmt], space=space, backend=backend) >>> # Test all formats in auto mode >>> for filefmt in formats.keys(): >>> for timer in ti.reset(f'kwimage.imread-{filefmt}-auto'): >>> with timer: >>> kwimage.imread(formats[filefmt], space=space, backend='auto') >>> ti.measures = ub.map_vals(ub.sorted_vals, ti.measures) >>> print('ti.measures = {}'.format(ub.urepr(ti.measures['min'], nl=2, align=':'))) Timed best=42891.504 µs, mean=44008.439 ± 1409.2 µs for imread-png-cv2 Timed best=33146.808 µs, mean=34185.172 ± 656.3 µs for imread-jpg-cv2 Timed best=40120.306 µs, mean=41220.927 ± 1010.9 µs for imread-jpg-gdal Timed best=30798.162 µs, mean=31573.070 ± 737.0 µs for imread-jpg-turbojpeg Timed best=6223.170 µs, mean=6370.462 ± 150.7 µs for imread-tif_raw-gdal Timed best=42459.404 µs, mean=46519.940 ± 5664.9 µs for imread-tif_lzw-gdal Timed best=36271.175 µs, mean=37301.108 ± 861.1 µs for imread-tif_deflate-gdal Timed best=5239.503 µs, mean=6566.574 ± 1086.2 µs for imread-tif_raw-skimage ti.measures = { 'imread-tif_raw-skimage' : 0.0052395030070329085, 'imread-tif_raw-gdal' : 0.006223169999429956, 'imread-jpg-turbojpeg' : 0.030798161998973228, 'imread-jpg-cv2' : 0.03314680799667258, 'imread-tif_deflate-gdal': 0.03627117499127053, 'imread-jpg-gdal' : 0.040120305988239124, 'imread-tif_lzw-gdal' : 0.042459404008695856, 'imread-png-cv2' : 0.042891503995633684, }
- kwimage.imresize(img, scale=None, dsize=None, max_dim=None, min_dim=None, interpolation=None, grow_interpolation=None, letterbox=False, return_info=False, antialias=False, border_value=0)[source]¶
Resize an image via a scale factor, final size, or size and aspect ratio.
Wraps and generalizes cv2.resize, allows for specification of either a scale factor, a final size, or the final size for a particular dimension.
Note
As described in [ResizeConfusion], this each entry in the image array as representing the center of a pixel. This is the pixels_are=’area’ approach, or align_corners=False in pytorch. It is equivalent to a shift and scale in warp_affine (which by default uses align corners).
Note
The border mode cannot be specified here and seems to always be reflect in the underlying cv2 implementation.
- Parameters:
img (ndarray) – image to resize
scale (float | Tuple[float, float]) – Desired floating point scale factor. If a tuple, the dimension ordering is x,y. Mutually exclusive with dsize, min_dim, max_dim.
dsize (Tuple[int | None, int | None] | None) – The desired width and height of the new image. If a dimension is None, then it is automatically computed to preserve aspect ratio. Mutually exclusive with scale, min_dim, max_dim.
max_dim (int) – New size of the maximum dimension, the other dimension is scaled to maintain aspect ratio. Mutually exclusive with scale, dsize, min_dim.
min_dim (int) – New size of the minimum dimension, the other dimension is scaled to maintain aspect ratio. Mutually exclusive with scale, dsize, max_dim.
interpolation (str | int) – The interpolation key or code (e.g. linear lanczos). By default “area” is used if the image is shrinking and “lanczos” is used if the image is growing. Note, if this is explicitly set, then it will be used regardless of if the image is growing or shrinking. Set
grow_interpolationto change the default for an enlarging interpolation.grow_interpolation (str | int) – The interpolation key or code to use when the image is being enlarged. Does nothing if “interpolation” is explicitly given. If “interpolation” is not specified “area” is used when shrinking. Defaults to “lanczos”.
letterbox (bool) – If used in conjunction with dsize, then the image is scaled and translated to fit in the center of the new image while maintaining aspect ratio. Border padding is added if necessary. Defaults to False.
return_info (bool) – if True returns information about the final transformation in a dictionary. If there is an offset, the scale is applied before the offset when transforming to the new resized space. Defaults to False.
antialias (bool) – if True blurs to anti-alias before downsampling. Defaults to False.
border_value (int | float | Iterable[int | float]) – if letterbox is True, this is used as the constant fill value.
- Returns:
the new image and optionally an info dictionary if return_info=True
- Return type:
ndarray | Tuple[ndarray, Dict]
References
Example
>>> import kwimage >>> import numpy as np >>> # Test scale >>> img = np.zeros((16, 10, 3), dtype=np.uint8) >>> new_img, info = kwimage.imresize(img, scale=.85, >>> interpolation='area', >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['scale'].tolist() == [.8, 0.875] >>> # Test dsize without None >>> new_img, info = kwimage.imresize(img, dsize=(5, 12), >>> interpolation='area', >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['scale'].tolist() == [0.5 , 0.75] >>> # Test dsize with None >>> new_img, info = kwimage.imresize(img, dsize=(6, None), >>> interpolation='area', >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['scale'].tolist() == [0.6, 0.625] >>> # Test max_dim >>> new_img, info = kwimage.imresize(img, max_dim=6, >>> interpolation='area', >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['scale'].tolist() == [0.4 , 0.375] >>> # Test min_dim >>> new_img, info = kwimage.imresize(img, min_dim=6, >>> interpolation='area', >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['scale'].tolist() == [0.6 , 0.625]
Example
>>> import kwimage >>> import numpy as np >>> # Test letterbox resize >>> img = np.ones((5, 10, 3), dtype=np.float32) >>> new_img, info = kwimage.imresize(img, dsize=(19, 19), >>> letterbox=True, >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['offset'].tolist() == [0, 4] >>> img = np.ones((10, 5, 3), dtype=np.float32) >>> new_img, info = kwimage.imresize(img, dsize=(19, 19), >>> letterbox=True, >>> return_info=True) >>> print('info = {!r}'.format(info)) >>> assert info['offset'].tolist() == [4, 0]
>>> import kwimage >>> import numpy as np >>> # Test letterbox resize >>> img = np.random.rand(100, 200) >>> new_img, info = kwimage.imresize(img, dsize=(300, 300), letterbox=True, return_info=True)
Example
>>> # Check aliasing >>> import kwimage >>> #img = kwimage.grab_test_image('checkerboard') >>> img = kwimage.grab_test_image('pm5644') >>> # test with nans >>> img = kwimage.ensure_float01(img) >>> img[100:200, 400:700] = np.nan >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dsize = (14, 14) >>> dsize = (64, 64) >>> # When we set "grow_interpolation" for a "shrinking" resize it should >>> # still do the "area" interpolation to antialias the results. But if we >>> # use explicit interpolation it should alias. >>> pnum_ = kwplot.PlotNums(nSubplots=12, nCols=4) >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, interpolation='area'), pnum=pnum_(), title='resize aa area') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, interpolation='linear'), pnum=pnum_(), title='resize aa linear') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, interpolation='nearest'), pnum=pnum_(), title='resize aa nearest') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, interpolation='cubic'), pnum=pnum_(), title='resize aa cubic')
>>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, grow_interpolation='area'), pnum=pnum_(), title='resize aa grow area') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, grow_interpolation='linear'), pnum=pnum_(), title='resize aa grow linear') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, grow_interpolation='nearest'), pnum=pnum_(), title='resize aa grow nearest') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=True, grow_interpolation='cubic'), pnum=pnum_(), title='resize aa grow cubic')
>>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=False, interpolation='area'), pnum=pnum_(), title='resize no-aa area') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=False, interpolation='linear'), pnum=pnum_(), title='resize no-aa linear') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=False, interpolation='nearest'), pnum=pnum_(), title='resize no-aa nearest') >>> kwplot.imshow(kwimage.imresize(img, dsize=dsize, antialias=False, interpolation='cubic'), pnum=pnum_(), title='resize no-aa cubic')

Example
>>> # Test single pixel resize >>> import kwimage >>> import numpy as np >>> assert kwimage.imresize(np.random.rand(1, 1, 3), scale=3).shape == (3, 3, 3) >>> assert kwimage.imresize(np.random.rand(1, 1), scale=3).shape == (3, 3)
# cv2.resize(np.random.rand(1, 1, 3), (3, 3))
Todo
[X] When interpolation is area and the number of channels > 4 cv2.resize will error but it is fine for linear interpolation
[ ] TODO: add padding options when letterbox=True
[ ] Allow for pre-clipping when letterbox=True
- kwimage.imscale(img, scale, interpolation=None, return_scale=False)[source]¶
DEPRECATED and removed: use imresize instead
- kwimage.imwrite(fpath, image, space='auto', backend='auto', **kwargs)[source]¶
Writes image data to disk.
- Parameters:
fpath (PathLike) – location to save the image
image (ndarray) – image data
space (str | None) – the colorspace of the image to save. Can by any colorspace accepted by convert_colorspace, or it can be ‘auto’, in which case we assume the input image is either RGB, RGBA or grayscale. If None, then absolutely no color modification is made and whatever backend is used writes the image as-is.
New in version 0.7.10: when the backend does not resolve to “cv2”, the “auto” space resolves to None, thus the image is saved as-is.
backend (str) – Which backend writer to use. By default the file extension is used to determine this. Valid backends are ‘gdal’, ‘skimage’, ‘itk’, and ‘cv2’.
**kwargs – args passed to the backend writer. When the backend is gdal, available options are: compress (str): Common options are auto, DEFLATE, LZW, JPEG. blocksize (int): size of tiled blocks (e.g. 256) overviews (None | str | int | list): Number of overviews. overview_resample (str): Common options NEAREST, CUBIC, LANCZOS options (List[str]): other gdal options. nodata (int): denotes a integer value as nodata. metadata (dict): the metadata for the default empty domain. transform (kwimage.Affine): Transform to CRS from pixel space crs (str): The coordinate reference system for transform. See
_imwrite_cloud_optimized_geotiff()for more details each options. When the backend is itk, seeitk.imwrite()for options When the backend is skimage, seeskimage.io.imsave()for options When the backend is cv2 seecv2.imwrite()for options.
- Returns:
path to the written file
- Return type:
Note
The image may be modified to preserve its colorspace depending on which backend is used to write the image.
When saving as a jpeg or png, the image must be encoded with the uint8 data type. When saving as a tiff, any data type is allowed.
The scikit-image backend is itself another multi-backend plugin-based image reader/writer.
- Raises:
Exception – if the image cannot be written
Example
>>> # xdoctest: +REQUIRES(--network) >>> # This should be moved to a unit test >>> from kwimage.im_io import _have_gdal # NOQA >>> import kwimage >>> import ubelt as ub >>> import uuid >>> dpath = ub.Path.appdir('kwimage/test/imwrite').ensuredir() >>> test_image_paths = [ >>> #ub.grabdata('https://ghostscript.com/doc/tiff/test/images/rgb-3c-16b.tiff', fname='pepper.tif'), >>> ub.grabdata('http://i.imgur.com/iXNf4Me.png', fname='ada.png'), >>> #ub.grabdata('http://www.topcoder.com/contest/problem/UrbanMapper3D/JAX_Tile_043_DTM.tif'), >>> ub.grabdata('https://upload.wikimedia.org/wikipedia/commons/f/fa/Grayscale_8bits_palette_sample_image.png', fname='parrot.png') >>> ] >>> for fpath in test_image_paths: >>> for space in ['auto', 'rgb', 'bgr', 'gray', 'rgba']: >>> img1 = kwimage.imread(fpath, space=space) >>> print('Test im-io consistency of fpath = {!r} in {} space, shape={}'.format(fpath, space, img1.shape)) >>> # Write the image in TIF and PNG format >>> tmp_tif_fpath = dpath / (str(uuid.uuid4()) + '.tif') >>> tmp_png_fpath = dpath / (str(uuid.uuid4()) + '.png') >>> kwimage.imwrite(tmp_tif_fpath, img1, space=space, backend='skimage') >>> kwimage.imwrite(tmp_png_fpath, img1, space=space) >>> tif_im = kwimage.imread(tmp_tif_fpath, space=space) >>> png_im = kwimage.imread(tmp_png_fpath, space=space) >>> assert np.all(tif_im == png_im), 'im-read/write inconsistency' >>> if _have_gdal: >>> tmp_tif2_fpath = dpath / (str(uuid.uuid4()) + '.tif') >>> kwimage.imwrite(tmp_tif2_fpath, img1, space=space, backend='gdal') >>> tif_im2 = kwimage.imread(tmp_tif2_fpath, space=space) >>> assert np.all(tif_im == tif_im2), 'im-read/write inconsistency' >>> tmp_tif2_fpath.delete() >>> if space == 'gray': >>> assert tif_im.ndim == 2 >>> assert png_im.ndim == 2 >>> elif space in ['rgb', 'bgr']: >>> assert tif_im.shape[2] == 3 >>> assert png_im.shape[2] == 3 >>> elif space in ['rgba', 'bgra']: >>> assert tif_im.shape[2] == 4 >>> assert png_im.shape[2] == 4 >>> tmp_tif_fpath.delete() >>> tmp_png_fpath.delete()
Benchmark
>>> import timerit >>> import os >>> import kwimage >>> import tempfile >>> # >>> img1 = kwimage.grab_test_image('astro', dsize=(1920, 1080)) >>> space = 'auto' >>> # >>> file_sizes = {} >>> # >>> ti = timerit.Timerit(10, bestof=3, verbose=2) >>> # >>> for timer in ti.reset('imwrite-skimage-tif'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='skimage') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-cv2-png'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.png') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='cv2') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-cv2-jpg'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.jpg') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='cv2') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-gdal-raw'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='gdal', compress='RAW') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-gdal-lzw'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='gdal', compress='LZW') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-gdal-zstd'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='gdal', compress='ZSTD') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-gdal-deflate'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='gdal', compress='DEFLATE') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> for timer in ti.reset('imwrite-gdal-jpeg'): >>> with timer: >>> tmp = tempfile.NamedTemporaryFile(suffix='.tif') >>> kwimage.imwrite(tmp.name, img1, space=space, backend='gdal', compress='JPEG') >>> file_sizes[ti.label] = os.stat(tmp.name).st_size >>> # >>> file_sizes = ub.sorted_vals(file_sizes) >>> import xdev >>> file_sizes_human = ub.map_vals(lambda x: xdev.byte_str(x, 'MB'), file_sizes) >>> print('ti.rankings = {}'.format(ub.urepr(ti.rankings, nl=2))) >>> print('file_sizes = {}'.format(ub.urepr(file_sizes_human, nl=1)))
Example
>>> # Test saving a multi-band file >>> import kwimage >>> import pytest >>> import ubelt as ub >>> dpath = ub.Path.appdir('kwimage/test/imwrite').ensuredir() >>> # In this case the backend will not resolve to cv2, so >>> # we should not need to specify space. >>> data = np.random.rand(32, 32, 13).astype(np.float32) >>> fpath = dpath / 'tmp1.tif' >>> kwimage.imwrite(fpath, data) >>> recon = kwimage.imread(fpath) >>> assert np.all(recon == data) >>> kwimage.imwrite(fpath, data, backend='skimage') >>> recon = kwimage.imread(fpath, backend='skimage') >>> assert np.all(recon == data) >>> # xdoctest: +REQUIRES(module:osgeo) >>> # gdal should error when trying to read an image written by skimage >>> with pytest.raises(NotImplementedError): >>> kwimage.imread(fpath, backend='gdal') >>> # In this case the backend will resolve to cv2, and thus we expect >>> # a failure >>> fpath = dpath / 'tmp1.png' >>> with pytest.raises(NotImplementedError): >>> kwimage.imwrite(fpath, data)
Example
>>> import ubelt as ub >>> import kwimage >>> dpath = ub.Path.appdir('kwimage/badwrite').ensuredir() >>> dpath.delete().ensuredir() >>> imdata = kwimage.ensure_uint255(kwimage.grab_test_image())[:, :, 0] >>> import pytest >>> fpath = dpath / 'does-not-exist/img.jpg' >>> with pytest.raises(IOError): ... kwimage.imwrite(fpath, imdata, backend='cv2') >>> with pytest.raises(IOError): ... kwimage.imwrite(fpath, imdata, backend='skimage') >>> # xdoctest: +SKIP >>> # TODO: run tests conditionally >>> with pytest.raises(IOError): ... kwimage.imwrite(fpath, imdata, backend='gdal') >>> with pytest.raises((IOError, RuntimeError)): ... kwimage.imwrite(fpath, imdata, backend='itk')
- kwimage.load_image_shape(fpath, backend='auto', include_channels=True)[source]¶
Determine the height/width/channels of an image without reading the entire file.
- Parameters:
fpath (str) – path to an image
backend (str) – can be “auto”, “pil”, or “gdal”.
include_channels (bool) – if False, only reads the height, width.
- Returns:
- Tuple[int, int, int] - shape of the image
Recall this library uses the convention that “shape” is refers to height,width,channels array-style ordering and “size” is width,height cv2-style ordering.
Example
>>> # xdoctest: +REQUIRES(module:osgeo) >>> # Test the loading the shape works the same as loading the image and >>> # testing the shape >>> import kwimage >>> temp_dpath = ub.Path.appdir('kwimage/tests/load_image_shape').ensuredir() >>> data = kwimage.grab_test_image() >>> datas = { >>> 'rgb255': kwimage.ensure_uint255(data), >>> 'rgb01': kwimage.ensure_float01(data), >>> 'rgba01': kwimage.ensure_alpha_channel(data), >>> } >>> results = {} >>> # These should be consistent >>> # The was a problem where CV2_IMREAD_UNCHANGED read the alpha band, >>> # but PIL did not, but maybe this is fixed now? >>> for key, imdata in datas.items(): >>> fpath = temp_dpath / f'{key}.png' >>> kwimage.imwrite(fpath, imdata) >>> shapes = {} >>> shapes['pil_load_shape'] = kwimage.load_image_shape(fpath, backend='pil') >>> shapes['gdal_load_shape'] = kwimage.load_image_shape(fpath, backend='gdal') >>> shapes['auto_load_shape'] = kwimage.load_image_shape(fpath, backend='auto') >>> shapes['pil'] = kwimage.imread(fpath, backend='pil').shape >>> shapes['cv2'] = kwimage.imread(fpath, backend='cv2').shape >>> shapes['gdal'] = kwimage.imread(fpath, backend='gdal').shape >>> shapes['skimage'] = kwimage.imread(fpath, backend='skimage').shape >>> results[key] = shapes >>> print('results = {}'.format(ub.urepr(results, nl=2, align=':', sort=0))) >>> for shapes in results.values(): >>> assert ub.allsame(shapes.values()) >>> temp_dpath.delete()
Benchmark
>>> # For large files, PIL is much faster >>> # xdoctest: +REQUIRES(module:osgeo) >>> from osgeo import gdal >>> from PIL import Image >>> import timerit >>> # >>> import kwimage >>> fpath = kwimage.grab_test_image_fpath() >>> # >>> ti = timerit.Timerit(100, bestof=10, verbose=2) >>> for timer in ti.reset('gdal'): >>> with timer: >>> gdal_dset = gdal.Open(fpath, gdal.GA_ReadOnly) >>> width = gdal_dset.RasterXSize >>> height = gdal_dset.RasterYSize >>> gdal_dset = None >>> # >>> for timer in ti.reset('PIL'): >>> with timer: >>> pil_img = Image.open(fpath) >>> width, height = pil_img.size >>> pil_img.close() >>> # xdoctest: +REQUIRES(module:imagesize) >>> # The imagesize module is quite fast >>> import imagesize >>> for timer in ti.reset('imagesize'): >>> with timer: >>> width, height = imagesize.get(fpath) Timed gdal for: 100 loops, best of 10 time per loop: best=54.423 µs, mean=72.761 ± 15.9 µs Timed PIL for: 100 loops, best of 10 time per loop: best=25.986 µs, mean=26.791 ± 1.2 µs Timed imagesize for: 100 loops, best of 10 time per loop: best=5.092 µs, mean=5.195 ± 0.1 µs
Example
>>> # xdoctest: +REQUIRES(module:osgeo) >>> import ubelt as ub >>> import kwimage >>> dpath = ub.Path.appdir('kwimage/tests', type='cache').ensuredir() >>> fpath = dpath / 'foo.tif' >>> kwimage.imwrite(fpath, np.random.rand(64, 64, 3)) >>> shape = kwimage.load_image_shape(fpath) >>> assert shape == (64, 64, 3)
- kwimage.make_channels_comparable(img1, img2, atleast3d=False)[source]¶
Broadcasts image arrays so they can have elementwise operations applied
- Parameters:
img1 (ndarray) – first image
img2 (ndarray) – second image
atleast3d (bool) – if true we ensure that the channel dimension exists (only relevant for 1-channel images). Defaults to False.
Example
>>> import itertools as it >>> wh_basis = [(5, 5), (3, 5), (5, 3), (1, 1), (1, 3), (3, 1)] >>> for w, h in wh_basis: >>> shape_basis = [(w, h), (w, h, 1), (w, h, 3)] >>> # Test all permutations of shap inputs >>> for shape1, shape2 in it.product(shape_basis, shape_basis): >>> print('* input shapes: %r, %r' % (shape1, shape2)) >>> img1 = np.empty(shape1) >>> img2 = np.empty(shape2) >>> img1, img2 = make_channels_comparable(img1, img2) >>> print('... output shapes: %r, %r' % (img1.shape, img2.shape)) >>> elem = (img1 + img2) >>> print('... elem(+) shape: %r' % (elem.shape,)) >>> assert elem.size == img1.size, 'outputs should have same size' >>> assert img1.size == img2.size, 'new imgs should have same size' >>> print('--------')
- kwimage.make_heatmask(probs, cmap='plasma', with_alpha=1.0, space='rgb', dsize=None)[source]¶
Colorizes a single-channel intensity mask (with an alpha channel)
- Parameters:
probs (ndarray) – 2D probability map with values between 0 and 1
cmap (str) – mpl colormap
with_alpha (float) – between 0 and 1, uses probs as the alpha multipled by this number.
space (str) – output colorspace
dsize (tuple) – if not None, then output is resized to W,H=dsize
- SeeAlso:
kwimage.overlay_alpha_images
Example
>>> # xdoctest: +REQUIRES(module:matplotlib) >>> from kwimage.im_draw import * # NOQA >>> probs = np.tile(np.linspace(0, 1, 10), (10, 1)) >>> heatmask = make_heatmask(probs, with_alpha=0.8, dsize=(100, 100)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(heatmask, fnum=1, doclf=True, colorspace='rgb', >>> title='make_heatmask') >>> kwplot.show_if_requested()
- kwimage.make_orimask(radians, mag=None, alpha=1.0)[source]¶
Makes a colormap in HSV space where the orientation changes color and mag changes the saturation/value.
- Parameters:
radians (ndarray) – orientation in radians
mag (ndarray) – magnitude (must be normalized between 0 and 1)
alpha (float | ndarray) – if False or None, then the image is returned without alpha if a float, then mag is scaled by this and used as the alpha channel if an ndarray, then this is explicilty set as the alpha channel
- Returns:
an rgb / rgba image in 01 space
- Return type:
ndarray[Any, Float32]
- SeeAlso:
kwimage.overlay_alpha_images
Example
>>> # xdoctest: +REQUIRES(module:matplotlib) >>> from kwimage.im_draw import * # NOQA >>> x, y = np.meshgrid(np.arange(64), np.arange(64)) >>> dx, dy = x - 32, y - 32 >>> radians = np.arctan2(dx, dy) >>> mag = np.sqrt(dx ** 2 + dy ** 2) >>> orimask = make_orimask(radians, mag) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(orimask, fnum=1, doclf=True, >>> colorspace='rgb', title='make_orimask') >>> kwplot.show_if_requested()
- kwimage.make_vector_field(dx, dy, stride=0.02, thresh=0.0, scale=1.0, alpha=1.0, color='strawberry', thickness=1, tipLength=0.1, line_type='aa')[source]¶
Create an image representing a 2D vector field.
- Parameters:
dx (ndarray) – grid of vector x components
dy (ndarray) – grid of vector y components
stride (int | float) – sparsity of vectors, int specifies stride step in pixels, a float specifies it as a percentage.
thresh (float) – only plot vectors with magnitude greater than thres
scale (float) – multiply magnitude for easier visualization
alpha (float) – alpha value for vectors. Non-vector regions receive 0 alpha (if False, no alpha channel is used)
color (str | tuple | kwimage.Color) – RGB color of the vectors
thickness (int) – thickness of arrows
tipLength (float) – fraction of line length
line_type (int | str) – either cv2.LINE_4, cv2.LINE_8, or cv2.LINE_AA or a string code.
- Returns:
vec_img - an rgb/rgba image in 0-1 space
- Return type:
ndarray[Any, Float32]
- SeeAlso:
kwimage.overlay_alpha_images
DEPRECATED USE: draw_vector_field instead
Example
>>> x, y = np.meshgrid(np.arange(512), np.arange(512)) >>> dx, dy = x - 256.01, y - 256.01 >>> radians = np.arctan2(dx, dy) >>> mag = np.sqrt(dx ** 2 + dy ** 2) >>> dx, dy = dx / mag, dy / mag >>> img = make_vector_field(dx, dy, scale=10, alpha=False) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img) >>> kwplot.show_if_requested()

- kwimage.morphology(data, mode, kernel=5, element='rect', iterations=1, border_mode='constant', border_value=0)[source]¶
Executes a morphological operation.
- Parameters:
input (ndarray[dtype=uint8 | float64]) – data (note if mode is hitmiss data must be uint8)
mode (str) – morphology mode, can be one of: ‘erode’, ‘dilate’, ‘open’, ‘close’, ‘gradient’, ‘tophat’, ‘blackhat’, or ‘hitmiss’.
kernel (ndarray | int | Tuple[int, int]) – size of the morphology kernel (w, h) to be constructed according to “element”. If the kernel size is 0, this function returns a copy of the data. Can also be a 2D array which is a custom structuring element. In this case “element” is ignored.
element (str) – structural element, can be ‘rect’, ‘cross’, or ‘ellipse’.
iterations (int) – numer of times to repeat the operation
border_mode (str | int) – Border code or cv2 integer. Border codes are constant (default) replicate, reflect, wrap, reflect101, and transparent.
border_value (int | float | Iterable[int | float]) – Used as the fill value if border_mode is constant. Otherwise this is ignored.
Example
>>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> #image = kwimage.grab_test_image(dsize=(380, 380)) >>> image = kwimage.Mask.demo().data * 255 >>> basis = { >>> 'mode': ['dilate'], >>> 'kernel': [5, (3, 7)], >>> 'element': ['rect', 'cross', 'ellipse'], >>> #'mode': ['dilate', 'erode'], >>> } >>> grid = list(ub.named_product(basis)) >>> grid += [{'mode': 'dilate', 'kernel': 0, 'element': 'rect', }] >>> grid += [{'mode': 'dilate', 'kernel': 'random', 'element': 'custom'}] >>> results = {} >>> for params in grid: ... key = ub.urepr(params, compact=1, si=0, nl=1) ... if params['kernel'] == 'random': ... params['kernel'] = np.random.rand(5, 5) ... results[key] = morphology(image, **params) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> to_stack = [] >>> canvas = image >>> canvas = kwimage.imresize(canvas, dsize=(380, 380), interpolation='nearest') >>> canvas = kwimage.draw_header_text(canvas, 'input', color='kitware_green') >>> to_stack.append(canvas) >>> for key, result in results.items(): >>> canvas = result >>> canvas = kwimage.imresize(canvas, dsize=(380, 380), interpolation='nearest') >>> canvas = kwimage.draw_header_text(canvas, key, color='kitware_green') >>> to_stack.append(canvas) >>> canvas = kwimage.stack_images_grid(to_stack, pad=10, bg_value='kitware_blue') >>> canvas = kwimage.draw_header_text(canvas, '--- kwimage.morphology demo ---', color='kitware_green') >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()

Example
>>> from kwimage.im_cv2 import * # NOQA >>> from kwimage.im_cv2 import _CV2_MORPH_MODES # NOQA >>> from kwimage.im_cv2 import _CV2_STRUCT_ELEMENTS # NOQA >>> #shape = (32, 32) >>> shape = (64, 64) >>> data = (np.random.rand(*shape) > 0.5).astype(np.uint8) >>> import kwimage >>> data = kwimage.gaussian_patch(shape) >>> data = data / data.max() >>> data = kwimage.ensure_uint255(data) >>> results = {} >>> kernel = 5 >>> for mode in _CV2_MORPH_MODES.keys(): ... for element in _CV2_STRUCT_ELEMENTS.keys(): ... results[f'{mode}+{element}'] = morphology(data, mode, kernel=kernel, element=element, iterations=2) >>> results['raw'] = data >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nCols=3, nSubplots=len(results)) >>> for k, result in results.items(): >>> kwplot.imshow(result, pnum=pnum_(), title=k) >>> kwplot.show_if_requested()
References
- kwimage.nodata_checkerboard(canvas, square_shape=8, on_value='auto', off_value='auto')[source]¶
Fills nans or masked values with a checkerbord pattern.
- Parameters:
canvas (ndarray) – A 2D image with any number of channels that may be a masked array or contain nan values.
square_shape (int) – the pixel size of the checkers
on_value (Number | str) – The value of one checker. Defaults to 1 for floats and 255 for ints.
off_value (Number | str) – The value off the other checker. Defaults to 0.
- Returns:
- an output array with imputed values.
if the input was a masked array, the mask will still exist.
- Return type:
ndarray
- SeeAlso:
fill_nans_with_checkers()- similar, but only operates on nan values.
Example
>>> import kwimage >>> # Test a masked array WITH nan values >>> data = kwimage.grab_test_image(space='rgb') >>> na_circle = kwimage.Polygon.circle((256 - 96, 256), 128) >>> ma_circle = kwimage.Polygon.circle((256 + 96, 256), 128) >>> ma_mask = na_circle.fill(np.zeros(data.shape, dtype=np.uint8), value=1).astype(bool) >>> na_mask = ma_circle.fill(np.zeros(data.shape, dtype=np.uint8), value=1).astype(bool) >>> # Hack the channels to make a ven diagram >>> ma_mask[..., [0, 1]] = False >>> na_mask[..., [0, 2]] = False >>> data = kwimage.ensure_float01(data) >>> data[na_mask] = np.nan >>> canvas = np.ma.MaskedArray(data, ma_mask) >>> kwimage.draw_text_on_image(canvas, 'masked values', (256 - 96, 256 - 128), halign='center', valign='bottom', border=2) >>> kwimage.draw_text_on_image(canvas, 'nan values', (256 + 96, 256 + 128), halign='center', valign='top', border=2) >>> kwimage.draw_text_on_image(canvas, 'kwimage.nodata_checkerboard', (256, 5), halign='center', valign='top', border=2) >>> kwimage.draw_text_on_image(canvas, '(pip install kwimage)', (512, 512 - 10), halign='right', valign='bottom', border=2, fontScale=0.8) >>> result = kwimage.nodata_checkerboard(canvas, on_value=0.5) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(result) >>> kwplot.show_if_requested()

Example
>>> # Simple test with a masked array >>> import kwimage >>> data = kwimage.grab_test_image(space='rgb', dsize=(64, 64)) >>> data = kwimage.ensure_uint255(data) >>> circle = kwimage.Polygon.circle((32, 32), 16) >>> mask = circle.fill(np.zeros(data.shape, dtype=np.uint8), value=1).astype(bool) >>> img = np.ma.MaskedArray(data, mask) >>> canvas = img.copy() >>> result = kwimage.nodata_checkerboard(canvas) >>> canvas.data is result.data >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(result, title='nodata_checkers with masked uint8') >>> kwplot.show_if_requested()
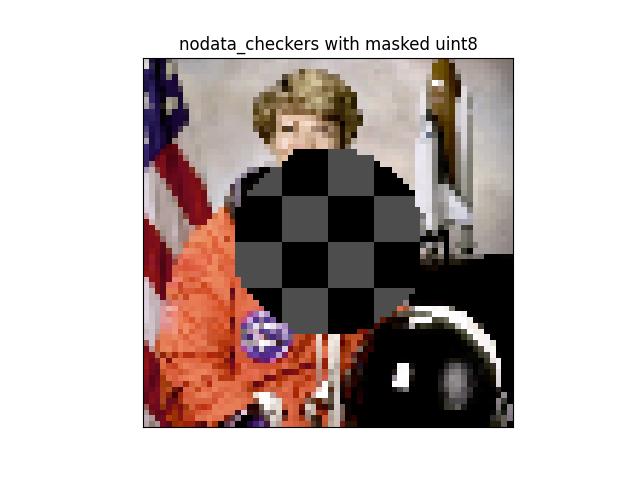

Example
>>> # Simple test with a bad mask >>> import kwimage >>> data = kwimage.grab_test_image(space='rgb', dsize=(64, 64)) >>> data = kwimage.ensure_uint255(data) >>> circle = kwimage.Polygon.circle((32, 32), 16) >>> mask = circle.fill(np.zeros(data.shape, dtype=np.uint8), value=1).astype(bool) >>> img = np.ma.MaskedArray(data, mask) >>> img.__dict__['_mask'] = np.empty((), dtype=bool) >>> import pytest >>> with pytest.raises(Exception): ... result = kwimage.nodata_checkerboard(img)
- kwimage.non_max_supression(ltrb, scores, thresh, bias=0.0, classes=None, impl='auto', device_id=None)[source]¶
Non-Maximum Suppression - remove redundant bounding boxes
Based on information from [CythonNMS] and [NMSPython].
- Parameters:
ltrb (ndarray[Any, Float32]) – Float32 array of shape Nx4 representing boxes in ltrb format
scores (ndarray[Any, Float32]) – Float32 array of shape N representing scores for each box
thresh (float) – iou threshold. Boxes are removed if they overlap greater than this threshold (i.e. Boxes are removed if iou > threshold). Thresh = 0 is the most strict, resulting in the fewest boxes, and 1 is the most permissive resulting in the most.
bias (float) – bias for iou computation either 0 or 1
classes (ndarray[Shape[’’], Int64] | None*) – integer classes. If specified NMS is done on a perclass basis.
impl (str) – implementation can be “auto”, “python”, “cython_cpu”, “gpu”, “torch”, or “torchvision”.
device_id (int) – used if impl is gpu, device id to work on. If not specified torch.cuda.current_device() is used.
Note
Using impl=’cython_gpu’ may result in an CUDA memory error that is not exposed to the python processes. In other words your program will hard crash if impl=’cython_gpu’, and you feed it too many bounding boxes. Ideally this will be fixed in the future.
TODO: SoftNMS [SoftNMS].
References
CommandLine
xdoctest -m ~/code/kwimage/kwimage/algo/algo_nms.py non_max_supression
Example
>>> from kwimage.algo.algo_nms import * >>> from kwimage.algo.algo_nms import _impls >>> ltrb = np.array([ >>> [0, 0, 100, 100], >>> [100, 100, 10, 10], >>> [10, 10, 100, 100], >>> [50, 50, 100, 100], >>> ], dtype=np.float32) >>> scores = np.array([.1, .5, .9, .1]) >>> keep = non_max_supression(ltrb, scores, thresh=0.5, impl='numpy') >>> print('keep = {!r}'.format(keep)) >>> assert keep == [2, 1, 3] >>> thresh = 0.0 >>> non_max_supression(ltrb, scores, thresh, impl='numpy') >>> if 'numpy' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='numpy') >>> assert list(keep) == [2, 1] >>> if 'cython_cpu' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='cython_cpu') >>> assert list(keep) == [2, 1] >>> if 'cython_gpu' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='cython_gpu') >>> assert list(keep) == [2, 1] >>> if 'torch' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='torch') >>> assert set(keep.tolist()) == {2, 1} >>> if 'torchvision' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='torchvision') # note torchvision has no bias >>> assert list(keep) == [2] >>> thresh = 1.0 >>> if 'numpy' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='numpy') >>> assert list(keep) == [2, 1, 3, 0] >>> if 'cython_cpu' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='cython_cpu') >>> assert list(keep) == [2, 1, 3, 0] >>> if 'cython_gpu' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='cython_gpu') >>> assert list(keep) == [2, 1, 3, 0] >>> if 'torch' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='torch') >>> assert set(keep.tolist()) == {2, 1, 3, 0} >>> if 'torchvision' in available_nms_impls(): >>> keep = non_max_supression(ltrb, scores, thresh, impl='torchvision') # note torchvision has no bias >>> assert set(kwarray.ArrayAPI.tolist(keep)) == {2, 1, 3, 0}
Example
>>> import ubelt as ub >>> ltrb = np.array([ >>> [0, 0, 100, 100], >>> [100, 100, 10, 10], >>> [10, 10, 100, 100], >>> [50, 50, 100, 100], >>> [100, 100, 150, 101], >>> [120, 100, 180, 101], >>> [150, 100, 200, 101], >>> ], dtype=np.float32) >>> scores = np.linspace(0, 1, len(ltrb)) >>> thresh = .2 >>> solutions = {} >>> if not _impls._funcs: >>> _impls._lazy_init() >>> for impl in _impls._funcs: >>> keep = non_max_supression(ltrb, scores, thresh, impl=impl) >>> solutions[impl] = sorted(keep) >>> assert 'numpy' in solutions >>> print('solutions = {}'.format(ub.urepr(solutions, nl=1))) >>> assert ub.allsame(solutions.values())
CommandLine
xdoctest -m ~/code/kwimage/kwimage/algo/algo_nms.py non_max_supression
Example
>>> import ubelt as ub >>> # Check that zero-area boxes are ok >>> ltrb = np.array([ >>> [0, 0, 0, 0], >>> [0, 0, 0, 0], >>> [10, 10, 10, 10], >>> ], dtype=np.float32) >>> scores = np.array([1, 2, 3], dtype=np.float32) >>> thresh = .2 >>> solutions = {} >>> if not _impls._funcs: >>> _impls._lazy_init() >>> for impl in _impls._funcs: >>> keep = non_max_supression(ltrb, scores, thresh, impl=impl) >>> solutions[impl] = sorted(keep) >>> assert 'numpy' in solutions >>> print('solutions = {}'.format(ub.urepr(solutions, nl=1))) >>> assert ub.allsame(solutions.values())
- kwimage.normalize(arr, mode='linear', alpha=None, beta=None, out=None)[source]¶
Rebalance pixel intensities via contrast stretching.
By default linearly stretches pixel intensities to minimum and maximum values.
Note
DEPRECATED: this function has been MOVED to
kwarray.normalize
- kwimage.normalize_intensity(imdata, return_info=False, nodata=None, axis=None, dtype=<class 'numpy.float32'>, params='auto', mask=None)[source]¶
Normalize data intensities using heuristics to help put sensor data with extremely high or low contrast into a visible range.
This function is designed with an emphasis on getting something that is reasonable for visualization.
Todo
[x] Move to kwarray and renamed to robust_normalize?
[ ] Support for M-estimators?
- Parameters:
imdata (ndarray) – raw intensity data
return_info (bool) – if True, return information about the chosen normalization heuristic.
params (str | dict) – can contain keys, low, high, or center e.g. {‘low’: 0.1, ‘center’: 0.8, ‘high’: 0.9}
axis (None | int) – The axis to normalize over, if unspecified, normalize jointly
nodata (None | int) – A value representing nodata to leave unchanged during normalization, for example 0
dtype (type) – can be float32 or float64
mask (ndarray | None) – A mask indicating what pixels are valid and what pixels should be considered nodata. Mutually exclusive with
nodataargument. A mask value of 1 indicates a VALID pixel. A mask value of 0 indicates an INVALID pixel.
- Returns:
a floating point array with values between 0 and 1.
- Return type:
ndarray
Example
>>> from kwimage.im_core import * # NOQA >>> import ubelt as ub >>> import kwimage >>> import kwarray >>> s = 512 >>> bit_depth = 11 >>> dtype = np.uint16 >>> max_val = int(2 ** bit_depth) >>> min_val = int(0) >>> rng = kwarray.ensure_rng(0) >>> background = np.random.randint(min_val, max_val, size=(s, s), dtype=dtype) >>> poly1 = kwimage.Polygon.random(rng=rng).scale(s / 2) >>> poly2 = kwimage.Polygon.random(rng=rng).scale(s / 2).translate(s / 2) >>> forground = np.zeros_like(background, dtype=np.uint8) >>> forground = poly1.fill(forground, value=255) >>> forground = poly2.fill(forground, value=122) >>> forground = (kwimage.ensure_float01(forground) * max_val).astype(dtype) >>> imdata = background + forground >>> normed, info = normalize_intensity(imdata, return_info=True) >>> print('info = {}'.format(ub.urepr(info, nl=1))) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(imdata, pnum=(1, 2, 1), fnum=1) >>> kwplot.imshow(normed, pnum=(1, 2, 2), fnum=1)

Example
>>> from kwimage.im_core import * # NOQA >>> import ubelt as ub >>> import kwimage >>> # Test on an image that is already normalized to test how it >>> # degrades >>> imdata = kwimage.grab_test_image() / 255
>>> quantile_basis = { >>> 'mode': ['linear', 'sigmoid'], >>> 'high': [0.8, 0.9, 1.0], >>> } >>> quantile_grid = list(ub.named_product(quantile_basis)) >>> quantile_grid += ['auto'] >>> rows = [] >>> rows.append({'key': 'orig', 'result': imdata}) >>> for params in quantile_grid: >>> key = ub.urepr(params, compact=1) >>> result, info = normalize_intensity(imdata, return_info=True, params=params) >>> print('key = {}'.format(key)) >>> print('info = {}'.format(ub.urepr(info, nl=1))) >>> rows.append({'key': key, 'info': info, 'result': result}) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nSubplots=len(rows)) >>> for row in rows: >>> _, ax = kwplot.imshow(row['result'], fnum=1, pnum=pnum_()) >>> ax.set_title(row['key'])

- kwimage.num_channels(img)[source]¶
Returns the number of color channels in an image.
Assumes images are 2D and the the channels are the trailing dimension. Returns 1 in the case with no trailing channel dimension, otherwise simply returns
img.shape[2].- Parameters:
img (ndarray) – an image with 2 or 3 dimensions.
- Returns:
the number of color channels (1, 3, or 4)
- Return type:
Example
>>> H = W = 3 >>> assert num_channels(np.empty((W, H))) == 1 >>> assert num_channels(np.empty((W, H, 1))) == 1 >>> assert num_channels(np.empty((W, H, 3))) == 3 >>> assert num_channels(np.empty((W, H, 4))) == 4 >>> assert num_channels(np.empty((W, H, 2))) == 2
- kwimage.overlay_alpha_images(img1, img2, keepalpha=True, dtype=<class 'numpy.float32'>, impl='inplace')[source]¶
Places img1 on top of img2 respecting alpha channels. Works like the Photoshop layers with opacity.
- Parameters:
img1 (ndarray) – top image to overlay over img2
img2 (ndarray) – base image to superimpose on
keepalpha (bool) – if False, the alpha channel is removed after blending
dtype (np.dtype) – format for blending computation (defaults to float32)
impl (str) – code specifying the backend implementation
- Returns:
raster: the blended images
- Return type:
ndarray
Todo
[ ] Make fast C++ version of this function
Example
>>> import kwimage >>> img1 = kwimage.grab_test_image('astro', dsize=(100, 100)) >>> img2 = kwimage.grab_test_image('carl', dsize=(100, 100)) >>> img1 = kwimage.ensure_alpha_channel(img1, alpha=.5) >>> img3 = kwimage.overlay_alpha_images(img1, img2) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(img3) >>> kwplot.show_if_requested()

- kwimage.overlay_alpha_layers(layers, keepalpha=True, dtype=<class 'numpy.float32'>)[source]¶
Stacks a sequences of layers on top of one another. The first item is the topmost layer and the last item is the bottommost layer.
- Parameters:
layers (Sequence[ndarray]) – stack of images
keepalpha (bool) – if False, the alpha channel is removed after blending
dtype (np.dtype) – format for blending computation (defaults to float32)
- Returns:
raster: the blended images
- Return type:
ndarray
Example
>>> import kwimage >>> keys = ['astro', 'carl', 'stars'] >>> layers = [kwimage.grab_test_image(k, dsize=(100, 100)) for k in keys] >>> layers = [kwimage.ensure_alpha_channel(g, alpha=.5) for g in layers] >>> stacked = kwimage.overlay_alpha_layers(layers) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(stacked) >>> kwplot.show_if_requested()

- kwimage.padded_slice(data, in_slice, pad=None, padkw=None, return_info=False)[source]¶
Allows slices with out-of-bound coordinates. Any out of bounds coordinate will be sampled via padding.
DEPRECATED FOR THE VERSION IN KWARRAY (slices are more array-ish than image-ish)
Note
Negative slices have a different meaning here then they usually do. Normally, they indicate a wrap-around or a reversed stride, but here they index into out-of-bounds space (which depends on the pad mode). For example a slice of -2:1 literally samples two pixels to the left of the data and one pixel from the data, so you get two padded values and one data value.
- Parameters:
data (Sliceable) – data to slice into. Any channels must be the last dimension.
in_slice (slice | Tuple[slice, …]) – slice for each dimensions
ndim (int) – number of spatial dimensions
pad (List[int|Tuple]) – additional padding of the slice
padkw (Dict) – if unspecified defaults to
{'mode': 'constant'}return_info (bool) – if True, return extra information about the transform. Defaults to False.
- SeeAlso:
_padded_slice_embed - finds the embedded slice and padding _padded_slice_apply - applies padding to sliced data
- Returns:
- data_sliced: subregion of the input data (possibly with padding,
depending on if the original slice went out of bounds)
- Tuple[Sliceable, Dict] :
data_sliced : as above
transform : information on how to return to the original coordinates
- Currently a dict containing:
- st_dims: a list indicating the low and high space-time
coordinate values of the returned data slice.
The structure of this dictionary mach change in the future
- Return type:
Sliceable
Example
>>> data = np.arange(5) >>> in_slice = [slice(-2, 7)]
>>> data_sliced = padded_slice(data, in_slice) >>> print(ub.urepr(data_sliced, with_dtype=False)) np.array([0, 0, 0, 1, 2, 3, 4, 0, 0])
>>> data_sliced = padded_slice(data, in_slice, pad=(3, 3)) >>> print(ub.urepr(data_sliced, with_dtype=False)) np.array([0, 0, 0, 0, 0, 0, 1, 2, 3, 4, 0, 0, 0, 0, 0])
>>> data_sliced = padded_slice(data, slice(3, 4), pad=[(1, 0)]) >>> print(ub.urepr(data_sliced, with_dtype=False)) np.array([2, 3])
- kwimage.radial_fourier_mask(img_hwc, radius=11, axis=None, clip=None)[source]¶
In [1] they use a radius of 11.0 on CIFAR-10.
- Parameters:
img_hwc (ndarray) – assumed to be float 01
References
[1] Jo and Bengio “Measuring the tendency of CNNs to Learn Surface Statistical Regularities” 2017. https://docs.opencv.org/3.0-beta/doc/py_tutorials/py_imgproc/py_transforms/py_fourier_transform/py_fourier_transform.html
Example
>>> from kwimage.im_filter import * # NOQA >>> import kwimage >>> img_hwc = kwimage.grab_test_image() >>> img_hwc = kwimage.ensure_float01(img_hwc) >>> out_hwc = radial_fourier_mask(img_hwc, radius=11) >>> # xdoctest: REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> def keepdim(func): >>> def _wrap(im): >>> needs_transpose = (im.shape[0] == 3) >>> if needs_transpose: >>> im = im.transpose(1, 2, 0) >>> out = func(im) >>> if needs_transpose: >>> out = out.transpose(2, 0, 1) >>> return out >>> return _wrap >>> @keepdim >>> def rgb_to_lab(im): >>> return kwimage.convert_colorspace(im, src_space='rgb', dst_space='lab') >>> @keepdim >>> def lab_to_rgb(im): >>> return kwimage.convert_colorspace(im, src_space='lab', dst_space='rgb') >>> @keepdim >>> def rgb_to_yuv(im): >>> return kwimage.convert_colorspace(im, src_space='rgb', dst_space='yuv') >>> @keepdim >>> def yuv_to_rgb(im): >>> return kwimage.convert_colorspace(im, src_space='yuv', dst_space='rgb') >>> def show_data(img_hwc): >>> # dpath = ub.ensuredir('./fouriertest') >>> kwplot.imshow(img_hwc, fnum=1) >>> pnum_ = kwplot.PlotNums(nRows=4, nCols=5) >>> for r in range(0, 17): >>> imgt = radial_fourier_mask(img_hwc, r, clip=(0, 1)) >>> kwplot.imshow(imgt, pnum=pnum_(), fnum=2) >>> plt.gca().set_title('r = {}'.format(r)) >>> kwplot.set_figtitle('RGB') >>> # plt.gcf().savefig(join(dpath, '{}_{:08d}.png'.format('rgb', x))) >>> pnum_ = kwplot.PlotNums(nRows=4, nCols=5) >>> for r in range(0, 17): >>> imgt = lab_to_rgb(radial_fourier_mask(rgb_to_lab(img_hwc), r)) >>> kwplot.imshow(imgt, pnum=pnum_(), fnum=3) >>> plt.gca().set_title('r = {}'.format(r)) >>> kwplot.set_figtitle('LAB') >>> # plt.gcf().savefig(join(dpath, '{}_{:08d}.png'.format('lab', x))) >>> pnum_ = kwplot.PlotNums(nRows=4, nCols=5) >>> for r in range(0, 17): >>> imgt = yuv_to_rgb(radial_fourier_mask(rgb_to_yuv(img_hwc), r)) >>> kwplot.imshow(imgt, pnum=pnum_(), fnum=4) >>> plt.gca().set_title('r = {}'.format(r)) >>> kwplot.set_figtitle('YUV') >>> # plt.gcf().savefig(join(dpath, '{}_{:08d}.png'.format('yuv', x))) >>> show_data(img_hwc) >>> kwplot.show_if_requested()




- kwimage.remove_homog(pts, mode='divide')[source]¶
Remove homogenous coordinate to a point array.
This is a convinience function, it is not particularly efficient.
- SeeAlso:
cv2.convertPointsFromHomogeneous
Example
>>> homog_pts = np.random.rand(10, 3) >>> remove_homog(homog_pts, 'divide') >>> remove_homog(homog_pts, 'drop')
- kwimage.rle_translate(rle, offset, output_shape=None)[source]¶
Translates a run-length encoded image in RLE-space.
- Parameters:
rle (dict) – an enconding dict returned by
kwimage.encode_run_length()offset (Tuple[int, int]) – x, y integer offsets.
output_shape (Tuple[int, int]) – h,w of transformed mask. If unspecified the input rle shape is used.
- SeeAlso:
# ITK has some RLE code that looks like it can perform translations https://github.com/KitwareMedical/ITKRLEImage/blob/master/include/itkRLERegionOfInterestImageFilter.h
Doctest
>>> # test that translate works on all zero images >>> img = np.zeros((7, 8), dtype=np.uint8) >>> rle = encode_run_length(img, binary=True, order='F') >>> new_rle = rle_translate(rle, (1, 2), (6, 9)) >>> assert np.all(new_rle['counts'] == [54])
Example
>>> from kwimage.im_runlen import * # NOQA >>> img = np.array([ >>> [1, 1, 1, 1], >>> [0, 1, 0, 0], >>> [0, 1, 0, 1], >>> [1, 1, 1, 1],], dtype=np.uint8) >>> rle = encode_run_length(img, binary=True, order='C') >>> offset = (1, -1) >>> output_shape = (3, 5) >>> new_rle = rle_translate(rle, offset, output_shape) >>> decoded = decode_run_length(**new_rle) >>> print(decoded) [[0 0 1 0 0] [0 0 1 0 1] [0 1 1 1 1]]
Example
>>> from kwimage.im_runlen import * # NOQA >>> img = np.array([ >>> [0, 0, 0], >>> [0, 1, 0], >>> [0, 0, 0]], dtype=np.uint8) >>> rle = encode_run_length(img, binary=True, order='C') >>> new_rle = rle_translate(rle, (1, 0)) >>> decoded = decode_run_length(**new_rle) >>> print(decoded) [[0 0 0] [0 0 1] [0 0 0]] >>> new_rle = rle_translate(rle, (0, 1)) >>> decoded = decode_run_length(**new_rle) >>> print(decoded) [[0 0 0] [0 0 0] [0 1 0]]
- kwimage.smooth_prob(prob, k=3, inplace=False, eps=1e-09)[source]¶
Smooths the probability map, but preserves the magnitude of the peaks.
Note
even if inplace is true, we still need to make a copy of the input array, however, we do ensure that it is cleaned up before we leave the function scope.
sigma=0.8 @ k=3, sigma=1.1 @ k=5, sigma=1.4 @ k=7
- kwimage.stack_images(images, axis=0, resize=None, interpolation=None, overlap=0, return_info=False, bg_value=None, pad=None, allow_casting=True)[source]¶
Make a new image with the input images side-by-side
- Parameters:
images (Iterable[ndarray]) – image data
axis (int) – axis to stack on (either 0 or 1)
resize (int | str | None) – if None image sizes are not modified, otherwise resize resize can be either 0 or 1. We resize the resize-th image to match the 1 - resize-th image. In other words, resize=0 means the current image is resized to match the next image, and resize=1 means the next image is resized to match the current image. Can also be strings “larger” or “smaller”. If resize=”larger”, the current image is only resized if the next image is larger. If resize=’smaller’ the current image is resized only if the next image is smaller. TODO: this needs to be reworked to be more intuitive.
interpolation (int | str) – string or cv2-style interpolation type. only used if resize or overlap > 0
overlap (int) – number of pixels to overlap. Using a negative number results in a border.
pad (int) – if specified overrides overlap as a the number of pixels to pad between images.
return_info (bool) – if True, returns transforms (scales and translations) to map from original image to its new location.
bg_value (Number | ndarray | str) – background value or color, if specified, uses this as a fill value.
allow_casting (bool) – if True, then if “uint255” and “float01” format images are given they are converted to “float01”. Defaults to True.
- Returns:
an image of stacked images side by side
- Tuple[ndarray, List]: where the first item is the aformentioned stacked
image and the second item is a list of transformations for each input image mapping it to its location in the returned image.
- Return type:
ndarray
Todo
- [ ] This is currently implemented by calling the “stack_two_images”
function multiple times. This should be optimized by allocating the entire canvas first.
Example
>>> import kwimage >>> img1 = kwimage.grab_test_image('carl', space='rgb') >>> img2 = kwimage.grab_test_image('astro', space='rgb') >>> images = [img1, img2] >>> imgB, transforms = kwimage.stack_images( >>> images, axis=0, resize='larger', pad=10, return_info=True) >>> print('imgB.shape = {}'.format(imgB.shape)) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> import kwimage >>> kwplot.autompl() >>> kwplot.imshow(imgB, colorspace='rgb') >>> wh1 = np.multiply(img1.shape[0:2][::-1], transforms[0].scale) >>> wh2 = np.multiply(img2.shape[0:2][::-1], transforms[1].scale) >>> xoff1, yoff1 = transforms[0].translation >>> xoff2, yoff2 = transforms[1].translation >>> xywh1 = (xoff1, yoff1, wh1[0], wh1[1]) >>> xywh2 = (xoff2, yoff2, wh2[0], wh2[1]) >>> kwplot.draw_boxes(kwimage.Boxes([xywh1], 'xywh'), color=(1.0, 0, 0)) >>> kwplot.draw_boxes(kwimage.Boxes([xywh2], 'xywh'), color=(1.0, 0, 0)) >>> kwplot.show_if_requested()

- kwimage.stack_images_grid(images, chunksize=None, axis=0, overlap=0, pad=None, return_info=False, bg_value=None, resize=None, allow_casting=True)[source]¶
Stacks images in a grid. Optionally return transforms of original image positions in the output image.
- Parameters:
images (Iterable[ndarray]) – image data
chunksize (int) – number of rows per column or columns per row depending on the value of axis. If unspecified, computes this as int(sqrt(len(images))).
axis (int) – If 0, chunksize is columns per row. If 1, chunksize is rows per column. Defaults to 0.
overlap (int) – number of pixels to overlap. Using a negative number results in a border.
pad (int) – if specified overrides overlap as a the number of pixels to pad between images.
return_info (bool) – if True, returns transforms (scales and translations) to map from original image to its new location.
resize (int | str | None) – if None image sizes are not modified, otherwise can be set to “larger” or “smaller” to resize the images in each stack direction.
bg_value (Number | ndarray | str) – background value or color, if specified, uses this as a fill value.
allow_casting (bool) – if True, then if “uint255” and “float01” format images are given they are converted to “float01”. Defaults to True.
- Returns:
an image of stacked images in a grid pattern
- Tuple[ndarray, List]: where the first item is the aformentioned stacked
image and the second item is a list of transformations for each input image mapping it to its location in the returned image.
- Return type:
ndarray
- SeeAlso:
Example
>>> import kwimage >>> img1 = kwimage.grab_test_image('carl') >>> img2 = kwimage.grab_test_image('astro') >>> img3 = kwimage.grab_test_image('airport') >>> img4 = kwimage.grab_test_image('paraview')[..., 0:3] >>> img5 = kwimage.grab_test_image('pm5644') >>> images = [img1, img2, img3, img4, img5] >>> canvas, transforms = kwimage.stack_images_grid( ... images, chunksize=3, axis=0, pad=10, bg_value='kitware_blue', ... return_info=True, resize='larger') >>> print('canvas.shape = {}'.format(canvas.shape)) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> import kwimage >>> kwplot.autompl() >>> kwplot.imshow(canvas) >>> kwplot.show_if_requested()
- kwimage.subpixel_accum(dst, src, index, interp_axes=None)[source]¶
Add the source values array into the destination array at a particular subpixel index.
- Parameters:
dst (ArrayLike) – destination accumulation array
src (ArrayLike) – source array containing values to add
index (Tuple[slice]) – subpixel slice into dst that corresponds with src
interp_axes (tuple) – specify which axes should be spatially interpolated
TextArt
Inputs: +---+---+---+---+---+ dst.shape = (5,) +---+---+ src.shape = (2,) |=======| index = 1.5:3.5 Subpixel shift the source by -0.5. When the index is non-integral, pad the aligned src with an extra value to ensure all dst pixels that would be influenced by the smaller subpixel shape are influenced by the aligned src. Note that we are not scaling. +---+---+---+ aligned_src.shape = (3,) |===========| aligned_index = 1:4
Example
>>> dst = np.zeros(5) >>> src = np.ones(2) >>> index = [slice(1.5, 3.5)] >>> subpixel_accum(dst, src, index) >>> print(ub.urepr(dst, precision=2, with_dtype=0)) np.array([0. , 0.5, 1. , 0.5, 0. ])
Example
>>> dst = np.zeros((6, 6)) >>> src = np.ones((3, 3)) >>> index = (slice(1.5, 4.5), slice(1, 4)) >>> subpixel_accum(dst, src, index) >>> print(ub.urepr(dst, precision=2, with_dtype=0)) np.array([[0. , 0. , 0. , 0. , 0. , 0. ], [0. , 0.5, 0.5, 0.5, 0. , 0. ], [0. , 1. , 1. , 1. , 0. , 0. ], [0. , 1. , 1. , 1. , 0. , 0. ], [0. , 0.5, 0.5, 0.5, 0. , 0. ], [0. , 0. , 0. , 0. , 0. , 0. ]]) >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> dst = torch.zeros((1, 3, 6, 6)) >>> src = torch.ones((1, 3, 3, 3)) >>> index = (slice(None), slice(None), slice(1.5, 4.5), slice(1.25, 4.25)) >>> subpixel_accum(dst, src, index) >>> print(ub.urepr(dst.numpy()[0, 0], precision=2, with_dtype=0)) np.array([[0. , 0. , 0. , 0. , 0. , 0. ], [0. , 0.38, 0.5 , 0.5 , 0.12, 0. ], [0. , 0.75, 1. , 1. , 0.25, 0. ], [0. , 0.75, 1. , 1. , 0.25, 0. ], [0. , 0.38, 0.5 , 0.5 , 0.12, 0. ], [0. , 0. , 0. , 0. , 0. , 0. ]])
Doctest
>>> # TODO: move to a unit test file >>> subpixel_accum(np.zeros(5), np.ones(2), [slice(1.5, 3.5)]).tolist() [0.0, 0.5, 1.0, 0.5, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(2), [slice(0, 2)]).tolist() [1.0, 1.0, 0.0, 0.0, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(.5, 3.5)]).tolist() [0.5, 1.0, 1.0, 0.5, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(-1, 2)]).tolist() [1.0, 1.0, 0.0, 0.0, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(-1.5, 1.5)]).tolist() [1.0, 0.5, 0.0, 0.0, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(10, 13)]).tolist() [0.0, 0.0, 0.0, 0.0, 0.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(3.25, 6.25)]).tolist() [0.0, 0.0, 0.0, 0.75, 1.0] >>> subpixel_accum(np.zeros(5), np.ones(3), [slice(4.9, 7.9)]).tolist() [0.0, 0.0, 0.0, 0.0, 0.099...] >>> subpixel_accum(np.zeros(5), np.ones(9), [slice(-1.5, 7.5)]).tolist() [1.0, 1.0, 1.0, 1.0, 1.0] >>> subpixel_accum(np.zeros(5), np.ones(9), [slice(2.625, 11.625)]).tolist() [0.0, 0.0, 0.375, 1.0, 1.0] >>> subpixel_accum(np.zeros(5), 1, [slice(2.625, 11.625)]).tolist() [0.0, 0.0, 0.375, 1.0, 1.0]
- kwimage.subpixel_align(dst, src, index, interp_axes=None)[source]¶
Returns an aligned version of the source tensor and destination index.
- Used as the backend to implement other subpixel functions like:
subpixel_accum, subpixel_maximum.
- kwimage.subpixel_getvalue(img, pts, coord_axes=None, interp='bilinear', bordermode='edge')[source]¶
Get values at subpixel locations
- Parameters:
img (ArrayLike) – image to sample from
pts (ArrayLike) – subpixel rc-coordinates to sample
coord_axes (Sequence) – axes to perform interpolation on, if not specified the first d axes are interpolated, where d=pts.shape[-1]. IE: this indicates which axes each coordinate dimension corresponds to.
interp (str) – interpolation mode
bordermode (str) – how locations outside the image are handled
Example
>>> from kwimage.util_warp import * # NOQA >>> img = np.arange(3 * 3).reshape(3, 3) >>> pts = np.array([[1, 1], [1.5, 1.5], [1.9, 1.1]]) >>> subpixel_getvalue(img, pts) array([4. , 6. , 6.8]) >>> subpixel_getvalue(img, pts, coord_axes=(1, 0)) array([4. , 6. , 5.2]) >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> img = torch.Tensor(img) >>> pts = torch.Tensor(pts) >>> subpixel_getvalue(img, pts) tensor([4.0000, 6.0000, 6.8000]) >>> subpixel_getvalue(img.numpy(), pts.numpy(), interp='nearest') array([4., 8., 7.], dtype=float32) >>> subpixel_getvalue(img.numpy(), pts.numpy(), interp='nearest', coord_axes=[1, 0]) array([4., 8., 5.], dtype=float32) >>> subpixel_getvalue(img, pts, interp='nearest') tensor([4., 8., 7.])
References
[SO12729228]stackoverflow.com/questions/12729228/simple-binlin-interp-images-numpy
- SeeAlso:
cv2.getRectSubPix(image, patchSize, center[, patch[, patchType]])
- kwimage.subpixel_maximum(dst, src, index, interp_axes=None)[source]¶
Take the max of the source values array into and the destination array at a particular subpixel index. Modifies the destination array.
- Parameters:
dst (ArrayLike) – destination array to index into
src (ArrayLike) – source array that agrees with the index
index (Tuple[slice]) – subpixel slice into dst that corresponds with src
interp_axes (tuple) – specify which axes should be spatially interpolated
Example
>>> dst = np.array([0, 1.0, 1.0, 1.0, 0]) >>> src = np.array([2.0, 2.0]) >>> index = [slice(1.6, 3.6)] >>> subpixel_maximum(dst, src, index) >>> print(ub.urepr(dst, precision=2, with_dtype=0)) np.array([0. , 1. , 2. , 1.2, 0. ])
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> dst = torch.zeros((1, 3, 5, 5)) + .5 >>> src = torch.ones((1, 3, 3, 3)) >>> index = (slice(None), slice(None), slice(1.4, 4.4), slice(1.25, 4.25)) >>> subpixel_maximum(dst, src, index) >>> print(ub.urepr(dst.numpy()[0, 0], precision=2, with_dtype=0)) np.array([[0.5 , 0.5 , 0.5 , 0.5 , 0.5 ], [0.5 , 0.5 , 0.6 , 0.6 , 0.5 ], [0.5 , 0.75, 1. , 1. , 0.5 ], [0.5 , 0.75, 1. , 1. , 0.5 ], [0.5 , 0.5 , 0.5 , 0.5 , 0.5 ]])
- kwimage.subpixel_minimum(dst, src, index, interp_axes=None)[source]¶
Take the min of the source values array into and the destination array at a particular subpixel index. Modifies the destination array.
- Parameters:
dst (ArrayLike) – destination array to index into
src (ArrayLike) – source array that agrees with the index
index (Tuple[slice]) – subpixel slice into dst that corresponds with src
interp_axes (tuple) – specify which axes should be spatially interpolated
Example
>>> dst = np.array([0, 1.0, 1.0, 1.0, 0]) >>> src = np.array([2.0, 2.0]) >>> index = [slice(1.6, 3.6)] >>> subpixel_minimum(dst, src, index) >>> print(ub.urepr(dst, precision=2, with_dtype=0)) np.array([0. , 0.8, 1. , 1. , 0. ])
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> dst = torch.zeros((1, 3, 5, 5)) + .5 >>> src = torch.ones((1, 3, 3, 3)) >>> index = (slice(None), slice(None), slice(1.4, 4.4), slice(1.25, 4.25)) >>> subpixel_minimum(dst, src, index) >>> print(ub.urepr(dst.numpy()[0, 0], precision=2, with_dtype=0)) np.array([[0.5 , 0.5 , 0.5 , 0.5 , 0.5 ], [0.5 , 0.45, 0.5 , 0.5 , 0.15], [0.5 , 0.5 , 0.5 , 0.5 , 0.25], [0.5 , 0.5 , 0.5 , 0.5 , 0.25], [0.5 , 0.3 , 0.4 , 0.4 , 0.1 ]])
- kwimage.subpixel_set(dst, src, index, interp_axes=None)[source]¶
Add the source values array into the destination array at a particular subpixel index.
- Parameters:
dst (ArrayLike) – destination accumulation array
src (ArrayLike) – source array containing values to add
index (Tuple[slice]) – subpixel slice into dst that corresponds with src
interp_axes (tuple) – specify which axes should be spatially interpolated
Todo
[ ]: allow index to be a sequence indices
Example
>>> import kwimage >>> dst = np.zeros(5) + .1 >>> src = np.ones(2) >>> index = [slice(1.5, 3.5)] >>> kwimage.util_warp.subpixel_set(dst, src, index) >>> print(ub.urepr(dst, precision=2, with_dtype=0)) np.array([0.1, 0.5, 1. , 0.5, 0.1])
- kwimage.subpixel_setvalue(img, pts, value, coord_axes=None, interp='bilinear', bordermode='edge')[source]¶
Set values at subpixel locations
- Parameters:
img (ArrayLike) – image to set values in
pts (ArrayLike) – subpixel rc-coordinates to set
value (ArrayLike) – value to place in the image
coord_axes (Sequence) – axes to perform interpolation on, if not specified the first d axes are interpolated, where d=pts.shape[-1]. IE: this indicates which axes each coordinate dimension corresponds to.
interp (str) – interpolation mode
bordermode (str) – how locations outside the image are handled
Example
>>> from kwimage.util_warp import * # NOQA >>> img = np.arange(3 * 3).reshape(3, 3).astype(float) >>> pts = np.array([[1, 1], [1.5, 1.5], [1.9, 1.1]]) >>> interp = 'bilinear' >>> value = 0 >>> print('img = {!r}'.format(img)) >>> pts = np.array([[1.5, 1.5]]) >>> img2 = subpixel_setvalue(img.copy(), pts, value) >>> print('img2 = {!r}'.format(img2)) >>> pts = np.array([[1.0, 1.0]]) >>> img2 = subpixel_setvalue(img.copy(), pts, value) >>> print('img2 = {!r}'.format(img2)) >>> pts = np.array([[1.1, 1.9]]) >>> img2 = subpixel_setvalue(img.copy(), pts, value) >>> print('img2 = {!r}'.format(img2)) >>> img2 = subpixel_setvalue(img.copy(), pts, value, coord_axes=[1, 0]) >>> print('img2 = {!r}'.format(img2))
- kwimage.subpixel_slice(inputs, index)[source]¶
Take a subpixel slice from a larger image. The returned output is left-aligned with the requested slice.
- Parameters:
inputs (ArrayLike) – data
index (Tuple[slice]) – a slice to subpixel accuracy
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> import torch >>> # say we have a (576, 576) input space >>> # and a (9, 9) output space downsampled by 64x >>> ospc_feats = np.tile(np.arange(9 * 9).reshape(1, 9, 9), (1024, 1, 1)) >>> inputs = torch.from_numpy(ospc_feats) >>> # We detected a box in the input space >>> ispc_bbox = kwimage.Boxes([[64, 65, 100, 120]], 'ltrb') >>> # Get coordinates in the output space >>> ospc_bbox = ispc_bbox.scale(1 / 64) >>> tl_x, tl_y, br_x, br_y = ospc_bbox.data[0] >>> # Convert the box to a slice >>> index = [slice(None), slice(tl_y, br_y), slice(tl_x, br_x)] >>> # Note: I'm not 100% sure this work right with non-intergral slices >>> outputs = kwimage.subpixel_slice(inputs, index)
Example
>>> inputs = np.arange(5 * 5 * 3).reshape(5, 5, 3) >>> index = [slice(0, 3), slice(0, 3)] >>> outputs = subpixel_slice(inputs, index) >>> index = [slice(0.5, 3.5), slice(-0.5, 2.5)] >>> outputs = subpixel_slice(inputs, index)
>>> inputs = np.arange(5 * 5).reshape(1, 5, 5).astype(float) >>> index = [slice(None), slice(3, 6), slice(3, 6)] >>> outputs = subpixel_slice(inputs, index) >>> print(outputs) [[[18. 19. 0.] [23. 24. 0.] [ 0. 0. 0.]]] >>> index = [slice(None), slice(3.5, 6.5), slice(2.5, 5.5)] >>> outputs = subpixel_slice(inputs, index) >>> print(outputs) [[[20. 21. 10.75] [11.25 11.75 6. ] [ 0. 0. 0. ]]]
- kwimage.subpixel_translate(inputs, shift, interp_axes=None, output_shape=None)[source]¶
Translates an image by a subpixel shift value using bilinear interpolation
- Parameters:
inputs (ArrayLike) – data to translate
shift (Sequence) – amount to translate each dimension specified by interp_axes. Note: if inputs contains more than one “image” then all “images” are translated by the same amount. This function contains no mechanism for translating each image differently. Note that by default this is a y,x shift for 2 dimensions.
interp_axes (Sequence) – axes to perform interpolation on, if not specified the final n axes are interpolated, where n=len(shift)
output_shape (tuple) – if specified the output is returned with this shape, otherwise
Note
This function powers most other functions in this file. Speedups here can go a long way.
Example
>>> inputs = np.arange(5) + 1 >>> print(inputs.tolist()) [1, 2, 3, 4, 5] >>> outputs = subpixel_translate(inputs, 1.5) >>> print(outputs.tolist()) [0.0, 0.5, 1.5, 2.5, 3.5]
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> inputs = torch.arange(9).view(1, 1, 3, 3).float() >>> print(inputs.long()) tensor([[[[0, 1, 2], [3, 4, 5], [6, 7, 8]]]]) >>> outputs = subpixel_translate(inputs, (-.4, .5), output_shape=(1, 1, 2, 5)) >>> print(outputs) tensor([[[[0.6000, 1.7000, 2.7000, 1.6000, 0.0000], [2.1000, 4.7000, 5.7000, 3.1000, 0.0000]]]])
- kwimage.warp_affine(image, transform, dsize=None, antialias=False, interpolation='linear', border_mode=None, border_value=0, large_warp_dim=None, return_info=False)[source]¶
Applies an affine transformation to an image with optional antialiasing.
- Parameters:
image (ndarray) – the input image as a numpy array. Note: this is passed directly to cv2, so it is best to ensure that it is contiguous and using a dtype that cv2 can handle.
transform (ndarray | dict | kwimage.Affine) – a coercable affine matrix. See
kwimage.Affinefor details on what can be coerced.dsize (Tuple[int, int] | None | str) – A integer width and height tuple of the resulting “canvas” image. If None, then the input image size is used.
If specified as a string, dsize is computed based on the given heuristic.
If ‘positive’ (or ‘auto’), dsize is computed such that the positive coordinates of the warped image will fit in the new canvas. In this case, any pixel that maps to a negative coordinate will be clipped. This has the property that the input transformation is not modified. NOTE: there are issues with this when the transformation includes rotation of reflections.
If ‘content’ (or ‘max’), the transform is modified with an extra translation such that both the positive and negative coordinates of the warped image will fit in the new canvas.
antialias (bool) – if True determines if the transform is downsampling and applies antialiasing via gaussian a blur. Defaults to False
interpolation (str | int) – interpolation code or cv2 integer. Interpolation codes are linear, nearest, cubic, lancsoz, and area. Defaults to “linear”.
border_mode (str | int) – Border code or cv2 integer. Border codes are constant (default) replicate, reflect, wrap, reflect101, and transparent.
border_value (int | float | Iterable[int | float]) – Used as the fill value if border_mode is constant. Otherwise this is ignored. Defaults to 0, but can also be defaulted to nan. if border_value is a scalar and there are multiple channels, the value is applied to all channels. More than 4 unique border values for individual channels will cause an error. See OpenCV #22283 for details. In the future we may accept np.ma and return a masked array, but for now that is not implemented.
large_warp_dim (int | None | str) – If specified, perform the warp piecewise in chunks of the specified size. If “auto”, it is set to the maximum “short” value in numpy. This works around a limitation of cv2.warpAffine, which must have image dimensions < SHRT_MAX (=32767 in version 4.5.3)
return_info (bool) – if True, returns information about the operation. In the case where dsize=”content”, this includes the modified transformation.
- Returns:
the warped image, or if return info is True, the warped image and the info dictionary.
- Return type:
ndarray | Tuple[ndarray, Dict]
Todo
- [ ] When dsize=’positive’ but the transform contains an axis flip,
the width / height of the box will become negative. Should we adjust for this?
Example
>>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> from kwimage.transform import Affine >>> image = kwimage.grab_test_image('astro') >>> #image = kwimage.grab_test_image('checkerboard') >>> transform = Affine.random() @ Affine.scale(0.05) >>> transform = Affine.scale(0.02) >>> warped1 = warp_affine(image, transform, dsize='positive', antialias=1, interpolation='nearest') >>> warped2 = warp_affine(image, transform, dsize='positive', antialias=0) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=1, nCols=2) >>> kwplot.imshow(warped1, pnum=pnum_(), title='antialias=True') >>> kwplot.imshow(warped2, pnum=pnum_(), title='antialias=False') >>> kwplot.show_if_requested()

Example
>>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> from kwimage.transform import Affine >>> image = kwimage.grab_test_image('astro') >>> image = kwimage.grab_test_image('checkerboard') >>> transform = Affine.random() @ Affine.scale((.1, 1.2)) >>> warped1 = warp_affine(image, transform, dsize='positive', antialias=1) >>> warped2 = warp_affine(image, transform, dsize='positive', antialias=0) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=1, nCols=2) >>> kwplot.imshow(warped1, pnum=pnum_(), title='antialias=True') >>> kwplot.imshow(warped2, pnum=pnum_(), title='antialias=False') >>> kwplot.show_if_requested()
Example
>>> # Test the case where the input data is empty or the target canvas >>> # is empty, this should be handled like boundary effects >>> import kwimage >>> image = np.random.rand(1, 1, 3) >>> transform = kwimage.Affine.random() >>> result = kwimage.warp_affine(image, transform, dsize=(0, 0)) >>> assert result.shape == (0, 0, 3) >>> # >>> empty_image = np.random.rand(0, 1, 3) >>> result = kwimage.warp_affine(empty_image, transform, dsize=(10, 10)) >>> assert result.shape == (10, 10, 3) >>> # >>> empty_image = np.random.rand(0, 1, 3) >>> result = kwimage.warp_affine(empty_image, transform, dsize=(10, 0)) >>> assert result.shape == (0, 10, 3)
Example
>>> # Demo difference between positive and content dsize >>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> from kwimage.transform import Affine >>> image = kwimage.grab_test_image('astro', dsize=(512, 512)) >>> transform = Affine.coerce(offset=(-100, -50), scale=2, theta=0.1) >>> # When warping other images or geometry along with this image >>> # it is important to account for the modified transform when >>> # setting dsize='content'. If dsize='positive', the transform >>> # will remain unchanged wrt other aligned images / geometries. >>> poly = kwimage.Boxes([[350, 5, 130, 290]], 'xywh').to_polygons()[0] >>> # Apply the warping to the images >>> warped_pos, info_pos = warp_affine(image, transform, dsize='positive', return_info=True) >>> warped_con, info_con = warp_affine(image, transform, dsize='content', return_info=True) >>> assert info_pos['dsize'] == (919, 1072) >>> assert info_con['dsize'] == (1122, 1122) >>> assert info_pos['transform'] == transform >>> # Demo the correct and incorrect way to apply transforms >>> poly_pos = poly.warp(transform) >>> poly_con = poly.warp(info_con['transform']) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> # show original >>> kwplot.imshow(image, pnum=(1, 3, 1), title='original') >>> poly.draw(color='green', alpha=0.5, border=True) >>> # show positive warped >>> kwplot.imshow(warped_pos, pnum=(1, 3, 2), title='dsize=positive') >>> poly_pos.draw(color='purple', alpha=0.5, border=True) >>> # show content warped >>> ax = kwplot.imshow(warped_con, pnum=(1, 3, 3), title='dsize=content')[1] >>> poly_con.draw(color='dodgerblue', alpha=0.5, border=True) # correct >>> poly_pos.draw(color='orangered', alpha=0.5, border=True) # incorrect >>> cc = poly_con.to_shapely().centroid >>> cp = poly_pos.to_shapely().centroid >>> ax.text(cc.x, cc.y + 250, 'correctly transformed', color='dodgerblue', >>> backgroundcolor=(0, 0, 0, 0.7), horizontalalignment='center') >>> ax.text(cp.x, cp.y - 250, 'incorrectly transformed', color='orangered', >>> backgroundcolor=(0, 0, 0, 0.7), horizontalalignment='center') >>> kwplot.show_if_requested()

Example
>>> # Demo piecewise transform >>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> from kwimage.transform import Affine >>> image = kwimage.grab_test_image('pm5644') >>> transform = Affine.coerce(offset=(-100, -50), scale=2, theta=0.1) >>> warped_piecewise, info = warp_affine(image, transform, dsize='positive', return_info=True, large_warp_dim=32) >>> warped_normal, info = warp_affine(image, transform, dsize='positive', return_info=True, large_warp_dim=None) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image, pnum=(1, 3, 1), title='original') >>> kwplot.imshow(warped_normal, pnum=(1, 3, 2), title='normal warp') >>> kwplot.imshow(warped_piecewise, pnum=(1, 3, 3), title='piecewise warp')
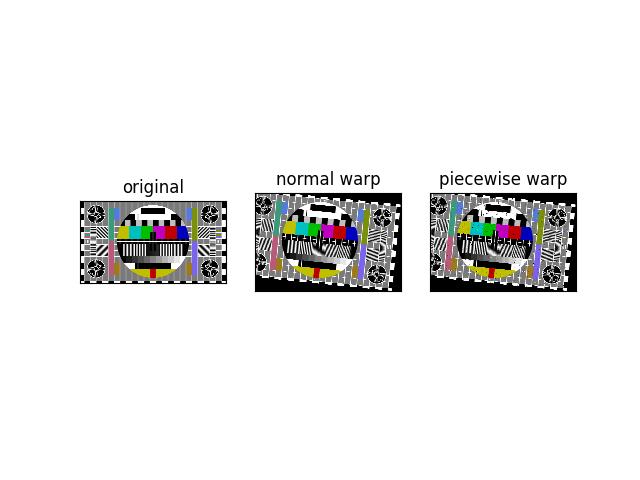
Example
>>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> # TODO: Explain why the bottom left is interpolated with 0's >>> # And not 2s, probably has to do with interpretation of pixels >>> # as points and not areas. >>> image = np.full((6, 6), fill_value=3, dtype=np.uint8) >>> transform = kwimage.Affine.eye() >>> transform = kwimage.Affine.coerce(offset=.5) @ transform >>> transform = kwimage.Affine.coerce(scale=2) @ transform >>> warped = kwimage.warp_affine(image, transform, dsize=(12, 12))
Example
>>> # Demo how nans are handled >>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> image = kwimage.grab_test_image('pm5644') >>> image = kwimage.ensure_float01(image) >>> image[100:300, 400:700] = np.nan >>> transform = kwimage.Affine.coerce(scale=0.05, offset=10.5, theta=0.3, shearx=0.2) >>> warped1 = warp_affine(image, transform, dsize='positive', antialias=1, interpolation='linear', border_value=0) >>> warped2 = warp_affine(image, transform, dsize='positive', antialias=0, border_value=np.nan) >>> assert np.isnan(warped1).any() >>> assert np.isnan(warped2).any() >>> assert warped1[np.isnan(warped1).any(axis=2)].all() >>> assert warped2[np.isnan(warped2).any(axis=2)].all() >>> print('warped1.shape = {!r}'.format(warped1.shape)) >>> print('warped2.shape = {!r}'.format(warped2.shape)) >>> assert warped2.shape == warped1.shape >>> warped2[np.isnan(warped2).any(axis=2)] >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=1, nCols=3) >>> image_canvas = kwimage.fill_nans_with_checkers(image) >>> warped1_canvas = kwimage.fill_nans_with_checkers(warped1) >>> warped2_canvas = kwimage.fill_nans_with_checkers(warped2) >>> kwplot.imshow(image_canvas, pnum=pnum_(), title='original') >>> kwplot.imshow(warped1_canvas, pnum=pnum_(), title='antialias=True, border=0') >>> kwplot.imshow(warped2_canvas, pnum=pnum_(), title='antialias=False, border=nan') >>> kwplot.show_if_requested()

Example
>>> # Demo how of how we also handle masked arrays >>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> _image = kwimage.grab_test_image('pm5644') >>> _image = kwimage.ensure_float01(_image) >>> _image[100:200, 400:700] = np.nan >>> mask = np.isnan(_image) >>> data = np.nan_to_num(_image) >>> image = np.ma.MaskedArray(data=data, mask=mask) >>> transform = kwimage.Affine.coerce(scale=0.05, offset=10.5, theta=0.3, shearx=0.2) >>> warped1 = warp_affine(image, transform, dsize='positive', antialias=1, interpolation='linear') >>> assert isinstance(warped1, np.ma.MaskedArray) >>> warped2 = warp_affine(image, transform, dsize='positive', antialias=0) >>> print('warped1.shape = {!r}'.format(warped1.shape)) >>> print('warped2.shape = {!r}'.format(warped2.shape)) >>> assert warped2.shape == warped1.shape >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=1, nCols=2) >>> kwplot.imshow(warped1, pnum=pnum_(), title='antialias=True') >>> kwplot.imshow(warped2, pnum=pnum_(), title='antialias=False') >>> kwplot.show_if_requested()

- kwimage.warp_image(image, transform, dsize=None, antialias=False, interpolation='linear', border_mode=None, border_value=0, large_warp_dim=None, return_info=False)[source]¶
Applies an transformation to an image with optional antialiasing.
- Parameters:
image (ndarray) – the input image as a numpy array. Note: this is passed directly to cv2, so it is best to ensure that it is contiguous and using a dtype that cv2 can handle.
transform (ndarray | dict | kwimage.Matrix) – a coercable affine or projective matrix. See
kwimage.Affineandkwimage.Projectivefor details on what can be coerced.dsize (Tuple[int, int] | None | str) – A integer width and height tuple of the resulting “canvas” image. If None, then the input image size is used.
If specified as a string, dsize is computed based on the given heuristic.
If ‘positive’ (or ‘auto’), dsize is computed such that the positive coordinates of the warped image will fit in the new canvas. In this case, any pixel that maps to a negative coordinate will be clipped. This has the property that the input transformation is not modified.
If ‘content’ (or ‘max’), the transform is modified with an extra translation such that both the positive and negative coordinates of the warped image will fit in the new canvas.
antialias (bool) – if True determines if the transform is downsampling and applies antialiasing via gaussian a blur. Defaults to False
interpolation (str | int) – interpolation code or cv2 integer. Interpolation codes are linear, nearest, cubic, lancsoz, and area. Defaults to “linear”.
border_mode (str | int) – Border code or cv2 integer. Border codes are constant (default) replicate, reflect, wrap, reflect101, and transparent.
border_value (int | float | Iterable[int | float]) – Used as the fill value if border_mode is constant. Otherwise this is ignored. Defaults to 0, but can also be defaulted to nan. if border_value is a scalar and there are multiple channels, the value is applied to all channels. More than 4 unique border values for individual channels will cause an error. See OpenCV #22283 for details. In the future we may accept np.ma and return a masked array, but for now that is not implemented.
large_warp_dim (int | None | str) – If specified, perform the warp piecewise in chunks of the specified size. If “auto”, it is set to the maximum “short” value in numpy. This works around a limitation of cv2.warpAffine, which must have image dimensions < SHRT_MAX (=32767 in version 4.5.3)
return_info (bool) – if True, returns information about the operation. In the case where dsize=”content”, this includes the modified transformation.
- Returns:
the warped image, or if return info is True, the warped image and the info dictionary.
- Return type:
ndarray | Tuple[ndarray, Dict]
Example
>>> from kwimage.im_cv2 import * # NOQA >>> import kwimage >>> image = kwimage.grab_test_image('paraview') >>> tf_homog = kwimage.Projective.random(rng=30342110) @ kwimage.Projective.coerce(uv=[0.001, 0.001]) >>> tf_aff = kwimage.Affine.coerce(ub.udict(tf_homog.decompose()) - {'uv'}) >>> tf_uv = kwimage.Projective.coerce(ub.udict(tf_homog.decompose()) & {'uv'}) >>> warped1 = kwimage.warp_image(image, tf_homog, dsize='positive') >>> warped2 = kwimage.warp_image(image, tf_aff, dsize='positive') >>> warped3 = kwimage.warp_image(image, tf_uv, dsize='positive') >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> pnum_ = kwplot.PlotNums(nRows=2, nCols=2) >>> kwplot.imshow(warped1, pnum=pnum_(), title='projective warp') >>> kwplot.imshow(warped2, pnum=pnum_(), title='affine warp') >>> kwplot.imshow(warped3, pnum=pnum_(), title='projective part') >>> kwplot.show_if_requested()

- kwimage.warp_points(matrix, pts, homog_mode='divide')[source]¶
Warp ND points / coordinates using a transformation matrix.
Homogoenous coordinates are added on the fly if needed. Works with both numpy and torch.
- Parameters:
matrix (ArrayLike) – [D1 x D2] transformation matrix. if using homogenous coordinates D2=D + 1, otherwise D2=D. if using homogenous coordinates and the matrix represents an Affine transformation, then either D1=D or D1=D2, i.e. the last row of zeros and a one is optional.
pts (ArrayLike) – [N1 x … x D] points (usually x, y). If points are already in homogenous space, then the output will be returned in homogenous space. D is the dimensionality of the points. The leading axis may take any shape, but usually, shape will be [N x D] where N is the number of points.
homog_mode (str) – what to do for homogenous coordinates. Can either divide, keep, or drop. Defaults do ‘divide’.
- Retrns:
new_pts (ArrayLike): the points after being transformed by the matrix
Example
>>> from kwimage.util_warp import * # NOQA >>> # --- with numpy >>> rng = np.random.RandomState(0) >>> pts = rng.rand(10, 2) >>> matrix = rng.rand(2, 2) >>> warp_points(matrix, pts) >>> # --- with torch >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> pts = torch.Tensor(pts) >>> matrix = torch.Tensor(matrix) >>> warp_points(matrix, pts)
Example
>>> from kwimage.util_warp import * # NOQA >>> # --- with numpy >>> pts = np.ones((10, 2)) >>> matrix = np.diag([2, 3, 1]) >>> ra = warp_points(matrix, pts) >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> rb = warp_points(torch.Tensor(matrix), torch.Tensor(pts)) >>> assert np.allclose(ra, rb.numpy())
Example
>>> from kwimage.util_warp import * # NOQA >>> # test different cases >>> rng = np.random.RandomState(0) >>> # Test 3x3 style projective matrices >>> pts = rng.rand(1000, 2) >>> matrix = rng.rand(3, 3) >>> ra33 = warp_points(matrix, pts) >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> rb33 = warp_points(torch.Tensor(matrix), torch.Tensor(pts)) >>> assert np.allclose(ra33, rb33.numpy()) >>> # Test opencv style affine matrices >>> pts = rng.rand(10, 2) >>> matrix = rng.rand(2, 3) >>> ra23 = warp_points(matrix, pts) >>> rb23 = warp_points(torch.Tensor(matrix), torch.Tensor(pts)) >>> assert np.allclose(ra33, rb33.numpy())
- kwimage.warp_projective(image, transform, dsize=None, antialias=False, interpolation='linear', border_mode=None, border_value=0, large_warp_dim=None, return_info=False)[source]¶
Applies an projective transformation to an image with optional antialiasing.
- Parameters:
image (ndarray) – the input image as a numpy array. Note: this is passed directly to cv2, so it is best to ensure that it is contiguous and using a dtype that cv2 can handle.
transform (ndarray | dict | kwimage.Projective) – a coercable projective matrix. See
kwimage.Projectivefor details on what can be coerced.dsize (Tuple[int, int] | None | str) – A integer width and height tuple of the resulting “canvas” image. If None, then the input image size is used.
If specified as a string, dsize is computed based on the given heuristic.
If ‘positive’ (or ‘auto’), dsize is computed such that the positive coordinates of the warped image will fit in the new canvas. In this case, any pixel that maps to a negative coordinate will be clipped. This has the property that the input transformation is not modified.
If ‘content’ (or ‘max’), the transform is modified with an extra translation such that both the positive and negative coordinates of the warped image will fit in the new canvas.
antialias (bool) – if True determines if the transform is downsampling and applies antialiasing via gaussian a blur. Defaults to False
interpolation (str | int) – interpolation code or cv2 integer. Interpolation codes are linear, nearest, cubic, lancsoz, and area. Defaults to “linear”.
border_mode (str | int) – Border code or cv2 integer. Border codes are constant (default) replicate, reflect, wrap, reflect101, and transparent.
border_value (int | float | Iterable[int | float]) – Used as the fill value if border_mode is constant. Otherwise this is ignored. Defaults to 0, but can also be defaulted to nan. if border_value is a scalar and there are multiple channels, the value is applied to all channels. More than 4 unique border values for individual channels will cause an error. See OpenCV #22283 for details. In the future we may accept np.ma and return a masked array, but for now that is not implemented.
large_warp_dim (int | None | str) – If specified, perform the warp piecewise in chunks of the specified size. If “auto”, it is set to the maximum “short” value in numpy. This works around a limitation of cv2.warpAffine, which must have image dimensions < SHRT_MAX (=32767 in version 4.5.3)
return_info (bool) – if True, returns information about the operation. In the case where dsize=”content”, this includes the modified transformation.
- Returns:
the warped image, or if return info is True, the warped image and the info dictionary.
- Return type:
ndarray | Tuple[ndarray, Dict]
- kwimage.warp_tensor(inputs, mat, output_dims, mode='bilinear', padding_mode='zeros', isinv=False, ishomog=None, align_corners=False, new_mode=False)[source]¶
A pytorch implementation of warp affine that works similarly to
cv2.warpAffine()andcv2.warpPerspective().It is possible to use 3x3 transforms to warp 2D image data. It is also possible to use 4x4 transforms to warp 3D volumetric data.
- Parameters:
inputs (Tensor) – tensor to warp. Up to 3 (determined by output_dims) of the trailing space-time dimensions are warped. Best practice is to use inputs with the shape in [B, C, *DIMS].
mat (Tensor) – either a 3x3 / 4x4 single transformation matrix to apply to all inputs or Bx3x3 or Bx4x4 tensor that specifies a transformation matrix for each batch item.
output_dims (Tuple[int, …]) – The output space-time dimensions. This can either be in the form (W,), (H, W), or (D, H, W).
mode (str) – Can be bilinear or nearest. See torch.nn.functional.grid_sample
padding_mode (str) – Can be zeros, border, or reflection. See torch.nn.functional.grid_sample.
isinv (bool) – Set to true if mat is the inverse transform
ishomog (bool) – Set to True if the matrix is non-affine
align_corners (bool) – Note the default of False does not work correctly with grid_sample in torch <= 1.2, but using align_corners=True isnt typically what you want either. We will be stuck with buggy functionality until torch 1.3 is released.
However, using align_corners=0 does seem to reasonably correspond with opencv behavior.
- Returns:
warped tensor
- Return type:
Tensor
Note
Also, it may be possible to speed up the code with F.affine_grid
- KNOWN ISSUE: There appears to some difference with cv2.warpAffine when
rotation or shear are non-zero. I’m not sure what the cause is. It may just be floating point issues, but Im’ not sure.
See issues in [TorchAffineTransform] and [TorchIssue15386].
Todo
[ ] FIXME: see example in Mask.scale where this algo breaks when the matrix is 2x3
[ ] Make this algo work when matrix ix 2x2
References
Example
>>> # Create a relatively simple affine matrix >>> # xdoctest: +REQUIRES(module:torch) >>> import skimage >>> import torch >>> mat = torch.FloatTensor(skimage.transform.AffineTransform( >>> translation=[1, -1], scale=[.532, 2], >>> rotation=0, shear=0, >>> ).params) >>> # Create inputs and an output dimension >>> input_shape = [1, 1, 4, 5] >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> output_dims = (11, 7) >>> # Warp with our code >>> result1 = warp_tensor(inputs, mat, output_dims=output_dims, align_corners=0) >>> print('result1 =\n{}'.format(ub.urepr(result1.cpu().numpy()[0, 0], precision=2))) >>> # Warp with opencv >>> import cv2 >>> cv2_M = mat.cpu().numpy()[0:2] >>> src = inputs[0, 0].cpu().numpy() >>> dsize = tuple(output_dims[::-1]) >>> result2 = cv2.warpAffine(src, cv2_M, dsize=dsize, flags=cv2.INTER_LINEAR) >>> print('result2 =\n{}'.format(ub.urepr(result2, precision=2))) >>> # Ensure the results are the same (up to floating point errors) >>> assert np.all(np.isclose(result1[0, 0].cpu().numpy(), result2, atol=1e-2, rtol=1e-2))
Example
>>> # Create a relatively simple affine matrix >>> # xdoctest: +REQUIRES(module:torch) >>> import skimage >>> import torch >>> mat = torch.FloatTensor(skimage.transform.AffineTransform( >>> rotation=0.01, shear=0.1).params) >>> # Create inputs and an output dimension >>> input_shape = [1, 1, 4, 5] >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> output_dims = (11, 7) >>> # Warp with our code >>> result1 = warp_tensor(inputs, mat, output_dims=output_dims) >>> print('result1 =\n{}'.format(ub.urepr(result1.cpu().numpy()[0, 0], precision=2, supress_small=True))) >>> print('result1.shape = {}'.format(result1.shape)) >>> # Warp with opencv >>> import cv2 >>> cv2_M = mat.cpu().numpy()[0:2] >>> src = inputs[0, 0].cpu().numpy() >>> dsize = tuple(output_dims[::-1]) >>> result2 = cv2.warpAffine(src, cv2_M, dsize=dsize, flags=cv2.INTER_LINEAR) >>> print('result2 =\n{}'.format(ub.urepr(result2, precision=2))) >>> print('result2.shape = {}'.format(result2.shape)) >>> # Ensure the results are the same (up to floating point errors) >>> # NOTE: The floating point errors seem to be significant for rotation / shear >>> assert np.all(np.isclose(result1[0, 0].cpu().numpy(), result2, atol=1, rtol=1e-2))
Example
>>> # Create a random affine matrix >>> # xdoctest: +REQUIRES(module:torch) >>> import skimage >>> import torch >>> rng = np.random.RandomState(0) >>> mat = torch.FloatTensor(skimage.transform.AffineTransform( >>> translation=rng.randn(2), scale=1 + rng.randn(2), >>> rotation=rng.randn() / 10., shear=rng.randn() / 10., >>> ).params) >>> # Create inputs and an output dimension >>> input_shape = [1, 1, 5, 7] >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> output_dims = (3, 11) >>> # Warp with our code >>> result1 = warp_tensor(inputs, mat, output_dims=output_dims, align_corners=0) >>> print('result1 =\n{}'.format(ub.urepr(result1.cpu().numpy()[0, 0], precision=2))) >>> # Warp with opencv >>> import cv2 >>> cv2_M = mat.cpu().numpy()[0:2] >>> src = inputs[0, 0].cpu().numpy() >>> dsize = tuple(output_dims[::-1]) >>> result2 = cv2.warpAffine(src, cv2_M, dsize=dsize, flags=cv2.INTER_LINEAR) >>> print('result2 =\n{}'.format(ub.urepr(result2, precision=2))) >>> # Ensure the results are the same (up to floating point errors) >>> # NOTE: The errors seem to be significant for rotation / shear >>> assert np.all(np.isclose(result1[0, 0].cpu().numpy(), result2, atol=1, rtol=1e-2))
Example
>>> # Test 3D warping with identity >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> mat = torch.eye(4) >>> input_dims = [2, 3, 3] >>> output_dims = (2, 3, 3) >>> input_shape = [1, 1] + input_dims >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> result = warp_tensor(inputs, mat, output_dims=output_dims) >>> print('result =\n{}'.format(ub.urepr(result.cpu().numpy()[0, 0], precision=2))) >>> assert torch.all(inputs == result)
Example
>>> # Test 3D warping with scaling >>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> mat = torch.FloatTensor([ >>> [0.8, 0, 0, 0], >>> [ 0, 1.0, 0, 0], >>> [ 0, 0, 1.2, 0], >>> [ 0, 0, 0, 1], >>> ]) >>> input_dims = [2, 3, 3] >>> output_dims = (2, 3, 3) >>> input_shape = [1, 1] + input_dims >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> result = warp_tensor(inputs, mat, output_dims=output_dims, align_corners=0) >>> print('result =\n{}'.format(ub.urepr(result.cpu().numpy()[0, 0], precision=2))) result = np.array([[[ 0. , 1.25, 1. ], [ 3. , 4.25, 2.5 ], [ 6. , 7.25, 4. ]], ... [[ 7.5 , 8.75, 4.75], [10.5 , 11.75, 6.25], [13.5 , 14.75, 7.75]]], dtype=np.float32)
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> mat = torch.eye(3) >>> input_dims = [5, 7] >>> output_dims = (11, 7) >>> for n_prefix_dims in [0, 1, 2, 3, 4, 5]: >>> input_shape = [2] * n_prefix_dims + input_dims >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> result = warp_tensor(inputs, mat, output_dims=output_dims) >>> #print('result =\n{}'.format(ub.urepr(result.cpu().numpy(), precision=2))) >>> print(result.shape)
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> mat = torch.eye(4) >>> input_dims = [5, 5, 5] >>> output_dims = (6, 6, 6) >>> for n_prefix_dims in [0, 1, 2, 3, 4, 5]: >>> input_shape = [2] * n_prefix_dims + input_dims >>> inputs = torch.arange(int(np.prod(input_shape))).reshape(*input_shape).float() >>> result = warp_tensor(inputs, mat, output_dims=output_dims) >>> #print('result =\n{}'.format(ub.urepr(result.cpu().numpy(), precision=2))) >>> print(result.shape)