kwimage.structs.coords module¶
Coordinates the fundamental “point” datatype. They do not contain metadata, only geometry. See the Points data type for a structure that maintains metadata on top of coordinate data.
- class kwimage.structs.coords.Coords(data=None, meta=None)[source]¶
-
A data structure to store n-dimensional coordinate geometry.
Currently it is up to the user to maintain what coordinate system this geometry belongs to.
Note
This class was designed to hold coordinates in r/c format, but in general this class is anostic to dimension ordering as long as you are consistent. However, there are two places where this matters: (1) drawing and (2) gdal/imgaug-warping. In these places we will assume x/y for legacy reasons. This may change in the future.
The term axes with resepct to
Coordsalways refers to the final numpy axis. In other words the final numpy-axis represents ALL of the coordinate-axes.CommandLine
xdoctest -m kwimage.structs.coords Coords
Example
>>> from kwimage.structs.coords import * # NOQA >>> import kwarray >>> rng = kwarray.ensure_rng(0) >>> self = Coords.random(num=4, dim=3, rng=rng) >>> print('self = {}'.format(self)) self = <Coords(data= array([[0.5488135 , 0.71518937, 0.60276338], [0.54488318, 0.4236548 , 0.64589411], [0.43758721, 0.891773 , 0.96366276], [0.38344152, 0.79172504, 0.52889492]]))> >>> matrix = rng.rand(4, 4) >>> self.warp(matrix) <Coords(data= array([[0.71037426, 1.25229659, 1.39498435], [0.60799503, 1.26483447, 1.42073131], [0.72106004, 1.39057144, 1.38757508], [0.68384299, 1.23914654, 1.29258196]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[3.5488135 , 3.71518937, 3.60276338], [3.54488318, 3.4236548 , 3.64589411], [3.43758721, 3.891773 , 3.96366276], [3.38344152, 3.79172504, 3.52889492]]))> >>> self.translate(3, inplace=True) <Coords(data= array([[6.5488135 , 6.71518937, 6.60276338], [6.54488318, 6.4236548 , 6.64589411], [6.43758721, 6.891773 , 6.96366276], [6.38344152, 6.79172504, 6.52889492]]))> >>> self.scale(2) <Coords(data= array([[13.09762701, 13.43037873, 13.20552675], [13.08976637, 12.8473096 , 13.29178823], [12.87517442, 13.783546 , 13.92732552], [12.76688304, 13.58345008, 13.05778984]]))> >>> # xdoctest: +REQUIRES(module:torch) >>> self.tensor() >>> self.tensor().tensor().numpy().numpy() >>> self.numpy() >>> #self.draw_on()
- property dtype¶
- property dim¶
- property shape¶
- classmethod random(num=1, dim=2, rng=None, meta=None)[source]¶
Makes random coordinates; typically for testing purposes
- compress(flags, axis=0, inplace=False)[source]¶
Filters items based on a boolean criterion
- Parameters:
flags (ArrayLike) – true for items to be kept. Extended type: ArrayLike[bool].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
filtered coords
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords.random(10, rng=0) >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self)) <Coords(data=array([], shape=(0, 2), dtype=float64))> >>> # xdoctest: +REQUIRES(module:torch) >>> self = self.tensor() >>> self.compress([True] * len(self)) >>> self.compress([False] * len(self))
- take(indices, axis=0, inplace=False)[source]¶
Takes a subset of items at specific indices
- Parameters:
indices (ArrayLike) – indexes of items to take. Extended type ArrayLike[int].
axis (int) – you usually want this to be 0
inplace (bool) – if True, modifies this object
- Returns:
filtered coords
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords(np.array([[25, 30, 15, 10]])) >>> self.take([0]) <Coords(data=array([[25, 30, 15, 10]]))> >>> self.take([]) <Coords(data=array([], shape=(0, 4), dtype=...))>
- astype(dtype, inplace=False)[source]¶
Changes the data type
- Parameters:
dtype – new type
inplace (bool) – if True, modifies this object
- Returns:
modified coordinates
- Return type:
- round(decimals=0, inplace=False)[source]¶
Rounds data to the specified decimal place
- Parameters:
inplace (bool) – if True, modifies this object
decimals (int) – number of decimal places to round to
- Returns:
modified coordinates
- Return type:
Example
>>> import kwimage >>> self = kwimage.Coords.random(3).scale(10) >>> self.round()
- view(*shape)[source]¶
Passthrough method to view or reshape
- Parameters:
*shape – new shape of the data
- Returns:
modified coordinates
- Return type:
Example
>>> self = Coords.random(6, dim=4).numpy() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4] >>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(6, dim=4).tensor() >>> assert list(self.view(3, 2, 4).data.shape) == [3, 2, 4]
- classmethod concatenate(coords, axis=0)[source]¶
Concatenates lists of coordinates together
- Parameters:
coords (Sequence[Coords]) – list of coords to concatenate
axis (int) – axis to stack on. Defaults to 0.
- Returns:
stacked coords
- Return type:
CommandLine
xdoctest -m kwimage.structs.coords Coords.concatenate
Example
>>> coords = [Coords.random(3) for _ in range(3)] >>> new = Coords.concatenate(coords) >>> assert len(new) == 9 >>> assert np.all(new.data[3:6] == coords[1].data)
- property device¶
If the backend is torch returns the data device, otherwise None
- property _impl¶
Returns the internal tensor/numpy ArrayAPI implementation
- tensor(device=NoParam)[source]¶
Converts numpy to tensors. Does not change memory if possible.
- Returns:
modified coordinates
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).numpy() >>> newself = self.tensor() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- numpy()[source]¶
Converts tensors to numpy. Does not change memory if possible.
- Returns:
modified coordinates
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Coords.random(3).tensor() >>> newself = self.numpy() >>> self.data[0, 0] = 0 >>> assert newself.data[0, 0] == 0 >>> self.data[0, 0] = 1 >>> assert self.data[0, 0] == 1
- reorder_axes(new_order, inplace=False)[source]¶
Change the ordering of the coordinate axes.
- Parameters:
new_order (Tuple[int]) –
new_order[i]should specify which axes in the original coordinates should be mapped to thei-thposition in the returned axes.inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Note
This is the ordering of the “columns” in final numpy axis, not the numpy axes themselves.
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords(data=np.array([ >>> [7, 11], >>> [13, 17], >>> [21, 23], >>> ])) >>> new = self.reorder_axes((1, 0)) >>> print('new = {!r}'.format(new)) new = <Coords(data= array([[11, 7], [17, 13], [23, 21]]))>
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.reorder_axes((1, 0)) >>> # Remapping using 1, 0 reverses the axes >>> assert np.all(new.data[:, 0] == self.data[:, 1]) >>> assert np.all(new.data[:, 1] == self.data[:, 0]) >>> # Remapping using 0, 1 does nothing >>> eye = self.reorder_axes((0, 1)) >>> assert np.all(eye.data == self.data) >>> # Remapping using 0, 0, destroys the 1-th column >>> bad = self.reorder_axes((0, 0)) >>> assert np.all(bad.data[:, 0] == self.data[:, 0]) >>> assert np.all(bad.data[:, 1] == self.data[:, 0])
- warp(transform, input_dims=None, output_dims=None, inplace=False)[source]¶
Generalized coordinate transform.
- Parameters:
transform (SKImageGeometricTransform | ArrayLike | Augmenter | Callable) – scikit-image tranform, a 3x3 transformation matrix, an imgaug Augmenter, or generic callable which transforms an NxD ndarray.
input_dims (Tuple) – shape of the image these objects correspond to (only needed / used when transform is an imgaug augmenter)
output_dims (Tuple) – unused in non-raster structures, only exists for compatibility.
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Note
Let D = self.dims
- transformation matrices can be either:
(D + 1) x (D + 1) # for homog
D x D # for scale / rotate
D x (D + 1) # for affine
Example
>>> from kwimage.structs.coords import * # NOQA >>> import skimage >>> self = Coords.random(10, rng=0) >>> transform = skimage.transform.AffineTransform(scale=(2, 2)) >>> new = self.warp(transform) >>> assert np.all(new.data == self.scale(2).data)
Doctest
>>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(np.eye(3)).data == self.data) >>> assert np.all(self.warp(np.eye(2)).data == self.data)
Doctest
>>> # xdoctest: +REQUIRES(module:osgeo) >>> from osgeo import osr >>> wgs84_crs = osr.SpatialReference() >>> wgs84_crs.ImportFromEPSG(4326) >>> dst_crs = osr.SpatialReference() >>> dst_crs.ImportFromEPSG(2927) >>> transform = osr.CoordinateTransformation(wgs84_crs, dst_crs) >>> self = Coords.random(10, rng=0) >>> new = self.warp(transform) >>> assert np.all(new.data != self.data)
>>> # Alternative using generic func >>> def _gdal_coord_tranform(pts): ... return np.array([transform.TransformPoint(x, y, 0)[0:2] ... for x, y in pts]) >>> alt = self.warp(_gdal_coord_tranform) >>> assert np.all(alt.data != self.data) >>> assert np.all(alt.data == new.data)
Doctest
>>> # can use a generic function >>> def func(xy): ... return np.zeros_like(xy) >>> self = Coords.random(10, rng=0) >>> assert np.all(self.warp(func).data == 0)
- _warp_imgaug(augmenter, input_dims, inplace=False)[source]¶
Warps by applying an augmenter from the imgaug library
Note
We are assuming you are using X/Y coordinates here.
- Parameters:
augmenter (imgaug.augmenters.Augmenter)
input_dims (Tuple) – h/w of the input image
inplace (bool) – if True, modifies data inplace
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/coords.py Coords._warp_imgaug
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> from kwimage.structs.coords import * # NOQA >>> import imgaug >>> input_dims = (10, 10) >>> self = Coords.random(10).scale(input_dims) >>> augmenter = imgaug.augmenters.Fliplr(p=1) >>> new = self._warp_imgaug(augmenter, input_dims) >>> # y coordinate should not change >>> assert np.allclose(self.data[:, 1], new.data[:, 1]) >>> assert np.allclose(input_dims[0] - self.data[:, 0], new.data[:, 0])
>>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> from matplotlib import pyplot as pl >>> ax = plt.gca() >>> ax.set_xlim(0, input_dims[0]) >>> ax.set_ylim(0, input_dims[1]) >>> self.draw(color='red', alpha=.4, radius=0.1) >>> new.draw(color='blue', alpha=.4, radius=0.1)
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> from kwimage.structs.coords import * # NOQA >>> import imgaug >>> input_dims = (32, 32) >>> inplace = 0 >>> self = Coords.random(1000, rng=142).scale(input_dims).scale(.8) >>> self.data = self.data.astype(np.int32).astype(np.float32) >>> augmenter = imgaug.augmenters.CropAndPad(px=(-4, 4), keep_size=1).to_deterministic() >>> new = self._warp_imgaug(augmenter, input_dims) >>> # Change should be linear >>> norm1 = (self.data - self.data.min(axis=0)) / (self.data.max(axis=0) - self.data.min(axis=0)) >>> norm2 = (new.data - new.data.min(axis=0)) / (new.data.max(axis=0) - new.data.min(axis=0)) >>> diff = norm1 - norm2 >>> assert np.allclose(diff, 0, atol=1e-6, rtol=1e-4) >>> #assert np.allclose(self.data[:, 1], new.data[:, 1]) >>> #assert np.allclose(input_dims[0] - self.data[:, 0], new.data[:, 0]) >>> # xdoctest: +REQUIRES(--show) >>> import kwimage >>> im = kwimage.imresize(kwimage.grab_test_image(), dsize=input_dims[::-1]) >>> new_im = augmenter.augment_image(im) >>> import kwplot >>> plt = kwplot.autoplt() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(im, pnum=(1, 2, 1), fnum=1) >>> self.draw(color='red', alpha=.8, radius=0.5) >>> kwplot.imshow(new_im, pnum=(1, 2, 2), fnum=1) >>> new.draw(color='blue', alpha=.8, radius=0.5, coord_axes=[1, 0])
- to_imgaug(input_dims)[source]¶
Translate to an imgaug object
- Returns:
imgaug data structure
- Return type:
imgaug.KeypointsOnImage
Example
>>> # xdoctest: +REQUIRES(module:imgaug) >>> import kwimage >>> import numpy as np >>> self = kwimage.Coords.random(10) >>> input_dims = (10, 10) >>> kpoi = self.to_imgaug(input_dims) >>> new = kwimage.Coords.from_imgaug(kpoi) >>> assert np.allclose(new.data, self.data)
- scale(factor, about=None, output_dims=None, inplace=False)[source]¶
Scale coordinates by a factor
- Parameters:
factor (float | Tuple[float, float]) – scale factor as either a scalar or per-dimension tuple.
about (Tuple | None) – if unspecified scales about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, rng=0) >>> new = self.scale(10) >>> assert new.data.max() <= 10
>>> self = Coords.random(10, rng=0) >>> self.data = (self.data * 10).astype(int) >>> new = self.scale(10) >>> assert new.data.dtype.kind == 'i' >>> new = self.scale(10.0) >>> assert new.data.dtype.kind == 'f'
- translate(offset, output_dims=None, inplace=False)[source]¶
Shift the coordinates
- Parameters:
offset (float | Tuple[float, float]) – transation offset as either a scalar or a per-dimension tuple.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=3, rng=0) >>> new = self.translate(10) >>> assert new.data.min() >= 10 >>> assert new.data.max() <= 11 >>> Coords.random(3, dim=3, rng=0) >>> Coords.random(3, dim=3, rng=0).translate((1, 2, 3))
- rotate(theta, about=None, output_dims=None, inplace=False)[source]¶
Rotate the coordinates about a point.
- Parameters:
theta (float) – rotation angle in radians
about (Tuple | None) – if unspecified rotates about the origin (0, 0), otherwise the rotation is about this point.
output_dims (Tuple) – unused in non-raster spatial structures
inplace (bool) – if True, modifies data inplace
- Returns:
modified coordinates
- Return type:
Todo
[ ] Generalized ND Rotations?
References
https://math.stackexchange.com/questions/197772/gen-rot-matrix
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=2, rng=0) >>> theta = np.pi / 2 >>> new = self.rotate(theta)
>>> # Test rotate agrees with warp >>> sin_ = np.sin(theta) >>> cos_ = np.cos(theta) >>> rot_ = np.array([[cos_, -sin_], [sin_, cos_]]) >>> new2 = self.warp(rot_) >>> assert np.allclose(new.data, new2.data)
>>> # >>> # Rotate about a custom point >>> theta = np.pi / 2 >>> new3 = self.rotate(theta, about=(0.5, 0.5)) >>> # >>> # Rotate about the center of mass >>> about = self.data.mean(axis=0) >>> new4 = self.rotate(theta, about=about) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.figure(fnum=1, doclf=True) >>> plt = kwplot.autoplt() >>> self.draw(radius=0.01, color='blue', alpha=.5, coord_axes=[1, 0], setlim='grow') >>> plt.gca().set_aspect('equal') >>> new3.draw(radius=0.01, color='red', alpha=.5, coord_axes=[1, 0], setlim='grow')
- _rectify_about(about)[source]¶
Ensures that about returns a specified point. Allows for special keys like center to be used.
Example
>>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10, dim=2, rng=0)
- fill(image, value, coord_axes=None, interp='bilinear')[source]¶
Sets sub-coordinate locations in a grid to a particular value
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns:
image with coordinates rasterized on it
- Return type:
ndarray
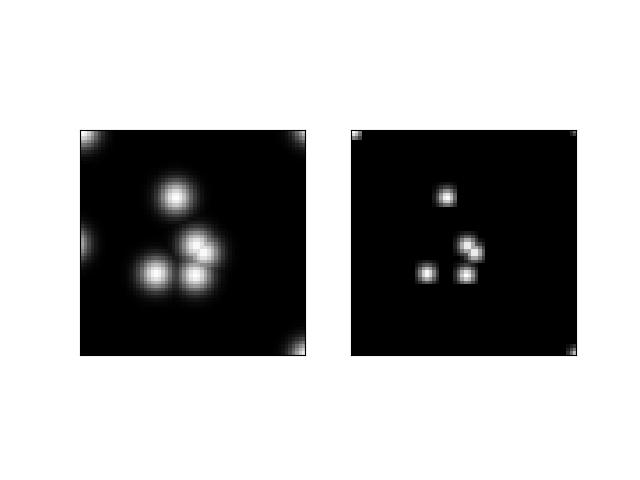
- soft_fill(image, coord_axes=None, radius=5)[source]¶
Used for drawing keypoint truth in heatmaps
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns:
image with coordinates rasterized on it
- Return type:
ndarray
References
https://stackoverflow.com/questions/54726703/generating-keypoint-heatmaps-in-tensorflow
Example
>>> from kwimage.structs.coords import * # NOQA >>> s = 64 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> # Put points on edges to to verify "edge cases" >>> self.data[1] = [0, 0] # top left >>> self.data[2] = [s, s] # bottom right >>> self.data[3] = [0, s + 10] # bottom left >>> self.data[4] = [-3, s // 2] # middle left >>> self.data[5] = [s + 1, -1] # top right >>> # Put points in the middle to verify overlap blending >>> self.data[6] = [32.5, 32.5] # middle >>> self.data[7] = [34.5, 34.5] # middle >>> fill_value = 1 >>> coord_axes = [1, 0] >>> radius = 10 >>> image1 = np.zeros((s, s)) >>> self.soft_fill(image1, coord_axes=coord_axes, radius=radius) >>> radius = 3.0 >>> image2 = np.zeros((s, s)) >>> self.soft_fill(image2, coord_axes=coord_axes, radius=radius) >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(image1, pnum=(1, 2, 1)) >>> kwplot.imshow(image2, pnum=(1, 2, 2))

- draw_on(image=None, fill_value=1, coord_axes=[1, 0], interp='bilinear')[source]¶
Note
unlike other methods, the defaults assume x/y internal data
- Parameters:
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
In other words the i-th entry in coord_axes specifies which row-major spatial dimension the i-th column of a coordinate corresponds to. The index is the coordinate dimension and the value is the axes dimension.
- Returns:
image with coordinates drawn on it
- Return type:
ndarray
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> s = 256 >>> self = Coords.random(10, meta={'shape': (s, s)}).scale(s) >>> self.data[0] = [10, 10] >>> self.data[1] = [20, 40] >>> image = np.zeros((s, s)) >>> fill_value = 1 >>> image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[0, 1], interp='nearest') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='bilinear') >>> # image = self.draw_on(image, fill_value, coord_axes=[1, 0], interp='nearest') >>> # xdoctest: +REQUIRES(--show) >>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(image) >>> self.draw(radius=3, alpha=.5, coord_axes=[1, 0])

- draw(color='blue', ax=None, alpha=None, coord_axes=[1, 0], radius=1, setlim=False)[source]¶
Draw these coordinates via matplotlib
Note
unlike other methods, the defaults assume x/y internal data
- Parameters:
setlim (bool) – if True ensures the limits of the axes contains the polygon
coord_axes (Tuple) – specify which image axes each coordinate dim corresponds to. For 2D images, if you are storing r/c data, set to [0,1], if you are storing x/y data, set to [1,0].
- Returns:
drawn matplotlib objects
- Return type:
List[mpl.collections.PatchCollection]
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> from kwimage.structs.coords import * # NOQA >>> self = Coords.random(10) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> plt = kwplot.autoplt() >>> self.draw(radius=0.05, alpha=0.8) >>> plt.gca().set_xlim(0, 1) >>> plt.gca().set_ylim(0, 1) >>> plt.gca().set_aspect('equal')