kwimage.structs.heatmap module¶
Todo
[ ] Remove doctest dependency on ndsampler?
- [ ] Remove the datakeys that tries to define what heatmap should represent
(e.g. class_probs, keypoints, etc…) and instead just focus on a data structure that stores a [C, H, W] or [H, W] tensor?
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/heatmap.py __doc__
Example
>>> # xdoctest: +REQUIRES(module:ndsampler)
>>> # xdoctest: +REQUIRES(--mask)
>>> from kwimage.structs.heatmap import * # NOQA
>>> import kwimage
>>> import ndsampler
>>> sampler = ndsampler.CocoSampler.demo('shapes')
>>> iminfo, anns = sampler.load_image_with_annots(1)
>>> image = iminfo['imdata']
>>> input_dims = image.shape[0:2]
>>> kp_classes = sampler.dset.keypoint_categories()
>>> dets = kwimage.Detections.from_coco_annots(
>>> anns, sampler.dset.dataset['categories'],
>>> sampler.catgraph, kp_classes, shape=input_dims)
>>> bg_size = [100, 100]
>>> heatmap = dets.rasterize(bg_size, input_dims, soften=2)
>>> # xdoctest: +REQUIRES(--show)
>>> import kwplot
>>> kwplot.autompl()
>>> kwplot.figure(fnum=1, doclf=True)
>>> kwplot.imshow(image)
>>> heatmap.draw(invert=True, kpts=[0, 1, 2, 3, 4])
Example
>>> # xdoctest: +REQUIRES(module:ndsampler)
>>> # xdoctest: +REQUIRES(--mask)
>>> from kwimage.structs.heatmap import * # NOQA
>>> from kwimage.structs.detections import _dets_to_fcmaps
>>> import kwimage
>>> import ndsampler
>>> sampler = ndsampler.CocoSampler.demo('shapes')
>>> iminfo, anns = sampler.load_image_with_annots(1)
>>> image = iminfo['imdata']
>>> input_dims = image.shape[0:2]
>>> kp_classes = sampler.dset.keypoint_categories()
>>> dets = kwimage.Detections.from_coco_annots(
>>> anns, sampler.dset.dataset['categories'],
>>> sampler.catgraph, kp_classes, shape=input_dims)
>>> bg_size = [100, 100]
>>> bg_idxs = sampler.catgraph.index('background')
>>> fcn_target = _dets_to_fcmaps(dets, bg_size, input_dims, bg_idxs)
>>> fcn_target.keys()
>>> print('fcn_target: ' + ub.urepr(ub.map_vals(lambda x: x.shape, fcn_target), nl=1))
>>> # xdoctest: +REQUIRES(--show)
>>> import kwplot
>>> kwplot.autompl()
>>> size_mask = fcn_target['size']
>>> dxdy_mask = fcn_target['dxdy']
>>> cidx_mask = fcn_target['cidx']
>>> kpts_mask = fcn_target['kpts']
>>> def _vizmask(dxdy_mask):
>>> dx, dy = dxdy_mask
>>> mag = np.sqrt(dx ** 2 + dy ** 2)
>>> mag /= (mag.max() + 1e-9)
>>> mask = (cidx_mask != 0).astype(np.float32)
>>> angle = np.arctan2(dy, dx)
>>> orimask = kwplot.make_orimask(angle, mask, alpha=mag)
>>> vecmask = kwplot.make_vector_field(
>>> dx, dy, stride=4, scale=0.1, thickness=1, tipLength=.2,
>>> line_type=16)
>>> return [vecmask, orimask]
>>> vecmask, orimask = _vizmask(dxdy_mask)
>>> raster = kwimage.overlay_alpha_layers(
>>> [vecmask, orimask, image], keepalpha=False)
>>> raster = dets.draw_on((raster * 255).astype(np.uint8),
>>> labels=True, alpha=None)
>>> kwplot.imshow(raster)
>>> kwplot.show_if_requested()
- class kwimage.structs.heatmap._HeatmapDrawMixin[source]¶
Bases:
objectmixin methods for drawing heatmap details
- colorize(channel=None, invert=False, with_alpha=1.0, interpolation='linear', imgspace=False, cmap=None)[source]¶
Creates a colorized version of a heatmap channel suitable for visualization
- Parameters:
channel (int | str) – index of category to visualize, or a special code indicating how to visualize multiple classes. Can be class_idx, class_probs, or class_energy.
imgspace (bool) – colorize the image after warping into the image space.
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/heatmap.py _HeatmapDrawMixin.colorize --show
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> self = Heatmap.random(rng=0, dims=(32, 32)) >>> colormask1 = self.colorize(0, imgspace=False) >>> colormask2 = self.colorize(0, imgspace=True) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(colormask1, pnum=(1, 2, 1), fnum=1, title='output space') >>> kwplot.imshow(colormask2, pnum=(1, 2, 2), fnum=1, title='image space') >>> kwplot.show_if_requested()
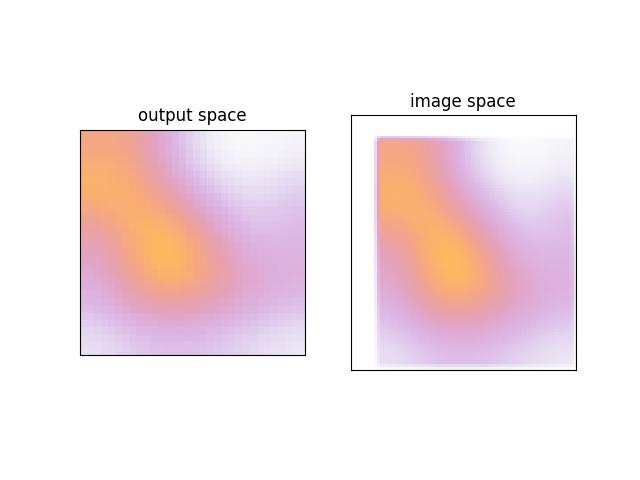
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> self = Heatmap.random(rng=0, dims=(32, 32)) >>> colormask1 = self.colorize('diameter', imgspace=False) >>> colormask2 = self.colorize('diameter', imgspace=True) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(colormask1, pnum=(1, 2, 1), fnum=1, title='output space') >>> kwplot.imshow(colormask2, pnum=(1, 2, 2), fnum=1, title='image space') >>> kwplot.show_if_requested()
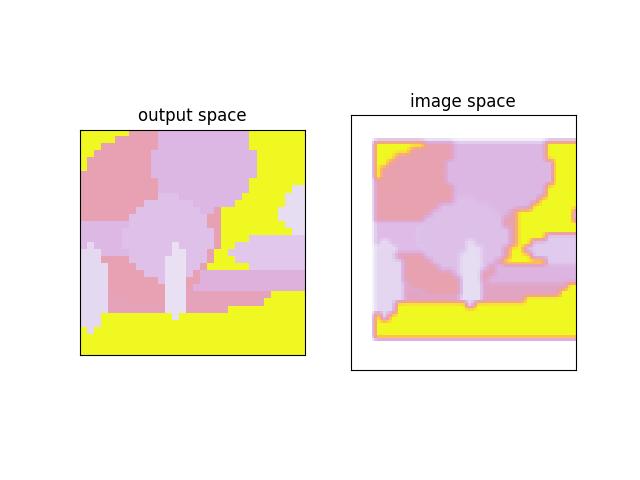
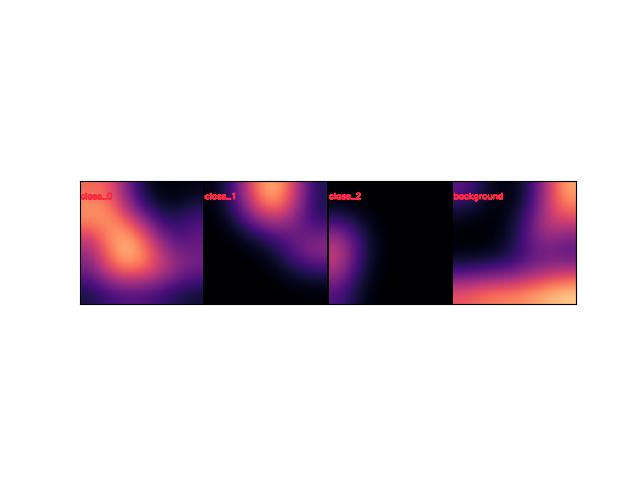
- draw_stacked(image=None, dsize=(224, 224), ignore_class_idxs={}, top=None, chosen_cxs=None)[source]¶
Draws per-class probabilities and stacks them into a single image
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> self = Heatmap.random(rng=0, dims=(32, 32)) >>> stacked = self.draw_stacked() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(stacked)
- draw(channel=None, image=None, imgspace=None, **kwargs)[source]¶
Accepts same args as draw_on, but uses maplotlib
- Parameters:
channel (int | str) – category index to visualize, or special key
- draw_on(image=None, channel=None, invert=False, with_alpha=1.0, interpolation='linear', vecs=False, kpts=None, imgspace=None)[source]¶
Overlays a heatmap channel on top of an image
- Parameters:
image (ndarray) – image to draw on, if unspecified one is created.
channel (int | str) – category index to visualize, or special key. special keys are: class_idx, class_probs, class_idx
imgspace (bool) – colorize the image after warping into the image space.
Todo
- [ ] Find a way to visualize offset, diameter, and class_probs
either individually or all at the same time
- CommandLine:
xdoctest -m /home/joncrall/code/kwimage/kwimage/structs/heatmap.py
Example
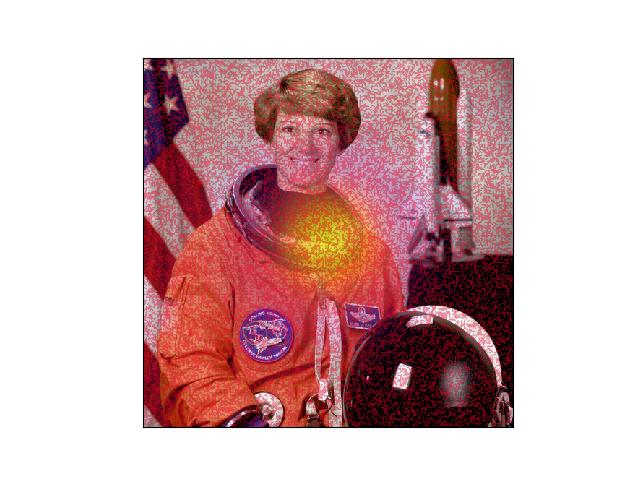
>>> # xdoctest: +REQUIRES(module:kwplot) >>> import kwarray >>> import kwimage >>> image = kwimage.grab_test_image('astro') >>> probs = kwimage.gaussian_patch(image.shape[0:2])[None, :] >>> probs = probs / probs.max() >>> class_probs = kwarray.ArrayAPI.cat([probs, 1 - probs], axis=0) >>> self = kwimage.Heatmap(class_probs=class_probs, offset=5 * np.random.randn(2, *probs.shape[1:])) >>> toshow = self.draw_on(image, 0, vecs=True, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(toshow)

Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> self = kwimage.Heatmap.random(dims=(200, 200), dets='coco', keypoints=True) >>> image = kwimage.grab_test_image('astro') >>> toshow = self.draw_on(image, 0, vecs=False, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(toshow)
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> self = kwimage.Heatmap.random(dims=(200, 200), dets='coco', keypoints=True) >>> kpts = [6] >>> self = self.warp(self.tf_data_to_img.params) >>> image = kwimage.grab_test_image('astro') >>> image = kwimage.ensure_alpha_channel(image) >>> toshow = self.draw_on(image, 0, with_alpha=0.85, kpts=kpts) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(toshow)
Example
>>> # xdoctest: +REQUIRES(module:kwplot) >>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> mask = np.random.rand(32, 32) >>> self = kwimage.Heatmap( >>> class_probs=mask, >>> img_dims=mask.shape[0:2], >>> tf_data_to_img=np.eye(3), >>> ) >>> canvas = self.draw_on() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(canvas)
- class kwimage.structs.heatmap._HeatmapWarpMixin[source]¶
Bases:
objectmixin method having to do with warping and aligning heatmaps
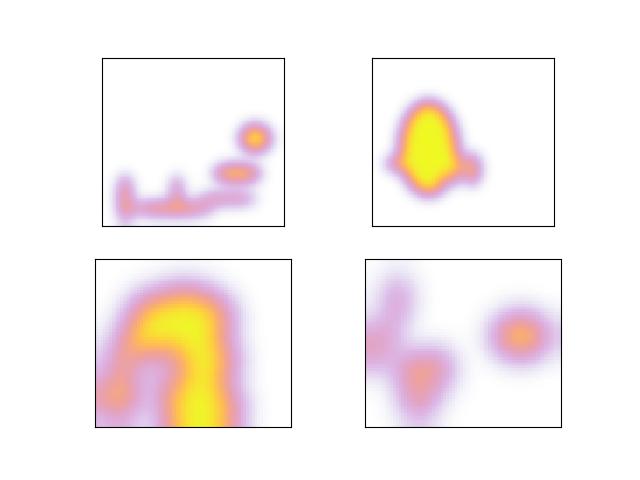
- _align_other(other)[source]¶
Warp another Heatmap (with the same underlying imgdims) into the same space as this heatmap. This lets us perform elementwise operations on the two heatmaps (like geometric mean).
- Parameters:
other (Heatmap) – the heatmap to align with self
- Returns:
warped version of other that aligns with self.
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> self = Heatmap.random((120, 130), img_dims=(200, 210), classes=2, nblips=10, rng=0) >>> other = Heatmap.random((60, 70), img_dims=(200, 210), classes=2, nblips=10, rng=1) >>> other2 = self._align_other(other) >>> assert self.shape != other.shape >>> assert self.shape == other2.shape >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(self.colorize(0, imgspace=False), fnum=1, pnum=(2, 2, 1)) >>> kwplot.imshow(self.colorize(1, imgspace=False), fnum=1, pnum=(2, 2, 2)) >>> kwplot.imshow(other.colorize(0, imgspace=False), fnum=1, pnum=(2, 2, 3)) >>> kwplot.imshow(other.colorize(1, imgspace=False), fnum=1, pnum=(2, 2, 4))
- _align(mask, interpolation='linear')[source]¶
Align a linear combination of heatmap channels with the original image
DEPRICATE
- upscale(channel=None, interpolation='linear')[source]¶
Warp the heatmap with the image dimensions
- Parameters:
channel (ndarray | None) – if None, use class probs, else chw data.
Todo
[ ] Needs refactor
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> self = Heatmap.random(rng=0, dims=(32, 32)) >>> colormask = self.upscale()
- warp(mat=None, input_dims=None, output_dims=None, interpolation='linear', modify_spatial_coords=True, int_interpolation='nearest', mat_is_xy=True, version=None)[source]¶
Warp all spatial maps. If the map contains spatial data, that data is also warped (ignoring the translation component).
- Parameters:
mat (ArrayLike) – transformation matrix
input_dims (tuple) – unused, only exists for compatibility
output_dims (tuple) – size of the output heatmap
interpolation (str) – see kwimage.warp_tensor
int_interpolation (str) – interpolation used for interger types (should be nearest)
mat_is_xy (bool) – set to false if the matrix is in yx space instead of xy space
- Returns:
this heatmap warped into a new spatial dimension
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> self = Heatmap.random(rng=0, keypoints=True) >>> S = 3.0 >>> mat = np.eye(3) * S >>> mat[-1, -1] = 1 >>> newself = self.warp(mat, np.array(self.dims) * S).numpy() >>> assert newself.offset.shape[0] == 2 >>> assert newself.diameter.shape[0] == 2 >>> f1 = newself.offset.max() / self.offset.max() >>> assert f1 == S >>> f2 = newself.diameter.max() / self.diameter.max() >>> assert f2 == S
Example
>>> import kwimage >>> # xdoctest: +REQUIRES(module:ndsampler) >>> self = kwimage.Heatmap.random(dims=(100, 100), dets='coco', keypoints=True) >>> image = np.zeros(self.img_dims) >>> # xdoctest: +REQUIRES(module:kwplot) >>> toshow = self.draw_on(image, 1, vecs=True, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(toshow)
- class kwimage.structs.heatmap._HeatmapAlgoMixin[source]¶
Bases:
objectAlgorithmic operations on heatmaps
- classmethod combine(heatmaps, root_index=None, dtype=<class 'numpy.float32'>)[source]¶
Combine multiple heatmaps into a single heatmap.
- Parameters:
heatmaps (Sequence[Heatmap]) – multiple heatmaps to combine into one
root_index (int) – which heatmap in the sequence to align other heatmaps with
- Returns:
the combined heatmap
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> a = Heatmap.random((120, 130), img_dims=(200, 210), classes=2, nblips=10, rng=0) >>> b = Heatmap.random((60, 70), img_dims=(200, 210), classes=2, nblips=10, rng=1) >>> c = Heatmap.random((40, 30), img_dims=(200, 210), classes=2, nblips=10, rng=1) >>> heatmaps = [a, b, c] >>> newself = Heatmap.combine(heatmaps, root_index=2) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(a.colorize(0, imgspace=1), fnum=1, pnum=(4, 2, 1)) >>> kwplot.imshow(a.colorize(1, imgspace=1), fnum=1, pnum=(4, 2, 2)) >>> kwplot.imshow(b.colorize(0, imgspace=1), fnum=1, pnum=(4, 2, 3)) >>> kwplot.imshow(b.colorize(1, imgspace=1), fnum=1, pnum=(4, 2, 4)) >>> kwplot.imshow(c.colorize(0, imgspace=1), fnum=1, pnum=(4, 2, 5)) >>> kwplot.imshow(c.colorize(1, imgspace=1), fnum=1, pnum=(4, 2, 6)) >>> kwplot.imshow(newself.colorize(0, imgspace=1), fnum=1, pnum=(4, 2, 7)) >>> kwplot.imshow(newself.colorize(1, imgspace=1), fnum=1, pnum=(4, 2, 8)) >>> # xdoctest: +REQUIRES(--show) >>> kwplot.imshow(a.colorize('offset', imgspace=1), fnum=2, pnum=(4, 1, 1)) >>> kwplot.imshow(b.colorize('offset', imgspace=1), fnum=2, pnum=(4, 1, 2)) >>> kwplot.imshow(c.colorize('offset', imgspace=1), fnum=2, pnum=(4, 1, 3)) >>> kwplot.imshow(newself.colorize('offset', imgspace=1), fnum=2, pnum=(4, 1, 4)) >>> # xdoctest: +REQUIRES(--show) >>> kwplot.imshow(a.colorize('diameter', imgspace=1), fnum=3, pnum=(4, 1, 1)) >>> kwplot.imshow(b.colorize('diameter', imgspace=1), fnum=3, pnum=(4, 1, 2)) >>> kwplot.imshow(c.colorize('diameter', imgspace=1), fnum=3, pnum=(4, 1, 3)) >>> kwplot.imshow(newself.colorize('diameter', imgspace=1), fnum=3, pnum=(4, 1, 4))
- detect(channel, invert=False, min_score=0.01, num_min=10, max_dims=None, min_dims=None, dim_thresh_space='image')[source]¶
Lossy conversion from a Heatmap to a Detections object.
For efficiency, the detections are returned in the same space as the heatmap, which usually some downsampled version of the image space. This is because it is more efficient to transform the detections into image-space after non-max supression is applied.
- Parameters:
channel (int | ArrayLike) – class index to detect objects in. Alternatively, channel can be a custom probability map as long as its dimension agree with the heatmap.
invert (bool) – if True, inverts the probabilities in the chosen channel. (Useful if you have a background channel but want to detect foreground objects).
min_score (float) – probability threshold required for a pixel to be converted into a detection. Defaults to 0.1
num_min (int) – always return at least nmin of the highest scoring detections even if they aren’t above the min_score threshold. Defaults to 10.
max_dims (Tuple[int, int]) – maximum height / width of detections By default these are expected to be in image-space.
min_dims (Tuple[int, int]) – minimum height / width of detections By default these are expected to be in image-space.
dim_thresh_space (str) – When dim_thresh_space==’native’, dimension thresholds (e.g. min_dims and max_dims) are specified in the native heatmap space (i.e. usually a downsampled space). If dim_thresh_space==’image’, then dimension thresholds are interpreted in the original image space. Defaults to ‘image’
- Returns:
raw detections.
Note that these detections will not have class_idx populated
It is the users responsbility to run non-max suppression on these results to remove duplicate detections.
- Return type:
- SeeAlso:
Detections.rasterize
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> from kwimage.structs.heatmap import * # NOQA >>> import ndsampler >>> self = Heatmap.random(rng=2, dims=(32, 32)) >>> dets = self.detect(channel=0, max_dims=7, num_min=None) >>> img_dets = dets.warp(self.tf_data_to_img) >>> assert img_dets.boxes.to_xywh().width.max() <= 7 >>> assert img_dets.boxes.to_xywh().height.max() <= 7 >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets1 = dets.sort().take(range(30)) >>> colormask1 = self.colorize(0, imgspace=False) >>> kwplot.imshow(colormask1, pnum=(1, 2, 1), fnum=1, title='output space') >>> dets1.draw() >>> # Transform heatmap and detections into image space. >>> dets2 = dets1.warp(self.tf_data_to_img) >>> colormask2 = self.colorize(0, imgspace=True) >>> kwplot.imshow(colormask2, pnum=(1, 2, 2), fnum=1, title='image space') >>> dets2.draw()
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> from kwimage.structs.heatmap import * # NOQA >>> import torch >>> import ndsampler >>> catgraph = ndsampler.CategoryTree.demo() >>> class_energy = torch.rand(len(catgraph), 32, 32) >>> class_probs = catgraph.hierarchical_softmax(class_energy, dim=0) >>> self = Heatmap.random(rng=0, dims=(32, 32), classes=catgraph, keypoints=True) >>> print(ub.urepr(ub.map_vals(lambda x: x.shape, self.data), nl=1)) >>> self.data['class_probs'] = class_probs.numpy() >>> channel = catgraph.index('background') >>> dets = self.detect(channel, invert=True) >>> class_idx, scores = catgraph.decision(dets.probs, dim=1) >>> dets.data['class_idx'] = class_idx >>> dets.data['scores'] = scores >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> dets1 = dets.sort().take(range(10)) >>> colormask1 = self.colorize(0, imgspace=False) >>> kwplot.imshow(colormask1, pnum=(1, 2, 1), fnum=1, title='output space') >>> dets1.draw(radius=1.0) >>> # Transform heatmap and detections into image space. >>> colormask2 = self.colorize(0, imgspace=True) >>> dets2 = dets1.warp(self.tf_data_to_img) >>> kwplot.imshow(colormask2, pnum=(1, 2, 2), fnum=1, title='image space') >>> dets2.draw(radius=1.0)
- class kwimage.structs.heatmap.Heatmap(data=None, meta=None, **kwargs)[source]¶
Bases:
Spatial,_HeatmapDrawMixin,_HeatmapWarpMixin,_HeatmapAlgoMixinKeeps track of a downscaled heatmap and how to transform it to overlay the original input image. Heatmaps generally are used to estimate class probabilites at each pixel. This data struction additionally contains logic to augment pixel with offset (dydx) and scale (diamter) information.
- Variables:
data (Dict[str, ArrayLike]) –
dictionary containing spatially aligned heatmap data. Valid keys are as follows.
- class_probs (ArrayLike[C, H, W] | ArrayLike[C, D, H, W]):
A probability map for each class. C is the number of classes.
- offset (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object center position offset in y,x / t,y,x coordinates
- diamter (ArrayLike[2, H, W] | ArrayLike[3, D, H, W], optional):
object bounding box sizes in h,w / d,h,w coordinates
- keypoints (ArrayLike[2, K, H, W] | ArrayLike[3, K, D, H, W], optional):
y/x offsets for K different keypoint classes
dictionary containing miscellanious metadata about the heatmap data. Valid keys are as follows.
- img_dims (Tuple[H, W] | Tuple[D, H, W]):
original image dimension
- tf_data_to_image (SKImageGeometricTransform):
transformation matrix (typically similarity or affine) that projects the given, heatmap onto the image dimensions such that the image and heatmap are spatially aligned.
- classes (List[str] | ndsampler.CategoryTree):
information about which index in
data['class_probs']corresponds to which semantic class.
dims (Tuple) – dimensions of the heatmap (See
image_dims) for the original image dimensions.**kwargs – any key that is accepted by the data or meta dictionaries can be specified as a keyword argument to this class and it will be properly placed in the appropriate internal dictionary.
CommandLine
xdoctest -m ~/code/kwimage/kwimage/structs/heatmap.py Heatmap --show
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> from kwimage.structs.heatmap import * # NOQA >>> import skimage >>> import kwimage >>> class_probs = kwimage.grab_test_image(dsize=(32, 32), space='gray')[None, ..., 0] / 255.0 >>> img_dims = (220, 220) >>> tf_data_to_img = skimage.transform.AffineTransform(translation=(-18, -18), scale=(8, 8)) >>> self = Heatmap(class_probs=class_probs, img_dims=img_dims, >>> tf_data_to_img=tf_data_to_img) >>> aligned = self.upscale() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(aligned[0]) >>> kwplot.show_if_requested()

Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> self = Heatmap.random() >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> self.draw()
- property shape¶
- property bounds¶
- property dims¶
space-time dimensions of this heatmap
- property _impl¶
Returns the internal tensor/numpy ArrayAPI implementation
- Returns:
kwarray.ArrayAPI
- classmethod random(dims=(10, 10), classes=3, diameter=True, offset=True, keypoints=False, img_dims=None, dets=None, nblips=10, noise=0.0, smooth_k=3, rng=None, ensure_background=True)[source]¶
Creates dummy data, suitable for use in tests and benchmarks
- Parameters:
dims (Tuple[int, int]) – dimensions of the heatmap
classes (int | List[str] | kwcoco.CategoryTree) – foreground classes
diameter (bool) – if True, include a “diameter” heatmap
offset (bool) – if True, include an “offset” heatmap
keypoints (bool)
smooth_k (int) – kernel size for gaussian blur to smooth out the heatmaps.
img_dims (Tuple) – dimensions of an upscaled image the heatmap corresponds to. (This should be removed and simply handled with a transform
in the future).
- Returns:
Heatmap
Example
>>> from kwimage.structs.heatmap import * # NOQA >>> self = Heatmap.random((128, 128), img_dims=(200, 200), >>> classes=3, nblips=10, rng=0, noise=0.1) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.imshow(self.colorize(0, imgspace=0), fnum=1, pnum=(1, 4, 1), doclf=1) >>> kwplot.imshow(self.colorize(1, imgspace=0), fnum=1, pnum=(1, 4, 2)) >>> kwplot.imshow(self.colorize(2, imgspace=0), fnum=1, pnum=(1, 4, 3)) >>> kwplot.imshow(self.colorize(3, imgspace=0), fnum=1, pnum=(1, 4, 4))
Example
>>> # xdoctest: +REQUIRES(module:ndsampler) >>> import kwimage >>> self = kwimage.Heatmap.random(dims=(50, 200), dets='coco', >>> keypoints=True) >>> image = np.zeros(self.img_dims) >>> # xdoctest: +REQUIRES(module:kwplot) >>> toshow = self.draw_on(image, 1, vecs=True, kpts=0, with_alpha=0.85) >>> # xdoctest: +REQUIRES(--show) >>> import kwplot >>> kwplot.autompl() >>> kwplot.figure(fnum=1, doclf=True) >>> kwplot.imshow(toshow)
- property class_probs¶
- property offset¶
- property diameter¶
- property img_dims¶
- property tf_data_to_img¶
- property classes¶
- kwimage.structs.heatmap._prob_to_dets(probs, diameter=None, offset=None, class_probs=None, keypoints=None, min_score=0.01, num_min=10, max_dims=None, min_dims=None)[source]¶
Directly convert a one-channel probability map into a Detections object.
Helper for Heatmap.detect
It does this by converting each pixel above a threshold in a probability map to a detection with a specified diameter.
- Parameters:
probs (ArrayLike[H, W]) – liklihood that each particular pixel should be detected as an object.
diameter (ArrayLike[2, H, W] | Tuple) – H, W sizes for the bounding box at each pixel location. If passed as a tuple, then all boxes receive that diameter.
offset (Tuple | ArrayLike[2, H, W]) – Y, X offsets from the pixel location to the bounding box center. If passed as a tuple, then all boxes receive that offset.
class_probs (ArrayLike[C, H, W], optional) – probabilities for each class at each pixel location. If specified, this will populate the probs attribute of the returned Detections object.
keypoints (ArrayLike[2, K, H, W], optional) – Keypoint predictions for all keypoint classes
min_score (float) – probability threshold required for a pixel to be converted into a detection. Defaults to 0.1
num_min (int) – always return at least nmin of the highest scoring detections even if they aren’t above the min_score threshold. Defaults to 10
- Returns:
- raw detections. It is the users responsbility to
run non-max suppression on these results to remove duplicate detections.
- Return type:
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import torch >>> rng = np.random.RandomState(0) >>> probs = rng.rand(3, 3).astype(np.float32) >>> min_score = .5 >>> diameter = [10, 10] >>> dets = _prob_to_dets(probs, diameter, min_score=min_score) >>> assert dets.boxes.data.dtype.kind == 'f' >>> assert len(dets) == 9 >>> dets = _prob_to_dets(torch.FloatTensor(probs), diameter, min_score=min_score) >>> assert dets.boxes.data.dtype.is_floating_point >>> assert len(dets) == 9
Example
>>> # xdoctest: +REQUIRES(module:torch) >>> import kwimage >>> from kwimage.structs.heatmap import * >>> from kwimage.structs.heatmap import _prob_to_dets >>> heatmap = kwimage.Heatmap.random(rng=0, dims=(3, 3), keypoints=True) >>> # Try with numpy >>> min_score = .5 >>> dets = _prob_to_dets(heatmap.class_probs[0], heatmap.diameter, >>> heatmap.offset, heatmap.class_probs, >>> heatmap.data['keypoints'], >>> min_score) >>> assert dets.boxes.data.dtype.kind == 'f' >>> assert 'keypoints' in dets.data >>> dets_np = dets >>> # Try with torch >>> heatmap = heatmap.tensor() >>> dets = _prob_to_dets(heatmap.class_probs[0], heatmap.diameter, >>> heatmap.offset, heatmap.class_probs, >>> heatmap.data['keypoints'], >>> min_score) >>> assert dets.boxes.data.dtype.is_floating_point >>> assert len(dets) == len(dets_np) >>> dets_torch = dets >>> assert np.all(dets_torch.numpy().boxes.data == dets_np.boxes.data)
Example
>>> heatmap = Heatmap.random(rng=0, dims=(3, 3), diameter=1) >>> probs = heatmap.class_probs[0] >>> diameter = heatmap.diameter >>> offset = heatmap.offset >>> class_probs = heatmap.class_probs >>> min_score = 0.5 >>> dets = _prob_to_dets(probs, diameter, offset, class_probs, None, min_score)
- kwimage.structs.heatmap.smooth_prob(prob, k=3, inplace=False, eps=1e-09)[source]¶
Smooths the probability map, but preserves the magnitude of the peaks.
Note
even if inplace is true, we still need to make a copy of the input array, however, we do ensure that it is cleaned up before we leave the function scope.
sigma=0.8 @ k=3, sigma=1.1 @ k=5, sigma=1.4 @ k=7
- kwimage.structs.heatmap._remove_translation(tf)[source]¶
Removes the translation component of a transform
Todo
[ ] Is this possible in more general cases? E.g. projective transforms?
- kwimage.structs.heatmap._gmean(a, axis=0, clobber=False)[source]¶
Compute the geometric mean along the specified axis.
Modification of the scipy.mstats method to be more memory efficient
- Example
>>> rng = np.random.RandomState(0) >>> C, H, W = 8, 32, 32 >>> axis = 0 >>> a = rng.rand(2, C, H, W) >>> _gmean(a)